Abstract
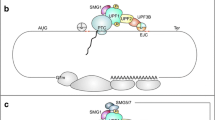
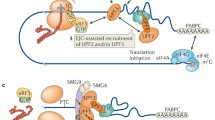
Nonsense-mediated mRNA decay (NMD) is a translation-dependent, multistep process that degrades irregular or faulty messenger RNAs (mRNAs). NMD mainly targets mRNAs with a truncated open reading frame (ORF) due to premature termination codons (PTCs). In addition, NMD also regulates the expression of different types of endogenous mRNA substrates. A multitude of factors are involved in the tight regulation of the NMD mechanism. In this review, we focus on the molecular mechanism of mammalian NMD. Based on the published data, we discuss the involvement of translation termination in NMD initiation. Furthermore, we provide a detailed overview of the core NMD machinery, as well as several peripheral NMD factors, and discuss their function. Finally, we present an overview of diseases associated with NMD factor mutations and summarize the current state of treatment for genetic disorders caused by nonsense mutations.




Similar content being viewed by others
References
Brogna S, Wen J (2009) Nonsense-mediated mRNA decay (NMD) mechanisms. Nat Struct Mol Biol 16:107–113. doi:10.1038/nsmb.1550
Fatscher T, Boehm V, Weiche B, Gehring NH (2014) The interaction of cytoplasmic poly(A)-binding protein with eukaryotic initiation factor 4G suppresses nonsense-mediated mRNA decay. RNA 20:1579–1592. doi:10.1261/rna.044933.114
Ivanov PV, Gehring NH, Kunz JB, Hentze MW, Kulozik AE (2008) Interactions between UPF1, eRFs, PABP and the exon junction complex suggest an integrated model for mammalian NMD pathways. EMBO J 27:736–747. doi:10.1038/emboj.2008.17
Joncourt R, Eberle AB, Rufener SC, Muhlemann O (2014) Eukaryotic initiation factor 4G suppresses nonsense-mediated mRNA decay by two genetically separable mechanisms. PLoS One 9:e104391. doi:10.1371/journal.pone.0104391
Singh G, Rebbapragada I, Lykke-Andersen J (2008) A competition between stimulators and antagonists of Upf complex recruitment governs human nonsense-mediated mRNA decay. PLoS Biol 6:e111. doi:10.1371/journal.pbio.0060111
Janzen DM, Geballe AP (2004) The effect of eukaryotic release factor depletion on translation termination in human cell lines. Nucleic Acids Res 32:4491–4502. doi:10.1093/nar/gkh791
Chauvin C, Salhi S, Le Goff C, Viranaicken W, Diop D, Jean-Jean O (2005) Involvement of human release factors eRF3a and eRF3b in translation termination and regulation of the termination complex formation. Mol Cell Biol 25:5801–5811. doi:10.1128/MCB.25.14.5801-5811.2005
Hoshino S, Imai M, Mizutani M, Kikuchi Y, Hanaoka F, Ui M, Katada T (1998) Molecular cloning of a novel member of the eukaryotic polypeptide chain-releasing factors (eRF) - Its identification as eRF3 interacting with eRF1. J Biol Chem 273:22254–22259. doi:10.1074/jbc.273.35.22254
Hoshino S, Miyazawa H, Enomoto T, Hanaoka F, Kikuchi Y, Kikuchi A, Ui M (1989) A human homolog of the yeast Gst1-gene codes for a Gtp-binding protein and is expressed in a proliferation-dependent manner in mammalian-cells. EMBO J 8:3807–3814
Chavatte L, Frolova L, Kisselev L, Favre A (2001) The polypeptide chain release factor eRF1 specifically contacts the s(4)UGA stop codon located in the A site of eukaryotic ribosomes. Eur J Biochem 268:2896–2904
Frolova L, Le Goff X, Rasmussen HH, Cheperegin S, Drugeon G, Kress M, Arman I, Haenni AL, Celis JE, Philippe M et al (1994) A highly conserved eukaryotic protein family possessing properties of polypeptide chain release factor. Nature 372:701–703. doi:10.1038/372701a0
Frolova LY, Tsivkovskii RY, Sivolobova GF, Oparina NY, Serpinsky OI, Blinov VM, Tatkov SI, Kisselev LL (1999) Mutations in the highly conserved GGQ motif of class 1 polypeptide release factors abolish ability of human eRF1 to trigger peptidyl-tRNA hydrolysis. RNA 5:1014–1020. doi:10.1017/s135583829999043x
Song H, Mugnier P, Das AK, Webb HM, Evans DR, Tuite MF, Hemmings BA, Barford D (2000) The crystal structure of human eukaryotic release factor eRF1—mechanism of stop codon recognition and peptidyl-tRNA hydrolysis. Cell 100:311–321
Zhouravleva G, Frolova L, Le Goff X, Le Guellec R, Inge-Vechtomov S, Kisselev L, Philippe M (1995) Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3. EMBO J 14:4065–4072
Salas-Marco J, Bedwell DM (2004) GTP hydrolysis by eRF3 facilitates stop codon decoding during eukaryotic translation termination. Mol Cell Biol 24:7769–7778. doi:10.1128/MCB.24.17.7769-7778.2004
Frolova L, LeGoff X, Zhouravleva G, Davydova E, Philippe M, Kisselev L (1996) Eukaryotic polypeptide chain release factor eRF3 is an eRF1- and ribosome-dependent guanosine triphosphatase. RNA 2:334–341
Alkalaeva EZ, Pisarev AV, Frolova LY, Kisselev LL, Pestova TV (2006) In vitro reconstitution of eukaryotic translation reveals cooperativity between release factors eRF1 and eRF3. Cell 125:1125–1136. doi:10.1016/j.cell.2006.04.035
Pisarev AV, Skabkin MA, Pisareva VP, Skabkina OV, Rakotondrafara AM, Hentze MW, Hellen CU, Pestova TV (2010) The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol Cell 37:196–210. doi:10.1016/j.molcel.2009.12.034
Barthelme D, Dinkelaker S, Albers SV, Londei P, Ermler U, Tampe R (2011) Ribosome recycling depends on a mechanistic link between the FeS cluster domain and a conformational switch of the twin-ATPase ABCE1. Proc Natl Acad Sci USA 108:3228–3233. doi:10.1073/pnas.1015953108
Becker T, Franckenberg S, Wickles S, Shoemaker CJ, Anger AM, Armache JP, Sieber H, Ungewickell C, Berninghausen O, Daberkow I, Karcher A, Thomm M, Hopfner KP, Green R, Beckmann R (2012) Structural basis of highly conserved ribosome recycling in eukaryotes and archaea. Nature 482:501–506. doi:10.1038/nature10829
Adam SA, Nakagawa T, Swanson MS, Woodruff TK, Dreyfuss G (1986) mRNA polyadenylate-binding protein: gene isolation and sequencing and identification of a ribonucleoprotein consensus sequence. Mol Cell Biol 6:2932–2943
Sachs AB, Bond MW, Kornberg RD (1986) A single gene from yeast for both nuclear and cytoplasmic polyadenylate-binding proteins—domain-structure and expression. Cell 45:827–835. doi:10.1016/0092-8674(86)90557-X
Deo RC, Bonanno JB, Sonenberg N, Burley SK (1999) Recognition of polyadenylate RNA by the poly(A)-binding protein. Cell 98:835–845
