Abstract
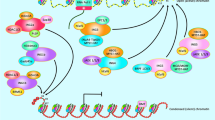
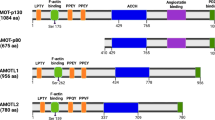
The INhibitor of Growth (ING) proteins belong to a well-conserved family which presents in diverse organisms with several structural and functional domains for each protein. The ING family members are found in association with many cellular processes. Thus, the ING family proteins are involved in regulation of gene transcription, DNA repair, tumorigenesis, apoptosis, cellular senescence and cell cycle arrest. The ING proteins have multiple domains that are potentially capable of binding to many partners. It is conceivable, therefore, that such proteins could function similarly within protein complexes. In this case, within this family, each function could be attributed to a specific domain. However, the role of ING domains is not definitively clear. In this review, we summarize recent advances in structure–function relationships in ING proteins. For each domain, we describe the known biological functions and the approaches utilized to identify the functions associated with ING proteins.

Similar content being viewed by others
References
Garkavtsev I, Kazarov A, Gudkov A, Riabowol K (1996) Suppression of the novel growth inhibitor p33ING1 promotes neoplastic transformation. Nat Genet 14:415–420
Garkavtsev I, Demetrick D, Riabowol K (1997) Cellular localization and chromosome mapping of a novel candidate tumor suppressor gene (ING1). Cytogenet Cell Genet 76:176–178
Zeremski M, Horrigan SK, Grigorian IA, Westbrook CA, Gudkov AV (1997) Localization of the candidate tumor suppressor gene ING1 to human chromosome 13q34. Somat Cell Mol Genet 23:233–236
Garkavtsev II (1999) Suppression of the novel growth inhibitor p33ING1 promotes neoplastic transformation. Nat Genet 23:373
Jager D, Stockert E, Scanlan MJ, Gure AO, Jager E, Knuth A, Old LJ, Chen YT (1999) Cancer-testis antigens and ING1 tumor suppressor gene product are breast cancer antigens: characterization of tissue-specific ING1 transcripts and a homologue gene. Cancer Res 59:6197–6204
Gunduz M, Ouchida M, Fukushima K, Hanafusa H, Etani T, Nishioka S, Nishizaki K, Shimizu K (2000) Genomic structure of the human ING1 gene and tumor-specific mutations detected in head and neck squamous cell carcinomas. Cancer Res 60:3143–3146
Saito A, Furukawa T, Fukushige S, Koyama S, Hoshi M, Hayashi Y, Horii A (2000) p24/ING1-ALT1 and p47/ING1-ALT2, distinct alternative transcripts of p33/ING1. J Hum Genet 45:177–181
Cheung KJ Jr, Li G (2001) The tumor suppressor ING1: structure and function. Exp Cell Res 268:1–6
Wagner MJ, Helbing CC (2005) Multiple variants of the ING1 and ING2 tumor suppressors are differentially expressed and thyroid hormone-responsive in Xenopus laevis. Gen Comp Endocrinol 144:38–50
Unoki M, Shen JC, Zheng ZM, Harris CC (2006) Novel splice variants of ING4 and their possible roles in the regulation of cell growth and motility. J Biol Chem 281:34677–34686
Shimada Y, Saito A, Suzuki M, Takahashi E, Horie M (1998) Cloning of a novel gene (ING1L) homologous to ING1, a candidate tumor suppressor. Cytogenet Cell Genet 83:232–235
Nagashima M, Shiseki M, Pedeux RM, Okamura S, Kitahama-Shiseki M, Miura K, Yokota J, Harris CC (2003) A novel PHD-finger motif protein, p47ING3, modulates p53-mediated transcription, cell cycle control, and apoptosis. Oncogene 22:343–350
Shiseki M, Nagashima M, Pedeux RM, Kitahama-Shiseki M, Miura K, Okamura S, Onogi H, Higashimoto Y, Appella E, Yokota J, Harris CC (2003) p29ING4 and p28ING5 bind to p53 and p300, and enhance p53 activity. Cancer Res 63:2373–2378