Cosson B, Berkova N, Couturier A, Chabelskaya S, Philippe M, Zhouravleva G (2002) Poly(A)-binding protein and eRF3 are associated in vivo in human and Xenopus cells. Biol Cell 94:205–216. doi:10.1016/S0248-4900(02)01194-2
Kozlov G, Trempe JF, Khaleghpour K, Kahvejian A, Ekiel I, Gehring K (2001) Structure and function of the C-terminal PABC domain of human poly(A)-binding protein. Proc Natl Acad Sci USA 98:4409–4413. doi:10.1073/pnas.071024998
Si Hoshino, Imai M, Kobayashi T, Uchida N, Katada T (1999) The eukaryotic polypeptide chain releasing factor (eRF3/GSPT) carrying the translation termination signal to the 3′-Poly(A) tail of mRNA. Direct association of eRF3/GSPT with polyadenylate-binding protein. J Biol Chem 274:16677–16680. doi:10.1074/jbc.274.24.16677
Uchida N, Hoshino S, Imataka H, Sonenberg N, Katada T (2002) A novel role of the mammalian GSPT/eRF3 associating with poly(A)-binding protein in Cap/Poly(A)-dependent translation. J Biol Chem 277:50286–50292. doi:10.1074/jbc.M203029200
Kozlov G, Gehring K (2010) Molecular basis of eRF3 recognition by the MLLE domain of poly(A)-binding protein. PLoS One 5:e10169. doi:10.1371/journal.pone.0010169
Tarun SZ Jr, Sachs AB (1996) Association of the yeast poly(A) tail binding protein with translation initiation factor eIF-4G. EMBO J 15:7168–7177
Safaee N, Kozlov G, Noronha AM, Xie J, Wilds CJ, Gehring K (2012) Interdomain allostery promotes assembly of the poly(A) mRNA complex with PABP and eIF4G. Mol Cell 48:375–386. doi:10.1016/j.molcel.2012.09.001
Marcotrigiano J, Gingras AC, Sonenberg N, Burley SK (1997) Cocrystal structure of the messenger RNA 5′ cap-binding protein (eIF4E) bound to 7-methyl-GDP. Cell 89:951–961
Amrani N, Ghosh S, Mangus DA, Jacobson A (2008) Translation factors promote the formation of two states of the closed-loop mRNP. Nature 453:1276–1280. doi:10.1038/nature06974
Kahvejian A, Svitkin YV, Sukarieh R, M’Boutchou MN, Sonenberg N (2005) Mammalian poly(A)-binding protein is a eukaryotic translation initiation factor, which acts via multiple mechanisms. Genes Dev 19:104–113. doi:10.1101/gad.1262905
Wells SE, Hillner PE, Vale RD, Sachs AB (1998) Circularization of mRNA by eukaryotic translation initiation factors. Mol Cell 2:135–140
Nagy E, Maquat LE (1998) A rule for termination-codon position within intron-containing genes: when nonsense affects RNA abundance. Trends Biochem Sci 23:198–199. doi:10.1016/S0968-0004(98)01208-0
Thermann R, Neu-Yilik G, Deters A, Frede U, Wehr K, Hagemeier C, Hentze MW, Kulozik AE (1998) Binary specification of nonsense codons by splicing and cytoplasmic translation. EMBO J 17:3484–3494. doi:10.1093/emboj/17.12.3484
Zhang J, Sun X, Qian Y, LaDuca JP (1998) At least one intron is required for the nonsense-mediated decay of triosephosphate isomerase mRNA: a possible link between nuclear splicing and cytoplasmic translation. Mol Cell Biol 18:5272–5283
Zhang J, Sun XL, Qian YM, Maquat LE (1998) Intron function in the nonsense-mediated decay of beta-globin mRNA: indications that pre-mRNA splicing in the nucleus can influence mRNA translation in the cytoplasm. RNA 4:801–815. doi:10.1017/S1355838298971849
Wang J, Gudikote JP, Olivas OR, Wilkinson MF (2002) Boundary-independent polar nonsense-mediated decay. EMBO Rep 3:274–279. doi:10.1093/embo-reports/kvf036
Brocke KS, Neu-Yilik G, Gehring NH, Hentze MW, Kulozik AE (2002) The human intronless melanocortin 4-receptor gene is NMD insensitive. Hum Mol Genet 11:331–335. doi:10.1093/hmg/11.3.331
Maquat LE, Li XJ (2001) Mammalian heat shock p70 and histone H4 transcripts, which derive from naturally intronless genes, are immune to nonsense-mediated decay. RNA 7:445–456. doi:10.1017/S1355838201002229
Neu-Yilik G, Gehring NH, Thermann R, Frede U, Hentze MW, Kulozik AE (2001) Splicing and 3′ end formation in the definition of nonsense-mediated decay-competent human beta-globin mRNPs. EMBO J 20:532–540. doi:10.1093/emboj/20.3.532
Dostie J, Dreyfuss G (2002) Translation is required to remove Y14 from mRNAs in the cytoplasm. Curr Biol 12:1060–1067
Lejeune F, Ishigaki Y, Li X, Maquat LE (2002) The exon junction complex is detected on CBP80-bound but not eIF4E-bound mRNA in mammalian cells: dynamics of mRNP remodeling. EMBO J 21:3536–3545. doi:10.1093/emboj/cdf345
Le Hir H, Izaurralde E, Maquat LE, Moore MJ (2000) The spliceosome deposits multiple proteins 20-24 nucleotides upstream of mRNA exon–exon junctions. EMBO J 19:6860–6869. doi:10.1093/emboj/19.24.6860
Steckelberg A-LL, Boehm V, Gromadzka AM, Gehring NH (2012) CWC22 connects pre-mRNA splicing and exon junction complex assembly. Cell Rep 2:454–461. doi:10.1016/j.celrep.2012.08.017
Gehring NH, Lamprinaki S, Kulozik AE, Hentze MW (2009) Disassembly of exon junction complexes by PYM. Cell 137:536–548. doi:10.1016/j.cell.2009.02.042
Ballut L, Marchadier B, Baguet A, Tomasetto C, Seraphin B, Le Hir H (2005) The exon junction core complex is locked onto RNA by inhibition of eIF4AIII ATPase activity. Nat Struct Mol Biol 12:861–869. doi:10.1038/nsmb990
Bono F, Ebert J, Lorentzen E, Conti E (2006) The crystal structure of the exon junction complex reveals how it maintains a stable grip on mRNA. Cell 126:713–725. doi:10.1016/j.cell.2006.08.006
Andersen CB, Ballut L, Johansen JS, Chamieh H, Nielsen KH, Oliveira CL, Pedersen JS, Seraphin B, Le Hir H, Andersen GR (2006) Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science 313:1968–1972. doi:10.1126/science.1131981
Kim VN, Kataoka N, Dreyfuss G (2001) Role of the nonsense-mediated decay factor hUpf3 in the splicing-dependent exon–exon junction complex. Science 293:1832–1836. doi:10.1126/science.1062829
Le Hir H, Gatfield D, Izaurralde E, Moore MJ (2001) The exon–exon junction complex provides a binding platform for factors involved in mRNA export and nonsense-mediated mRNA decay. EMBO J 20:4987–4997. doi:10.1093/emboj/20.17.4987
Gehring NH, Neu-Yilik G, Schell T, Hentze MW, Kulozik AE (2003) Y14 and hUpf3b form an NMD-activating complex. Mol Cell 11:939–949. doi:10.1016/S1097-2765(03)00142-4
Buhler M, Steiner S, Mohn F, Paillusson A, Muhlemann O (2006) EJC-independent degradation of nonsense immunoglobulin-mu mRNA depends on 3′ UTR length. Nat Struct Mol Biol 13:462–464. doi:10.1038/nsmb1081
Boehm V, Haberman N, Ottens F, Ule J, Gehring NH (2014) 3′ UTR length and messenger ribonucleoprotein composition determine endocleavage efficiencies at termination codons. Cell Rep 9:555–568. doi:10.1016/j.celrep.2014.09.012
Toma KG, Rebbapragada I, Durand S, Lykke-Andersen J (2015) Identification of elements in human long 3′ UTRs that inhibit nonsense-mediated decay. RNA 21:887–897. doi:10.1261/rna.048637.114
Hurt JA, Robertson AD, Burge CB (2013) Global analyses of UPF1 binding and function reveal expanded scope of nonsense-mediated mRNA decay. Genome Res 23:1636–1650. doi:10.1101/gr.157354.113
Kurosaki T, Maquat LE (2013) Rules that govern UPF1 binding to mRNA 3′ UTRs. Proc Natl Acad Sci USA 110:3357–3362. doi:10.1073/pnas.1219908110
Zund D, Gruber AR, Zavolan M, Muhlemann O (2013) Translation-dependent displacement of UPF1 from coding sequences causes its enrichment in 3′ UTRs. Nat Struct Mol Biol 20:936–943. doi:10.1038/nsmb.2635
Amrani N, Ganesan R, Kervestin S, Mangus DA, Ghosh S, Jacobson A (2004) A faux 3′-UTR promotes aberrant termination and triggers nonsense-mediated mRNA decay. Nature 432:112–118. doi:10.1038/nature03060
Muhlrad D, Parker R (1999) Aberrant mRNAs with extended 3′ UTRs are substrates for rapid degradation by mRNA surveillance. RNA 5:1299–1307
Ishigaki Y, Li X, Serin G, Maquat LE (2001) Evidence for a pioneer round of mRNA translation: mRNAs subject to nonsense-mediated decay in mammalian cells are bound by CBP80 and CBP20. Cell 106:607–617
Durand S, Lykke-Andersen J (2013) Nonsense-mediated mRNA decay occurs during eIF4F-dependent translation in human cells. Nat Struct Mol Biol 20:702–709. doi:10.1038/nsmb.2575
Rufener SC, Muhlemann O (2013) eIF4E-bound mRNPs are substrates for nonsense-mediated mRNA decay in mammalian cells. Nat Struct Mol Biol 20:710–717. doi:10.1038/nsmb.2576
Belgrader P, Cheng J, Maquat LE (1993) Evidence to implicate translation by ribosomes in the mechanism by which nonsense codons reduce the nuclear level of human triosephosphate isomerase mRNA. Proc Natl Acad Sci USA 90:482–486
Carter MS, Doskow J, Morris P, Li SL, Nhim RP, Sandstedt S, Wilkinson MF (1995) A regulatory mechanism that detects premature nonsense codons in T-cell receptor transcripts in-vivo is reversed by protein-synthesis inhibitors in-vitro. J Biol Chem 270:28995–29003
Gradi A, Svitkin YV, Imataka H, Sonenberg N (1998) Proteolysis of human eukaryotic translation initiation factor eIF4GII, but not eIF4GI, coincides with the shutoff of host protein synthesis after poliovirus infection. Proc Natl Acad Sci USA 95:11089–11094
Kuyumcu-Martinez NM, Joachims M, Lloyd RE (2002) Efficient cleavage of ribosome-associated poly(A)-binding protein by enterovirus 3C protease. J Virol 76:2062–2074
Zhang J, Maquat LE (1997) Evidence that translation reinitiation abrogates nonsense-mediated mRNA decay in mammalian cells. EMBO J 16:826–833. doi:10.1093/emboj/16.4.826
Neu-Yilik G, Amthor B, Gehring NH, Bahri S, Paidassi H, Hentze MW, Kulozik AE (2011) Mechanism of escape from nonsense-mediated mRNA decay of human beta-globin transcripts with nonsense mutations in the first exon. RNA 17:843–854. doi:10.1261/rna.2401811
Baserga SJ, Benz EJ Jr (1988) Nonsense mutations in the human beta-globin gene affect mRNA metabolism. Proc Natl Acad Sci USA 85:2056–2060
Pergolizzi R, Spritz RA, Spence S, Goossens M, Kan YW, Bank A (1981) Two cloned beta thalassemia genes are associated with amber mutations at codon 39. Nucleic Acids Res 9:7065–7072
Peixeiro I, Inacio A, Barbosa C, Silva AL, Liebhaber SA, Romao L (2012) Interaction of PABPC1 with the translation initiation complex is critical to the NMD resistance of AUG-proximal nonsense mutations. Nucleic Acids Res 40:1160–1173. doi:10.1093/nar/gkr820
Kashima I, Yamashita A, Izumi N, Kataoka N, Morishita R, Hoshino S, Ohno M, Dreyfuss G, Ohno S (2006) Binding of a novel SMG-1-Upf1-eRF1-eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev 20:355–367. doi:10.1101/gad.1389006
Yepiskoposyan H, Aeschimann F, Nilsson D, Okoniewski M, Muhlemann O (2011) Autoregulation of the nonsense-mediated mRNA decay pathway in human cells. RNA 17:2108–2118. doi:10.1261/rna.030247.111
Eberle AB, Stalder L, Mathys H, Orozco RZ, Muhlemann O (2008) Posttranscriptional gene regulation by spatial rearrangement of the 3′ untranslated region. PLoS Biol 6:e92. doi:10.1371/journal.pbio.0060092
Lee BJ, Worland PJ, Davis JN, Stadtman TC, Hatfield DL (1989) Identification of a selenocysteyl-tRNA(Ser) in mammalian cells that recognizes the nonsense codon, UGA. J Biol Chem 264:9724–9727
Berry MJ, Banu L, Harney JW, Larsen PR (1993) Functional characterization of the eukaryotic SECIS elements which direct selenocysteine insertion at UGA codons. EMBO J 12:3315–3322
Bermano G, Nicol F, Dyer JA, Sunde RA, Beckett GJ, Arthur JR, Hesketh JE (1995) Tissue-specific regulation of selenoenzyme gene expression during selenium deficiency in rats. Biochem J 311(Pt 2):425–430
Hadley KB, Sunde RA (2001) Selenium regulation of thioredoxin reductase activity and mRNA levels in rat liver. J Nutr Biochem 12:693–702
Seyedali A, Berry MJ (2014) Nonsense-mediated decay factors are involved in the regulation of selenoprotein mRNA levels during selenium deficiency. RNA 20:1248–1256. doi:10.1261/rna.043463.113
Mendell JT, Sharifi NA, Meyers JL, Martinez-Murillo F, Dietz HC (2004) Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat Genet 36:1073–1078. doi:10.1038/ng1429
Stockklausner C, Breit S, Neu-Yilik G, Echner N, Hentze MW, Kulozik AE, Gehring NH (2006) The uORF-containing thrombopoietin mRNA escapes nonsense-mediated decay (NMD). Nucleic Acids Res 34:2355–2363. doi:10.1093/nar/gkl277
Brett D, Hanke J, Lehmann G, Haase S, Delbruck S, Krueger S, Reich J, Bork P (2000) EST comparison indicates 38% of human mRNAs contain possible alternative splice forms. FEBS Lett 474:83–86
Lewis BP, Green RE, Brenner SE (2003) Evidence for the widespread coupling of alternative splicing and nonsense-mediated mRNA decay in humans. Proc Natl Acad Sci USA 100:189–192. doi:10.1073/pnas.0136770100
Sureau A, Gattoni R, Dooghe Y, Stevenin J, Soret J (2001) SC35 autoregulates its expression by promoting splicing events that destabilize its mRNAs. EMBO J 20:1785–1796. doi:10.1093/emboj/20.7.1785