Campos EI, Chin MY, Kuo WH, Li G (2004) Biological functions of the ING family tumor suppressors. Cell Mol Life Sci 61:2597–2613
Soliman MA, Riabowol K (2007) After a decade of study-ING, a PHD for a versatile family of proteins. Trends Biochem Sci 32:509–519
Toyama T, Iwase H, Watson P, Muzik H, Saettler E, Magliocco A, DiFrancesco L, Forsyth P, Garkavtsev I, Kobayashi S, Riabowol K (1999) Suppression of ING1 expression in sporadic breast cancer. Oncogene 18:5187–5193
Ohmori M, Nagai M, Tasaka T, Koeffler HP, Toyama T, Riabowol K, Takahara J (1999) Decreased expression of p33ING1 mRNA in lymphoid malignancies. Am J Hematol 62:118–119
Oki E, Maehara Y, Tokunaga E, Kakeji Y, Sugimachi K (1999) Reduced expression of p33(ING1) and the relationship with p53 expression in human gastric cancer. Cancer Lett 147:157–162
Tokunaga E, Maehara Y, Oki E, Kitamura K, Kakeji Y, Ohno S, Sugimachi K (2000) Diminished expression of ING1 mRNA and the correlation with p53 expression in breast cancers. Cancer Lett 152:15–22
Chen L, Matsubara N, Yoshino T, Nagasaka T, Hoshizima N, Shirakawa Y, Naomoto Y, Isozaki H, Riabowol K, Tanaka N (2001) Genetic alterations of candidate tumor suppressor ING1 in human esophageal squamous cell cancer. Cancer Res 61:4345–4349
Krishnamurthy J, Kannan K, Feng J, Mohanprasad BK, Tsuchida N, Shanmugam G (2001) Mutational analysis of the candidate tumor suppressor gene ING1 in Indian oral squamous cell carcinoma. Oral Oncol 37:222–224
Bromidge T, Lynas C (2002) Relative levels of alternative transcripts of the ING1 gene and lack of mutations of p33/ING1 in haematological malignancies. Leuk Res 26:631–635
Gunduz M, Ouchida M, Fukushima K, Ito S, Jitsumori Y, Nakashima T, Nagai N, Nishizaki K, Shimizu K (2002) Allelic loss and reduced expression of the ING3, a candidate tumor suppressor gene at 7q31, in human head and neck cancers. Oncogene 21:4462–4470
Ito K, Kinjo K, Nakazato T, Ikeda Y, Kizaki M (2002) Expression and sequence analyses of p33(ING1) gene in myeloid leukemia. Am J Hematol 69:141–143
Nouman GS, Anderson JJ, Wood KM, Lunec J, Hall AG, Reid MM, Angus B (2002) Loss of nuclear expression of the p33(ING1b) inhibitor of growth protein in childhood acute lymphoblastic leukaemia. J Clin Pathol 55:596–601
Chen B, Campos EI, Crawford R, Martinka M, Li G (2003) Analyses of the tumour suppressor ING1 expression and gene mutation in human basal cell carcinoma. Int J Oncol 22:927–931
Lu F, Dai DL, Martinka M, Ho V, Li G (2006) Nuclear ING2 expression is reduced in human cutaneous melanomas. Br J Cancer 95:80–86
Wang Y, Dai DL, Martinka M, Li G (2007) Prognostic significance of nuclear ING3 expression in human cutaneous melanoma. Clin Cancer Res 13:4111–4116
Li J, Martinka M, Li G (2008) Role of ING4 in human melanoma cell migration, invasion and patient survival. Carcinogenesis 29:1373–1379
Cheung KJ Jr, Li G (2002) p33(ING1) enhances UVB-induced apoptosis in melanoma cells. Exp Cell Res 279:291–298
Chin MY, Ng KC, Li G (2005) The novel tumor suppressor p33ING2 enhances UVB-induced apoptosis in human melanoma cells. Exp Cell Res 304:531–543
Pedeux R, Sengupta S, Shen JC, Demidov ON, Saito S, Onogi H, Kumamoto K, Wincovitch S, Garfield SH, McMenamin M, Nagashima M, Grossman SR, Appella E, Harris CC (2005) ING2 regulates the onset of replicative senescence by induction of p300-dependent p53 acetylation. Mol Cell Biol 25:6639–6648