Tani H, Torimura M, Akimitsu N (2013) The RNA degradation pathway regulates the function of GAS5 a non-coding RNA in mammalian cells. PLoS One 8:e55684. doi:10.1371/journal.pone.0055684
Chew GL, Pauli A, Rinn JL, Regev A, Schier AF, Valen E (2013) Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140:2828–2834. doi:10.1242/dev.098343
Culbertson MR, Underbrink KM, Fink GR (1980) Frameshift suppression Saccharomyces cerevisiae. II. Genetic properties of group II suppressors. Genetics 95:833–853
Cui Y, Hagan KW, Zhang S, Peltz SW (1995) Identification and characterization of genes that are required for the accelerated degradation of mRNAs containing a premature translational termination codon. Genes Dev 9:423–436
Leeds P, Peltz SW, Jacobson A, Culbertson MR (1991) The product of the yeast UPF1 gene is required for rapid turnover of mRNAs containing a premature translational termination codon. Genes Dev 5:2303–2314
Leeds P, Wood JM, Lee BS, Culbertson MR (1992) Gene products that promote mRNA turnover in Saccharomyces cerevisiae. Mol Cell Biol 12:2165–2177
Behm-Ansmant I, Kashima I, Rehwinkel J, Sauliere J, Wittkopp N, Izaurralde E (2007) mRNA quality control: an ancient machinery recognizes and degrades mRNAs with nonsense codons. FEBS Lett 581:2845–2853. doi:10.1016/j.febslet.2007.05.027
Culbertson MR, Leeds PF (2003) Looking at mRNA decay pathways through the window of molecular evolution. Curr Opin Genet Dev 13:207–214
Chen YH, Su LH, Sun CH (2008) Incomplete nonsense-mediated mRNA decay in Giardia lamblia. Int J Parasitol 38:1305–1317. doi:10.1016/j.ijpara.2008.02.006
Kadlec J, Guilligay D, Ravelli RB, Cusack S (2006) Crystal structure of the UPF2-interacting domain of nonsense-mediated mRNA decay factor UPF1. RNA 12:1817–1824. doi:10.1261/rna.177606
Cali BM, Kuchma SL, Latham J, Anderson P (1999) smg-7 is required for mRNA surveillance in Caenorhabditis elegans. Genetics 151:605–616
Hodgkin J, Papp A, Pulak R, Ambros V, Anderson P (1989) A new kind of informational suppression in the nematode Caenorhabditis elegans. Genetics 123:301–313
Pulak R, Anderson P (1993) mRNA surveillance by the Caenorhabditis elegans smg genes. Genes Dev 7:1885–1897
Applequist SE, Selg M, Raman C, Jack HM (1997) Cloning and characterization of HUPF1, a human homolog of the Saccharomyces cerevisiae nonsense mRNA-reducing UPF1 protein. Nucleic Acids Res 25:814–821
Aronoff R, Baran R, Hodgkin J (2001) Molecular identification of smg-4, required for mRNA surveillance in C. elegans. Gene 268:153–164
Denning G, Jamieson L, Maquat LE, Thompson EA, Fields AP (2001) Cloning of a novel phosphatidylinositol kinase-related kinase: characterization of the human SMG-1 RNA surveillance protein. J Biol Chem 276:22709–22714. doi:10.1074/jbc.C100144200
Lykke-Andersen J, Shu MD, Steitz JA (2000) Human Upf proteins target an mRNA for nonsense-mediated decay when bound downstream of a termination codon. Cell 103:1121–1131
Ohnishi T, Yamashita A, Kashima I, Schell T, Anders KR, Grimson A, Hachiya T, Hentze MW, Anderson P, Ohno S (2003) Phosphorylation of hUPF1 induces formation of mRNA surveillance complexes containing hSMG-5 and hSMG-7. Mol Cell 12:1187–1200
Page MF, Carr B, Anders KR, Grimson A, Anderson P (1999) SMG-2 is a phosphorylated protein required for mRNA surveillance in Caenorhabditis elegans and related to Upf1p of yeast. Mol Cell Biol 19:5943–5951
Yamashita A, Ohnishi T, Kashima I, Taya Y, Ohno S (2001) Human SMG-1, a novel phosphatidylinositol 3-kinase-related protein kinase, associates with components of the mRNA surveillance complex and is involved in the regulation of nonsense-mediated mRNA decay. Genes Dev 15:2215–2228. doi:10.1101/gad.913001
Casadio A, Longman D, Hug N, Delavaine L, Vallejos Baier R, Alonso CR, Caceres JF (2015) Identification and characterization of novel factors that act in the nonsense-mediated mRNA decay pathway in nematodes, flies and mammals. EMBO Rep 16:71–78. doi:10.15252/embr.201439183
Hug N, Caceres JF (2014) The RNA helicase DHX34 activates NMD by promoting a transition from the surveillance to the decay-inducing complex. Cell Rep 8:1845–1856. doi:10.1016/j.celrep.2014.08.020
Izumi N, Yamashita A, Ohno S (2012) Integrated regulation of PIKK-mediated stress responses by AAA+ proteins RUVBL1 and RUVBL2. Nucleus 3:29–43. doi:10.4161/nucl.18926
Longman D, Hug N, Keith M, Anastasaki C, Patton EE, Grimes G, Caceres JF (2013) DHX34 and NBAS form part of an autoregulatory NMD circuit that regulates endogenous RNA targets in human cells, zebrafish and Caenorhabditis elegans. Nucleic Acids Res 41:8319–8331. doi:10.1093/nar/gkt585
Longman D, Plasterk RH, Johnstone IL, Caceres JF (2007) Mechanistic insights and identification of two novel factors in the C. elegans NMD pathway. Genes Dev 21:1075–1085. doi:10.1101/gad.417707
Gregersen LH, Schueler M, Munschauer M, Mastrobuoni G, Chen W, Kempa S, Dieterich C, Landthaler M (2014) MOV10 Is a 5′ to 3′ RNA helicase contributing to UPF1 mRNA target degradation by translocation along 3′ UTRs. Mol Cell 54:573–585. doi:10.1016/j.molcel.2014.03.017
Yamashita A, Izumi N, Kashima I, Ohnishi T, Saari B, Katsuhata Y, Muramatsu R, Morita T, Iwamatsu A, Hachiya T, Kurata R, Hirano H, Anderson P, Ohno S (2009) SMG-8 and SMG-9, two novel subunits of the SMG-1 complex, regulate remodeling of the mRNA surveillance complex during nonsense-mediated mRNA decay. Genes Dev 23:1091–1105. doi:10.1101/gad.1767209
Riehs-Kearnan N, Gloggnitzer J, Dekrout B, Jonak C, Riha K (2012) Aberrant growth and lethality of Arabidopsis deficient in nonsense-mediated RNA decay factors is caused by autoimmune-like response. Nucleic Acids Res 40:5615–5624. doi:10.1093/nar/gks195
Medghalchi SM, Frischmeyer PA, Mendell JT, Kelly AG, Lawler AM, Dietz HC (2001) Rent1, a trans-effector of nonsense-mediated mRNA decay, is essential for mammalian embryonic viability. Hum Mol Genet 10:99–105
Wittkopp N, Huntzinger E, Weiler C, Sauliere J, Schmidt S, Sonawane M, Izaurralde E (2009) Nonsense-mediated mRNA decay effectors are essential for zebrafish embryonic development and survival. Mol Cell Biol 29:3517–3528. doi:10.1128/MCB.00177-09