Gozani O, Karuman P, Jones DR, Ivanov D, Cha J, Lugovskoy AA, Baird CL, Zhu H, Field SJ, Lessnick SL, Villasenor J, Mehrotra B, Chen J, Rao VR, Brugge JS, Ferguson CG, Payrastre B, Myszka DG, Cantley LC, Wagner G, Divecha N, Prestwich GD, Yuan J (2003) The PHD finger of the chromatin-associated protein ING2 functions as a nuclear phosphoinositide receptor. Cell 114:99–111
Kim S, Chin K, Gray JW, Bishop JM (2004) A screen for genes that suppress loss of contact inhibition: identification of ING4 as a candidate tumor suppressor gene in human cancer. Proc Natl Acad Sci U S A 101:16251–16256
Shen JC, Unoki M, Ythier D, Duperray A, Varticovski L, Kumamoto K, Pedeux R, Harris CC (2007) Inhibitor of growth 4 suppresses cell spreading and cell migration by interacting with a novel binding partner, liprin alpha1. Cancer Res 67:2552–2558
Helbing CC, Veillette C, Riabowol K, Johnston RN, Garkavtsev I (1997) A novel candidate tumor suppressor, ING1, is involved in the regulation of apoptosis. Cancer Res 57:1255–1258
Scott M, Boisvert FM, Vieyra D, Johnston RN, Bazett-Jones DP, Riabowol K (2001) UV induces nucleolar translocation of ING1 through two distinct nucleolar targeting sequences. Nucleic Acids Res 29:2052–2058
Wang J, Chin MY, Li G (2006) The novel tumor suppressor p33ING2 enhances nucleotide excision repair via inducement of histone H4 acetylation and chromatin relaxation. Cancer Res 66:1906–1911
Kuo WH, Wang Y, Wong RP, Campos EI, Li G (2007) The ING1b tumor suppressor facilitates nucleotide excision repair by promoting chromatin accessibility to XPA. Exp Cell Res 313:1628–1638
Cheung KJ Jr, Mitchell D, Lin P, Li G (2001) The tumor suppressor candidate p33(ING1) mediates repair of UV-damaged DNA. Cancer Res 61:4974–4977
Garkavtsev I, Riabowol K (1997) Extension of the replicative life span of human diploid fibroblasts by inhibition of the p33ING1 candidate tumor suppressor. Mol Cell Biol 17:2014–2019
Nagashima M, Shiseki M, Miura K, Hagiwara K, Linke SP, Pedeux R, Wang XW, Yokota J, Riabowol K, Harris CC (2001) DNA damage-inducible gene p33ING2 negatively regulates cell proliferation through acetylation of p53. Proc Natl Acad Sci U S A 98:9671–9676
Loewith R, Meijer M, Lees-Miller SP, Riabowol K, Young D (2000) Three yeast proteins related to the human candidate tumor suppressor p33(ING1) are associated with histone acetyltransferase activities. Mol Cell Biol 20:3807–3816
Kuzmichev A, Zhang Y, Erdjument-Bromage H, Tempst P, Reinberg D (2002) Role of the Sin3-histone deacetylase complex in growth regulation by the candidate tumor suppressor p33(ING1). Mol Cell Biol 22:835–848
Wagner MJ, Gogela-Spehar M, Skirrow RC, Johnston RN, Riabowol K, Helbing CC (2001) Expression of novel ING variants is regulated by thyroid hormone in the Xenopus laevis tadpole. J Biol Chem 276:47013–47020
Toyama T, Iwase H, Yamashita H, Hara Y, Sugiura H, Zhang Z, Fukai I, Miura Y, Riabowol K, Fujii Y (2003) p33(ING1b) stimulates the transcriptional activity of the estrogen receptor alpha via its activation function (AF) 2 domain. J Steroid Biochem Mol Biol 87:57–63
Garkavtsev I, Kozin SV, Chernova O, Xu L, Winkler F, Brown E, Barnett GH, Jain RK (2004) The candidate tumour suppressor protein ING4 regulates brain tumour growth and angiogenesis. Nature 428:328–332
Ozer A, Wu LC, Bruick RK (2005) The candidate tumor suppressor ING4 represses activation of the hypoxia inducible factor (HIF). Proc Natl Acad Sci U S A 102:7481–7486