Avery P, Vicente-Crespo M, Francis D, Nashchekina O, Alonso CR, Palacios IM (2011) Drosophila Upf1 and Upf2 loss of function inhibits cell growth and causes animal death in a Upf3-independent manner. RNA 17:624–638. doi:10.1261/rna.2404211
Bhattacharya A, Czaplinski K, Trifillis P, He F, Jacobson A, Peltz SW (2000) Characterization of the biochemical properties of the human Upf1 gene product that is involved in nonsense-mediated mRNA decay. RNA 6:1226–1235
Cheng Z, Muhlrad D, Lim MK, Parker R, Song H (2007) Structural and functional insights into the human Upf1 helicase core. EMBO J 26:253–264. doi:10.1038/sj.emboj.7601464
Fairman-Williams ME, Guenther UP, Jankowsky E (2010) SF1 and SF2 helicases: family matters. Curr Opin Struct Biol 20:313–324. doi:10.1016/j.sbi.2010.03.011
Singleton MR, Dillingham MS, Wigley DB (2007) Structure and mechanism of helicases and nucleic acid translocases. Annu Rev Biochem 76:23–50. doi:10.1146/annurev.biochem.76.052305.115300
Chamieh H, Ballut L, Bonneau F, Le Hir H (2008) NMD factors UPF2 and UPF3 bridge UPF1 to the exon junction complex and stimulate its RNA helicase activity. Nat Struct Mol Biol 15:85–93. doi:10.1038/nsmb1330
Chakrabarti S, Jayachandran U, Bonneau F, Fiorini F, Basquin C, Domcke S, Le Hir H, Conti E (2011) Molecular mechanisms for the RNA-dependent ATPase activity of Upf1 and its regulation by Upf2. Mol Cell 41:693–703. doi:10.1016/j.molcel.2011.02.010
Weng Y, Czaplinski K, Peltz SW (1996) Genetic and biochemical characterization of mutations in the ATPase and helicase regions of the Upf1 protein. Mol Cell Biol 16:5477–5490
Mendell JT, ap Rhys CM, Dietz HC (2002) Separable roles for rent1/hUpf1 in altered splicing and decay of nonsense transcripts. Science 298:419–422. doi:10.1126/science.1074428
Weng Y, Czaplinski K, Peltz SW (1996) Identification and characterization of mutations in the UPF1 gene that affect nonsense suppression and the formation of the Upf protein complex but not mRNA turnover. Mol Cell Biol 16:5491–5506
Franks TM, Singh G, Lykke-Andersen J (2010) Upf1 ATPase-dependent mRNP disassembly is required for completion of nonsense- mediated mRNA decay. Cell 143:938–950. doi:10.1016/j.cell.2010.11.043
Shigeoka T, Kato S, Kawaichi M, Ishida Y (2012) Evidence that the Upf1-related molecular motor scans the 3′-UTR to ensure mRNA integrity. Nucleic Acids Res 40:6887–6897. doi:10.1093/nar/gks344
Fiorini F, Boudvillain M, Le Hir H (2013) Tight intramolecular regulation of the human Upf1 helicase by its N- and C-terminal domains. Nucleic Acids Res 41:2404–2415. doi:10.1093/nar/gks1320
Ponting CP (2000) Novel eIF4G domain homologues linking mRNA translation with nonsense-mediated mRNA decay. Trends Biochem Sci 25:423–426
Clerici M, Deniaud A, Boehm V, Gehring NH, Schaffitzel C, Cusack S (2014) Structural and functional analysis of the three MIF4G domains of nonsense-mediated decay factor UPF2. Nucleic Acids Res 42:2673–2686. doi:10.1093/nar/gkt1197
Aravind L, Koonin EV (2000) Eukaryote-specific domains in translation initiation factors: implications for translation regulation and evolution of the translation system. Genome Res 10:1172–1184
Alexandrov A, Colognori D, Shu MD, Steitz JA (2012) Human spliceosomal protein CWC22 plays a role in coupling splicing to exon junction complex deposition and nonsense-mediated decay. Proc Natl Acad Sci USA 109:21313–21318. doi:10.1073/pnas.1219725110
Barbosa I, Haque N, Fiorini F, Barrandon C, Tomasetto C, Blanchette M, Le Hir H (2012) Human CWC22 escorts the helicase eIF4AIII to spliceosomes and promotes exon junction complex assembly. Nat Struct Mol Biol 19:983–990. doi:10.1038/nsmb.2380
Buchwald G, Schussler S, Basquin C, Le Hir H, Conti E (2013) Crystal structure of the human eIF4AIII-CWC22 complex shows how a DEAD-box protein is inhibited by a MIF4G domain. Proc Natl Acad Sci USA 110:E4611–E4618. doi:10.1073/pnas.1314684110
Steckelberg AL, Boehm V, Gromadzka AM, Gehring NH (2012) CWC22 connects pre-mRNA splicing and exon junction complex assembly. Cell Rep 2:454–461. doi:10.1016/j.celrep.2012.08.017
Kadlec J, Izaurralde E, Cusack S (2004) The structural basis for the interaction between nonsense-mediated mRNA decay factors UPF2 and UPF3. Nat Struct Mol Biol 11:330–337. doi:10.1038/nsmb741
Serin G, Gersappe A, Black JD, Aronoff R, Maquat LE (2001) Identification and characterization of human orthologues to Saccharomyces cerevisiae Upf2 protein and Upf3 protein (Caenorhabditis elegans SMG-4). Mol Cell Biol 21:209–223. doi:10.1128/MCB.21.1.209-223.2001
Melero R, Buchwald G, Castano R, Raabe M, Gil D, Lazaro M, Urlaub H, Conti E, Llorca O (2012) The cryo-EM structure of the UPF-EJC complex shows UPF1 poised toward the RNA 3′ end. Nat Struct Mol Biol 19(498–505):S491–S492. doi:10.1038/nsmb.2287
Fourati Z, Roy B, Millan C, Coureux PD, Kervestin S, van Tilbeurgh H, He F, Uson I, Jacobson A, Graille M (2014) A highly conserved region essential for NMD in the Upf2 N-terminal domain. J Mol Biol 426:3689–3702. doi:10.1016/j.jmb.2014.09.015
Chan WK, Bhalla AD, Le Hir H, Nguyen LS, Huang L, Gecz J, Wilkinson MF (2009) A UPF3-mediated regulatory switch that maintains RNA surveillance. Nat Struct Mol Biol 16:747–753. doi:10.1038/nsmb.1612
Kunz JB, Neu-Yilik G, Hentze MW, Kulozik AE, Gehring NH (2006) Functions of hUpf3a and hUpf3b in nonsense-mediated mRNA decay and translation. RNA 12:1015–1022. doi:10.1261/rna.12506
Turner JM (2007) Meiotic sex chromosome inactivation. Development 134:1823–1831. doi:10.1242/dev.000018
Buchwald G, Ebert J, Basquin C, Sauliere J, Jayachandran U, Bono F, Le Hir H, Conti E (2010) Insights into the recruitment of the NMD machinery from the crystal structure of a core EJC-UPF3b complex. Proc Natl Acad Sci USA 107:10050–10055. doi:10.1073/pnas.1000993107
Kashima I, Jonas S, Jayachandran U, Buchwald G, Conti E, Lupas AN, Izaurralde E (2010) SMG6 interacts with the exon junction complex via two conserved EJC-binding motifs (EBMs) required for nonsense-mediated mRNA decay. Genes Dev 24:2440–2450. doi:10.1101/gad.604610
Metze S, Herzog VA, Ruepp MD, Muhlemann O (2013) Comparison of EJC-enhanced and EJC-independent NMD in human cells reveals two partially redundant degradation pathways. RNA 19:1432–1448. doi:10.1261/rna.038893.113
Gatfield D, Unterholzner L, Ciccarelli FD, Bork P, Izaurralde E (2003) Nonsense-mediated mRNA decay in Drosophila: at the intersection of the yeast and mammalian pathways. EMBO J 22:3960–3970. doi:10.1093/emboj/cdg371
Spingola M, Grate L, Haussler D, Ares M Jr (1999) Genome-wide bioinformatic and molecular analysis of introns in Saccharomyces cerevisiae. RNA 5:221–234
Wen J, Brogna S (2010) Splicing-dependent NMD does not require the EJC in Schizosaccharomyces pombe. EMBO J 29:1537–1551. doi:10.1038/emboj.2010.48
Grimson A, O’Connor S, Newman CL, Anderson P (2004) SMG-1 is a phosphatidylinositol kinase-related protein kinase required for nonsense-mediated mRNA decay in Caenorhabditis elegans. Mol Cell Biol 24:7483–7490. doi:10.1128/MCB.24.17.7483-7490.2004
Melero R, Uchiyama A, Castano R, Kataoka N, Kurosawa H, Ohno S, Yamashita A, Llorca O (2014) Structures of SMG1-UPFs complexes: SMG1 contributes to regulate UPF2-dependent activation of UPF1 in NMD. Structure 22:1105–1119. doi:10.1016/j.str.2014.05.015
Arias-Palomo E, Yamashita A, Fernandez IS, Nunez-Ramirez R, Bamba Y, Izumi N, Ohno S, Llorca O (2011) The nonsense-mediated mRNA decay SMG-1 kinase is regulated by large-scale conformational changes controlled by SMG-8. Genes Dev 25:153–164. doi:10.1101/gad.606911
Fernandez IS, Yamashita A, Arias-Palomo E, Bamba Y, Bartolome RA, Canales MA, Teixido J, Ohno S, Llorca O (2011) Characterization of SMG-9, an essential component of the nonsense-mediated mRNA decay SMG1C complex. Nucleic Acids Res 39:347–358. doi:10.1093/nar/gkq749
Chakrabarti S, Bonneau F, Schussler S, Eppinger E, Conti E (2014) Phospho-dependent and phospho-independent interactions of the helicase UPF1 with the NMD factors SMG5-SMG7 and SMG6. Nucleic Acids Res 42:9447–9460. doi:10.1093/nar/gku578
Lasalde C, Rivera AV, Leon AJ, Gonzalez-Feliciano JA, Estrella LA, Rodriguez-Cruz EN, Correa ME, Cajigas IJ, Bracho DP, Vega IE, Wilkinson MF, Gonzalez CI (2014) Identification and functional analysis of novel phosphorylation sites in the RNA surveillance protein Upf1. Nucleic Acids Res 42:1916–1929. doi:10.1093/nar/gkt1049
Wang W, Cajigas IJ, Peltz SW, Wilkinson MF, Gonzalez CI (2006) Role for Upf2p phosphorylation in Saccharomyces cerevisiae nonsense-mediated mRNA decay. Mol Cell Biol 26:3390–3400. doi:10.1128/MCB.26.9.3390-3400.2006
Fukuhara N, Ebert J, Unterholzner L, Lindner D, Izaurralde E, Conti E (2005) SMG7 is a 14-3-3-like adaptor in the nonsense-mediated mRNA decay pathway. Mol Cell 17:537–547. doi:10.1016/j.molcel.2005.01.010
Gardino AK, Smerdon SJ, Yaffe MB (2006) Structural determinants of 14-3-3 binding specificities and regulation of subcellular localization of 14-3-3-ligand complexes: a comparison of the X-ray crystal structures of all human 14-3-3 isoforms. Semin Cancer Biol 16:173–182. doi:10.1016/j.semcancer.2006.03.007
Jonas S, Weichenrieder O, Izaurralde E (2013) An unusual arrangement of two 14-3-3-like domains in the SMG5-SMG7 heterodimer is required for efficient nonsense-mediated mRNA decay. Genes Dev 27:211–225. doi:10.1101/gad.206672.112
Obsil T, Obsilova V (2011) Structural basis of 14-3-3 protein functions. Semin Cell Dev Biol 22:663–672. doi:10.1016/j.semcdb.2011.09.001
Okada-Katsuhata Y, Yamashita A, Kutsuzawa K, Izumi N, Hirahara F, Ohno S (2012) N- and C-terminal Upf1 phosphorylations create binding platforms for SMG-6 and SMG-5:SMG-7 during NMD. Nucleic Acids Res 40:1251–1266. doi:10.1093/nar/gkr791
Unterholzner L, Izaurralde E (2004) SMG7 acts as a molecular link between mRNA surveillance and mRNA decay. Mol Cell 16:587–596. doi:10.1016/j.molcel.2004.10.013
Loh B, Jonas S, Izaurralde E (2013) The SMG5-SMG7 heterodimer directly recruits the CCR4-NOT deadenylase complex to mRNAs containing nonsense codons via interaction with POP2. Genes Dev 27:2125–2138. doi:10.1101/gad.226951.113
He F, Jacobson A (1995) Identification of a novel component of the nonsense-mediated mRNA decay pathway by use of an interacting protein screen. Genes Dev 9:437–454
Lykke-Andersen J (2002) Identification of a human decapping complex associated with hUpf proteins in nonsense-mediated decay. Mol Cell Biol 22:8114–8121
He F, Jacobson A (2001) Upf1p, Nmd2p, and Upf3p regulate the decapping and exonucleolytic degradation of both nonsense-containing mRNAs and wild-type mRNAs. Mol Cell Biol 21:1515–1530. doi:10.1128/MCB.21.5.1515-1530.2001
Lejeune F, Li X, Maquat LE (2003) Nonsense-mediated mRNA decay in mammalian cells involves decapping, deadenylating, and exonucleolytic activities. Mol Cell 12:675–687
Cho H, Kim KM, Kim YK (2009) Human proline-rich nuclear receptor coregulatory protein 2 mediates an interaction between mRNA surveillance machinery and decapping complex. Mol Cell 33:75–86. doi:10.1016/j.molcel.2008.11.022
Lai T, Cho H, Liu Z, Bowler MW, Piao S, Parker R, Kim YK, Song H (2012) Structural basis of the PNRC2-mediated link between mrna surveillance and decapping. Structure 20:2025–2037. doi:10.1016/j.str.2012.09.009
Cho H, Han S, Choe J, Park SG, Choi SS, Kim YK (2013) SMG5-PNRC2 is functionally dominant compared with SMG5-SMG7 in mammalian nonsense-mediated mRNA decay. Nucleic Acids Res 41:1319–1328. doi:10.1093/nar/gks1222
Anders KR, Grimson A, Anderson P (2003) SMG-5, required for C.elegans nonsense-mediated mRNA decay, associates with SMG-2 and protein phosphatase 2A. EMBO J 22:641–650. doi:10.1093/emboj/cdg056
Clissold PM, Ponting CP (2000) PIN domains in nonsense-mediated mRNA decay and RNAi. Curr Biol 10:R888–R890
Schoenberg DR (2011) Mechanisms of endonuclease-mediated mRNA decay. Wiley Interdiscip Rev RNA 2:582–600. doi:10.1002/wrna.78
Glavan F, Behm-Ansmant I, Izaurralde E, Conti E (2006) Structures of the PIN domains of SMG6 and SMG5 reveal a nuclease within the mRNA surveillance complex. EMBO J 25:5117–5125. doi:10.1038/sj.emboj.7601377
Chiu SY, Serin G, Ohara O, Maquat LE (2003) Characterization of human Smg5/7a: a protein with similarities to Caenorhabditis elegans SMG5 and SMG7 that functions in the dephosphorylation of Upf1. RNA 9:77–87
Gatfield D, Izaurralde E (2004) Nonsense-mediated messenger RNA decay is initiated by endonucleolytic cleavage in Drosophila. Nature 429:575–578. doi:10.1038/nature02559
Huntzinger E, Kashima I, Fauser M, Sauliere J, Izaurralde E (2008) SMG6 is the catalytic endonuclease that cleaves mRNAs containing nonsense codons in metazoan. RNA 14:2609–2617. doi:10.1261/rna.1386208