Piche B, Li G (2010) Inhibitor of growth tumor suppressors in cancer progression. Cell Mol Life Sci 67:1987–1999
Campos EI, Martinka M, Mitchell DL, Dai DL, Li G (2004) Mutations of the ING1 tumor suppressor gene detected in human melanoma abrogate nucleotide excision repair. Int J Oncol 25:73–80
Vieyra D, Senger DL, Toyama T, Muzik H, Brasher PM, Johnston RN, Riabowol K, Forsyth PA (2003) Altered subcellular localization and low frequency of mutations of ING1 in human brain tumors. Clin Cancer Res 9:5952–5961
Gunduz M, Nagatsuka H, Demircan K, Gunduz E, Cengiz B, Ouchida M, Tsujigiwa H, Yamachika E, Fukushima K, Beder L, Hirohata S, Ninomiya Y, Nishizaki K, Shimizu K, Nagai N (2005) Frequent deletion and down-regulation of ING4, a candidate tumor suppressor gene at 12p13, in head and neck squamous cell carcinomas. Gene 356:109–117
Cengiz B, Gunduz E, Gunduz M, Beder LB, Tamamura R, Bagci C, Yamanaka N, Shimizu K, Nagatsuka H (2010) Tumor-specific mutation and down-regulation of ING5 detected in oral squamous cell carcinoma. Int J Cancer. doi: 10.1002/ijc.25224
Coles AH, Liang H, Zhu Z, Marfella CG, Kang J, Imbalzano AN, Jones SN (2007) Deletion of p37Ing1 in mice reveals a p53-independent role for Ing1 in the suppression of cell proliferation, apoptosis, and tumorigenesis. Cancer Res 67:2054–2061
Doyon Y, Cayrou C, Ullah M, Landry AJ, Cote V, Selleck W, Lane WS, Tan S, Yang XJ, Cote J (2006) ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol Cell 21:51–64
Feng X, Hara Y, Riabowol K (2002) Different HATS of the ING1 gene family. Trends Cell Biol 12:532–538
Berardi P, Russell M, El-Osta A, Riabowol K (2004) Functional links between transcription, DNA repair and apoptosis. Cell Mol Life Sci 61:2173–2180
Ransom M, Dennehey BK, Tyler JK (2010) Chaperoning histones during DNA replication and repair. Cell 140:183–195
Cunliffe VT (2003) Memory by modification: the influence of chromatin structure on gene expression during vertebrate development. Gene 305:141–150
Jelinsky SA, Estep P, Church GM, Samson LD (2000) Regulatory networks revealed by transcriptional profiling of damaged Saccharomyces cerevisiae cells: Rpn4 links base excision repair with proteasomes. Mol Cell Biol 20:8157–8167
Amundson SA, Bittner M, Meltzer P, Trent J, Fornace AJ Jr (2001) Induction of gene expression as a monitor of exposure to ionizing radiation. Radiat Res 156:657–661
Sesto A, Navarro M, Burslem F, Jorcano JL (2002) Analysis of the ultraviolet B response in primary human keratinocytes using oligonucleotide microarrays. Proc Natl Acad Sci U S A 99:2965–2970
Wood C, Snijders A, Williamson J, Reynolds C, Baldwin J, Dickman M (2009) Post-translational modifications of the linker histone variants and their association with cell mechanisms. FEBS J 276:3685–3697
Garcia-Dominguez M, Reyes JC (2009) SUMO association with repressor complexes, emerging routes for transcriptional control. Biochim Biophys Acta 1789:451–459
Campos EI, Reinberg D (2009) Histones: annotating chromatin. Annu Rev Genet 43:559–599
Wu RS, Panusz HT, Hatch CL, Bonner WM (1986) Histones and their modifications. CRC Crit Rev Biochem 20:201–263
Scott M, Bonnefin P, Vieyra D, Boisvert FM, Young D, Bazett-Jones DP, Riabowol K (2001) UV-induced binding of ING1 to PCNA regulates the induction of apoptosis. J Cell Sci 114:3455–3462
Li J, Wang Y, Wong RP, Li G (2009) The role of ING tumor suppressors in UV stress response and melanoma progression. Curr Drug Targets 10:455–464