Eberle AB, Lykke-Andersen S, Muhlemann O, Jensen TH (2009) SMG6 promotes endonucleolytic cleavage of nonsense mRNA in human cells. Nat Struct Mol Biol 16:49–55. doi:10.1038/nsmb.1530
Nicholson P, Josi C, Kurosawa H, Yamashita A, Muhlemann O (2014) A novel phosphorylation-independent interaction between SMG6 and UPF1 is essential for human NMD. Nucleic Acids Res 42:9217–9235. doi:10.1093/nar/gku645
Kurosaki T, Li W, Hoque M, Popp MW, Ermolenko DN, Tian B, Maquat LE (2014) A post-translational regulatory switch on UPF1 controls targeted mRNA degradation. Genes Dev 28:1900–1916. doi:10.1101/gad.245506.114
Lykke-Andersen S, Chen Y, Ardal BR, Lilje B, Waage J, Sandelin A, Jensen TH (2014) Human nonsense-mediated RNA decay initiates widely by endonucleolysis and targets snoRNA host genes. Genes Dev 28:2498–2517. doi:10.1101/gad.246538.114
Schmidt SA, Foley PL, Jeong DH, Rymarquis LA, Doyle F, Tenenbaum SA, Belasco JG, Green PJ (2014) Identification of SMG6 cleavage sites and a preferred RNA cleavage motif by global analysis of endogenous NMD targets in human cells. Nucleic Acids Res. doi:10.1093/nar/gku1258
Hwang J, Maquat LE (2011) Nonsense-mediated mRNA decay (NMD) in animal embryogenesis: to die or not to die, that is the question. Curr Opin Genet Dev 21:422–430. doi:10.1016/j.gde.2011.03.008
Weischenfeldt J, Damgaard I, Bryder D, Theilgaard-Monch K, Thoren LA, Nielsen FC, Jacobsen SE, Nerlov C, Porse BT (2008) NMD is essential for hematopoietic stem and progenitor cells and for eliminating by-products of programmed DNA rearrangements. Genes Dev 22:1381–1396. doi:10.1101/gad.468808
McIlwain DR, Pan Q, Reilly PT, Elia AJ, McCracken S, Wakeham AC, Itie-Youten A, Blencowe BJ, Mak TW (2010) Smg1 is required for embryogenesis and regulates diverse genes via alternative splicing coupled to nonsense-mediated mRNA decay. Proc Natl Acad Sci USA 107:12186–12191. doi:10.1073/pnas.1007336107
Thoren LA, Norgaard GA, Weischenfeldt J, Waage J, Jakobsen JS, Damgaard I, Bergstrom FC, Blom AM, Borup R, Bisgaard HC, Porse BT (2010) UPF2 is a critical regulator of liver development, function and regeneration. PLoS One 5:e11650. doi:10.1371/journal.pone.0011650
Li T, Shi Y, Wang P, Guachalla LM, Sun B, Joerss T, Chen YS, Groth M, Krueger A, Platzer M, Yang YG, Rudolph KL, Wang ZQ (2015) Smg6/Est1 licenses embryonic stem cell differentiation via nonsense-mediated mRNA decay. EMBO J. doi:10.15252/embj.201489947
Isken O, Maquat LE (2008) The multiple lives of NMD factors: balancing roles in gene and genome regulation. Nat Rev Genet 9:699–712. doi:10.1038/nrg2402
Nicholson P, Yepiskoposyan H, Metze S, Zamudio Orozco R, Kleinschmidt N, Muhlemann O (2010) Nonsense-mediated mRNA decay in human cells: mechanistic insights, functions beyond quality control and the double-life of NMD factors. Cell Mol Life Sci 67:677–700. doi:10.1007/s00018-009-0177-1
Tarpey PS, Raymond FL, Nguyen LS, Rodriguez J, Hackett A, Vandeleur L, Smith R, Shoubridge C, Edkins S, Stevens C, O’Meara S, Tofts C, Barthorpe S, Buck G, Cole J, Halliday K, Hills K, Jones D, Mironenko T, Perry J, Varian J, West S, Widaa S, Teague J, Dicks E, Butler A, Menzies A, Richardson D, Jenkinson A, Shepherd R, Raine K, Moon J, Luo Y, Parnau J, Bhat SS, Gardner A, Corbett M, Brooks D, Thomas P, Parkinson-Lawrence E, Porteous ME, Warner JP, Sanderson T, Pearson P, Simensen RJ, Skinner C, Hoganson G, Superneau D, Wooster R, Bobrow M, Turner G, Stevenson RE, Schwartz CE, Futreal PA, Srivastava AK, Stratton MR, Gecz J (2007) Mutations in UPF3B, a member of the nonsense-mediated mRNA decay complex, cause syndromic and nonsyndromic mental retardation. Nat Genet 39:1127–1133. doi:10.1038/ng2100
Laumonnier F, Shoubridge C, Antar C, Nguyen LS, Van Esch H, Kleefstra T, Briault S, Fryns JP, Hamel B, Chelly J, Ropers HH, Ronce N, Blesson S, Moraine C, Gecz J, Raynaud M (2010) Mutations of the UPF3B gene, which encodes a protein widely expressed in neurons, are associated with nonspecific mental retardation with or without autism. Mol Psychiatry 15:767–776. doi:10.1038/mp.2009.14
Addington AM, Gauthier J, Piton A, Hamdan FF, Raymond A, Gogtay N, Miller R, Tossell J, Bakalar J, Inoff-Germain G, Gochman P, Long R, Rapoport JL, Rouleau GA (2011) A novel frameshift mutation in UPF3B identified in brothers affected with childhood onset schizophrenia and autism spectrum disorders. Mol Psychiatry 16:238–239. doi:10.1038/mp.2010.59
Lynch SA, Nguyen LS, Ng LY, Waldron M, McDonald D, Gecz J (2012) Broadening the phenotype associated with mutations in UPF3B: two further cases with renal dysplasia and variable developmental delay. Eur J Med Genet 55:476–479. doi:10.1016/j.ejmg.2012.03.010
Xu X, Zhang L, Tong P, Xun G, Su W, Xiong Z, Zhu T, Zheng Y, Luo S, Pan Y, Xia K, Hu Z (2013) Exome sequencing identifies UPF3B as the causative gene for a Chinese non-syndrome mental retardation pedigree. Clin Genet 83:560–564. doi:10.1111/cge.12014
Nguyen LS, Jolly L, Shoubridge C, Chan WK, Huang L, Laumonnier F, Raynaud M, Hackett A, Field M, Rodriguez J, Srivastava AK, Lee Y, Long R, Addington AM, Rapoport JL, Suren S, Hahn CN, Gamble J, Wilkinson MF, Corbett MA, Gecz J (2012) Transcriptome profiling of UPF3B/NMD-deficient lymphoblastoid cells from patients with various forms of intellectual disability. Mol Psychiatry 17:1103–1115. doi:10.1038/mp.2011.163
Chan WK, Huang L, Gudikote JP, Chang YF, Imam JS, MacLean JA 2nd, Wilkinson MF (2007) An alternative branch of the nonsense-mediated decay pathway. EMBO J 26:1820–1830. doi:10.1038/sj.emboj.7601628
Nguyen LS, Kim HG, Rosenfeld JA, Shen Y, Gusella JF, Lacassie Y, Layman LC, Shaffer LG, Gecz J (2013) Contribution of copy number variants involving nonsense-mediated mRNA decay pathway genes to neuro-developmental disorders. Hum Mol Genet 22:1816–1825. doi:10.1093/hmg/ddt035
Gulsuner S, Walsh T, Watts AC, Lee MK, Thornton AM, Casadei S, Rippey C, Shahin H, Consortium on the Genetics of S, Group PS, Nimgaonkar VL, Go RC, Savage RM, Swerdlow NR, Gur RE, Braff DL, King MC, McClellan JM (2013) Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network. Cell 154:518–529. doi:10.1016/j.cell.2013.06.049