Larrieu D, Ythier D, Binet R, Brambilla C, Brambilla E, Sengupta S, Pedeux R (2009) ING2 controls the progression of DNA replication forks to maintain genome stability. EMBO Rep 10:1168–1174
He GH, Helbing CC, Wagner MJ, Sensen CW, Riabowol K (2005) Phylogenetic analysis of the ING family of PHD finger proteins. Mol Biol Evol 22:104–116
Ythier D, Larrieu D, Brambilla C, Brambilla E, Pedeux R (2008) The new tumor suppressor genes ING: genomic structure and status in cancer. Int J Cancer 123:1483–1490
Loewith R, Smith JS, Meijer M, Williams TJ, Bachman N, Boeke JD, Young D (2001) Pho23 is associated with the Rpd3 histone deacetylase and is required for its normal function in regulation of gene expression and silencing in Saccharomyces cerevisiae. J Biol Chem 276:24068–24074
Wang Y, Wang J, Li G (2006) Leucine zipper-like domain is required for tumor suppressor ING2-mediated nucleotide excision repair and apoptosis. FEBS Lett 580:3787–3793
Han X, Feng X, Rattner JB, Smith H, Bose P, Suzuki K, Soliman MA, Scott MS, Burke BE, Riabowol K (2008) Tethering by lamin A stabilizes and targets the ING1 tumour suppressor. Nat Cell Biol 10:1333–1340
Ha S, Park S, Yun CH, Choi Y (2002) Characterization of nuclear localization signal in mouse ING1 homolog protein. Biochem Biophys Res Commun 293:163–166
Gong W, Suzuki K, Russell M, Riabowol K (2005) Function of the ING family of PHD proteins in cancer. Int J Biochem Cell Biol 37:1054–1065
Zhang X, Wang KS, Wang ZQ, Xu LS, Wang QW, Chen F, Wei DZ, Han ZG (2005) Nuclear localization signal of ING4 plays a key role in its binding to p53. Biochem Biophys Res Commun 331:1032–1038
Russell MW, Soliman MA, Schriemer D, Riabowol K (2008) ING1 protein targeting to the nucleus by karyopherins is necessary for activation of p21. Biochem Biophys Res Commun 374:490–495
Sutherland JJ, Higgs RE, Watson I, Vieth M (2008) Chemical fragments as foundations for understanding target space and activity prediction. J Med Chem 51:2689–2700
Matthews JM, Bhati M, Lehtomaki E, Mansfield RE, Cubeddu L, Mackay JP (2009) It takes two to tango: the structure and function of LIM, RING, PHD and MYND domains. Curr Pharm Des 15:3681–3696
Aasland R, Gibson TJ, Stewart AF (1995) The PHD finger: implications for chromatin-mediated transcriptional regulation. Trends Biochem Sci 20:56–59
Pascual J, Martinez-Yamout M, Dyson HJ, Wright PE (2000) Structure of the PHD zinc finger from human Williams-Beuren syndrome transcription factor. J Mol Biol 304:723–729
Bienz M (2006) The PHD finger, a nuclear protein-interaction domain. Trends Biochem Sci 31:35–40
Capili AD, Schultz DC, Rauscher IF, Borden KL (2001) Solution structure of the PHD domain from the KAP-1 corepressor: structural determinants for PHD, RING and LIM zinc-binding domains. EMBO J 20:165–177
Kwan AH, Gell DA, Verger A, Crossley M, Matthews JM, Mackay JP (2003) Engineering a protein scaffold from a PHD finger. Structure 11:803–813
Bottomley MJ, Stier G, Pennacchini D, Legube G, Simon B, Akhtar A, Sattler M, Musco G (2005) NMR structure of the first PHD finger of autoimmune regulator protein (AIRE1). Insights into autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED) disease. J Biol Chem 280:11505–11512
Gibbons RJ, Bachoo S, Picketts DJ, Aftimos S, Asenbauer B, Bergoffen J, Berry SA, Dahl N, Fryer A, Keppler K, Kurosawa K, Levin ML, Masuno M, Neri G, Pierpont ME, Slaney SF, Higgs DR (1997) Mutations in transcriptional regulator ATRX establish the functional significance of a PHD-like domain. Nat Genet 17:146–148