Liu C, Karam R, Zhou Y, Su F, Ji Y, Li G, Xu G, Lu L, Wang C, Song M, Zhu J, Wang Y, Zhao Y, Foo WC, Zuo M, Valasek MA, Javle M, Wilkinson MF, Lu Y (2014) The UPF1 RNA surveillance gene is commonly mutated in pancreatic adenosquamous carcinoma. Nat Med 20:596–598. doi:10.1038/nm.3548
Albers CA, Paul DS, Schulze H, Freson K, Stephens JC, Smethurst PA, Jolley JD, Cvejic A, Kostadima M, Bertone P, Breuning MH, Debili N, Deloukas P, Favier R, Fiedler J, Hobbs CM, Huang N, Hurles ME, Kiddle G, Krapels I, Nurden P, Ruivenkamp CA, Sambrook JG, Smith K, Stemple DL, Strauss G, Thys C, van Geet C, Newbury-Ecob R, Ouwehand WH, Ghevaert C (2012) Compound inheritance of a low-frequency regulatory SNP and a rare null mutation in exon-junction complex subunit RBM8A causes TAR syndrome. Nat Genet 44(435–439):S431–S432. doi:10.1038/ng.1083
Greenhalgh KL, Howell RT, Bottani A, Ancliff PJ, Brunner HG, Verschuuren-Bemelmans CC, Vernon E, Brown KW, Newbury-Ecob RA (2002) Thrombocytopenia-absent radius syndrome: a clinical genetic study. J Med Genet 39:876–881
Richieri-Costa A, Pereira SC (1992) Short stature, Robin sequence, cleft mandible, pre/postaxial hand anomalies, and clubfoot: a new autosomal recessive syndrome. Am J Med Genet 42:681–687. doi:10.1002/ajmg.1320420511
Favaro FP, Alvizi L, Zechi-Ceide RM, Bertola D, Felix TM, de Souza J, Raskin S, Twigg SR, Weiner AM, Armas P, Margarit E, Calcaterra NB, Andersen GR, McGowan SJ, Wilkie AO, Richieri-Costa A, de Almeida ML, Passos-Bueno MR (2014) A noncoding expansion in EIF4A3 causes Richieri-Costa-Pereira syndrome, a craniofacial disorder associated with limb defects. Am J Hum Genet 94:120–128. doi:10.1016/j.ajhg.2013.11.020
Culbertson MR (1999) RNA surveillance. Unforeseen consequences for gene expression, inherited genetic disorders and cancer. Trends Genet 15:74–80
Peltz SW, Morsy M, Welch EM, Jacobson A (2013) Ataluren as an agent for therapeutic nonsense suppression. Annu Rev Med 64:407–425. doi:10.1146/annurev-med-120611-144851
Mort M, Ivanov D, Cooper DN, Chuzhanova NA (2008) A meta-analysis of nonsense mutations causing human genetic disease. Hum Mutat 29:1037–1047. doi:10.1002/humu.20763
Bhuvanagiri M, Schlitter AM, Hentze MW, Kulozik AE (2010) NMD: RNA biology meets human genetic medicine. Biochem J 430:365–377. doi:10.1042/BJ20100699
Hall GW, Thein S (1994) Nonsense codon mutations in the terminal exon of the beta-globin gene are not associated with a reduction in beta-mRNA accumulation: a mechanism for the phenotype of dominant beta-thalassemia. Blood 83:2031–2037
Holbrook JA, Neu-Yilik G, Hentze MW, Kulozik AE (2004) Nonsense-mediated decay approaches the clinic. Nat Genet 36:801–808. doi:10.1038/ng1403
Kerr TP, Sewry CA, Robb SA, Roberts RG (2001) Long mutant dystrophins and variable phenotypes: evasion of nonsense-mediated decay? Hum Genet 109:402–407. doi:10.1007/s004390100598
Kerem E (2004) Pharmacologic therapy for stop mutations: how much CFTR activity is enough? Curr Opin Pulm Med 10:547–552
Hermann T (2007) Aminoglycoside antibiotics: old drugs and new therapeutic approaches. Cell Mol Life Sci 64:1841–1852. doi:10.1007/s00018-007-7034-x
Bedwell DM, Kaenjak A, Benos DJ, Bebok Z, Bubien JK, Hong J, Tousson A, Clancy JP, Sorscher EJ (1997) Suppression of a CFTR premature stop mutation in a bronchial epithelial cell line. Nat Med 3:1280–1284
Kovesi TA, Swartz R, MacDonald N (1998) Transient renal failure due to simultaneous ibuprofen and aminoglycoside therapy in children with cystic fibrosis. N Engl J Med 338:65–66. doi:10.1056/NEJM199801013380115
Welch EM, Barton ER, Zhuo J, Tomizawa Y, Friesen WJ, Trifillis P, Paushkin S, Patel M, Trotta CR, Hwang S, Wilde RG, Karp G, Takasugi J, Chen G, Jones S, Ren H, Moon YC, Corson D, Turpoff AA, Campbell JA, Conn MM, Khan A, Almstead NG, Hedrick J, Mollin A, Risher N, Weetall M, Yeh S, Branstrom AA, Colacino JM, Babiak J, Ju WD, Hirawat S, Northcutt VJ, Miller LL, Spatrick P, He F, Kawana M, Feng H, Jacobson A, Peltz SW, Sweeney HL (2007) PTC124 targets genetic disorders caused by nonsense mutations. Nature 447:87–91. doi:10.1038/nature05756
Auld DS, Thorne N, Maguire WF, Inglese J (2009) Mechanism of PTC124 activity in cell-based luciferase assays of nonsense codon suppression. Proc Natl Acad Sci USA 106:3585–3590. doi:10.1073/pnas.0813345106
McElroy SP, Nomura T, Torrie LS, Warbrick E, Gartner U, Wood G, McLean WH (2013) A lack of premature termination codon read-through efficacy of PTC124 (Ataluren) in a diverse array of reporter assays. PLoS Biol 11:e1001593. doi:10.1371/journal.pbio.1001593
Chang JC, Kan YW (1979) Beta 0 thalassemia, a nonsense mutation in man. Proc Natl Acad Sci USA 76:2886–2889
Moriarty PM, Reddy CC, Maquat LE (1998) Selenium deficiency reduces the abundance of mRNA for Se-dependent glutathione peroxidase 1 by a UGA-dependent mechanism likely to be nonsense codon-mediated decay of cytoplasmic mRNA. Mol Cell Biol 18:2932–2939
Schmidt SA, Foley PL, Jeong DH, Rymarquis LA, Doyle F, Tenenbaum SA, Belasco JG, Green PJ (2015) Identification of SMG6 cleavage sites and a preferred RNA cleavage motif by global analysis of endogenous NMD targets in human cells. Nucleic Acids Res 43:309–323. doi:10.1093/nar/gku1258
Jones RB, Wang F, Luo Y, Yu C, Jin C, Suzuki T, Kan M, McKeehan WL (2001) The nonsense-mediated decay pathway and mutually exclusive expression of alternatively spliced FGFR2IIIb and -IIIc mRNAs. J Biol Chem 276:4158–4167. doi:10.1074/jbc.M006151200
Sharma K, D’Souza RC, Tyanova S, Schaab C, Wisniewski JR, Cox J, Mann M (2014) Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep 8:1583–1594. doi:10.1016/j.celrep.2014.07.036
Acknowledgments
This research was funded by grants from the Fritz Thyssen Stiftung and the Deutsche Forschungsgemeinschaft (SFB635, project B06) to N.H.G.
Author information
Authors and Affiliations
Corresponding author
Additional information
T. Fatscher and V. Boehm contributed equally to this work.
Rights and permissions
About this article
Cite this article
Fatscher, T., Boehm, V. & Gehring, N.H. Mechanism, factors, and physiological role of nonsense-mediated mRNA decay. Cell. Mol. Life Sci. 72, 4523–4544 (2015). https://doi.org/10.1007/s00018-015-2017-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-015-2017-9