Saugier-Veber P, Drouot N, Wolf LM, Kuhn JM, Frebourg T, Lefebvre H (2001) Identification of a novel mutation in the autoimmune regulator (AIRE-1) gene in a French family with autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy. Eur J Endocrinol 144:347–351
Elkin SK, Ivanov D, Ewalt M, Ferguson CG, Hyberts SG, Sun ZY, Prestwich GD, Yuan J, Wagner G, Oettinger MA, Gozani OP (2005) A PHD finger motif in the C terminus of RAG2 modulates recombination activity. J Biol Chem 280:28701–28710
Gozani O, Field SJ, Ferguson CG, Ewalt M, Mahlke C, Cantley LC, Prestwich GD, Yuan J (2005) Modification of protein sub-nuclear localization by synthetic phosphoinositides: evidence for nuclear phosphoinositide signaling mechanisms. Adv Enzyme Regul 45:171–185
Ragvin A, Valvatne H, Erdal S, Arskog V, Tufteland KR, Breen K, Yan AM, Eberharter A, Gibson TJ, Becker PB, Aasland R (2004) Nucleosome binding by the bromodomain and PHD finger of the transcriptional cofactor p300. J Mol Biol 337:773–788
Denslow SA, Wade PA (2007) The human Mi-2/NuRD complex and gene regulation. Oncogene 26:5433–5438
Misra S, Miller GJ, Hurley JH (2001) Recognizing phosphatidylinositol 3-phosphate. Cell 107:559–562
Stenmark H, Aasland R, Driscoll PC (2002) The phosphatidylinositol 3-phosphate-binding FYVE finger. FEBS Lett 513:77–84
Coscoy L, Sanchez DJ, Ganem D (2001) A novel class of herpesvirus-encoded membrane-bound E3 ubiquitin ligases regulates endocytosis of proteins involved in immune recognition. J Cell Biol 155:1265–1273
Lu Z, Xu S, Joazeiro C, Cobb MH, Hunter T (2002) The PHD domain of MEKK1 acts as an E3 ubiquitin ligase and mediates ubiquitination and degradation of ERK1/2. Mol Cell 9:945–956
Jones DR, Divecha N (2004) Linking lipids to chromatin. Curr Opin Genet Dev 14:196–202
Uchida D, Hatakeyama S, Matsushima A, Han H, Ishido S, Hotta H, Kudoh J, Shimizu N, Doucas V, Nakayama KI, Kuroda N, Matsumoto M (2004) AIRE functions as an E3 ubiquitin ligase. J Exp Med 199:167–172
Aravind L, Iyer LM, Koonin EV (2003) Scores of RINGS but no PHDs in ubiquitin signaling. Cell Cycle 2:123–126
Scheel H, Hofmann K (2003) No evidence for PHD fingers as ubiquitin ligases. Trends Cell Biol 13:285–287, author reply 287–288
Bunce MW, Bergendahl K, Anderson RA (2006) Nuclear PI(4,5)P(2): a new place for an old signal. Biochim Biophys Acta 1761:560–569
Eberharter A, Vetter I, Ferreira R, Becker PB (2004) ACF1 improves the effectiveness of nucleosome mobilization by ISWI through PHD-histone contacts. EMBO J 23:4029–4039
Shi X, Hong T, Walter KL, Ewalt M, Michishita E, Hung T, Carney D, Pena P, Lan F, Kaadige MR, Lacoste N, Cayrou C, Davrazou F, Saha A, Cairns BR, Ayer DE, Kutateladze TG, Shi Y, Cote J, Chua KF, Gozani O (2006) ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442:96–99
Pena PV, Musselman CA, Kuo AJ, Gozani O, Kutateladze TG (2009) NMR assignments and histone specificity of the ING2 PHD finger. Magn Reson Chem 47:352–358
Pena PV, Davrazou F, Shi X, Walter KL, Verkhusha VV, Gozani O, Zhao R, Kutateladze TG (2006) Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442:100–103
Lan F, Collins RE, De Cegli R, Alpatov R, Horton JR, Shi X, Gozani O, Cheng X, Shi Y (2007) Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature 448:718–722
Laherty CD, Billin AN, Lavinsky RM, Yochum GS, Bush AC, Sun JM, Mullen TM, Davie JR, Rose DW, Glass CK, Rosenfeld MG, Ayer DE, Eisenman RN (1998) SAP30, a component of the mSin3 corepressor complex involved in N-CoR-mediated repression by specific transcription factors. Mol Cell 2:33–42
Campos EI, Xiao H, Li G (2004) Generation of a polyclonal antibody specifically against the p33(ING1b) tumor suppressor. J Immunoassay Immunochem 25:71–80
Sugasawa K, Akagi J, Nishi R, Iwai S, Hanaoka F (2009) Two-step recognition of DNA damage for mammalian nucleotide excision repair: directional binding of the XPC complex and DNA strand scanning. Mol Cell 36:642–653
Feng X, Bonni S, Riabowol K (2006) HSP70 induction by ING proteins sensitizes cells to tumor necrosis factor alpha receptor-mediated apoptosis. Mol Cell Biol 26:9244–9255
Huang W, Zhang H, Davrazou F, Kutateladze TG, Shi X, Gozani O, Prestwich GD (2007) Stabilized phosphatidylinositol-5-phosphate analogues as ligands for the nuclear protein ING2: chemistry, biology, and molecular modeling. J Am Chem Soc 129:6498–6506
Kaadige MR, Ayer DE (2006) The polybasic region that follows the plant homeodomain zinc finger 1 of Pf1 is necessary and sufficient for specific phosphoinositide binding. J Biol Chem 281:28831–28836
Martin DG, Baetz K, Shi X, Walter KL, MacDonald VE, Wlodarski MJ, Gozani O, Hieter P, Howe L (2006) The Yng1p plant homeodomain finger is a methyl-histone binding module that recognizes lysine 4-methylated histone H3. Mol Cell Biol 26:7871–7879
Palacios A, Garcia P, Padro D, Lopez-Hernandez E, Martin I, Blanco FJ (2006) Solution structure and NMR characterization of the binding to methylated histone tails of the plant homeodomain finger of the tumour suppressor ING4. FEBS Lett 580:6903–6908
Jones DR, Bultsma Y, Keune WJ, Halstead JR, Elouarrat D, Mohammed S, Heck AJ, D’Santos CS, Divecha N (2006) Nuclear PtdIns5P as a transducer of stress signaling: an in vivo role for PIP4Kbeta. Mol Cell 23:685–695
Shi X, Gozani O (2005) The fellowships of the INGs. J Cell Biochem 96:1127–1136
Bunce MW, Gonzales ML, Anderson RA (2006) Stress-ING out: phosphoinositides mediate the cellular stress response. Sci STKE 2006:pe46
Irvine RF (2003) Nuclear lipid signalling. Nat Rev Mol Cell Biol 4:349–360
Gonzalez L, Freije JM, Cal S, Lopez-Otin C, Serrano M, Palmero I (2006) A functional link between the tumour suppressors ARF and p33ING1. Oncogene 25:5173–5179
Xin H, Yoon HG, Singh PB, Wong J, Qin J (2004) Components of a pathway maintaining histone modification and heterochromatin protein 1 binding at the pericentric heterochromatin in mammalian cells. J Biol Chem 279:9539–9546
Pena PV, Hom RA, Hung T, Lin H, Kuo AJ, Wong RP, Subach OM, Champagne KS, Zhao R, Verkhusha VV, Li G, Gozani O, Kutateladze TG (2008) Histone H3K4me3 binding is required for the DNA repair and apoptotic activities of ING1 tumor suppressor. J Mol Biol 380:303–312
Vieyra D, Loewith R, Scott M, Bonnefin P, Boisvert FM, Cheema P, Pastyryeva S, Meijer M, Johnston RN, Bazett-Jones DP, McMahon S, Cole MD, Young D, Riabowol K (2002) Human ING1 proteins differentially regulate histone acetylation. J Biol Chem 277:29832–29839
Goeman F, Otto K, Kyrylenko S, Schmidt O, Baniahmad A (2008) ING2 recruits histone methyltransferase activity with methylation site specificity distinct from histone H3 lysines 4 and 9. Biochim Biophys Acta 1783:1673–1680
Kataoka H, Bonnefin P, Vieyra D, Feng X, Hara Y, Miura Y, Joh T, Nakabayashi H, Vaziri H, Harris CC, Riabowol K (2003) ING1 represses transcription by direct DNA binding and through effects on p53. Cancer Res 63:5785–5792
Skowyra D, Zeremski M, Neznanov N, Li M, Choi Y, Uesugi M, Hauser CA, Gu W, Gudkov AV, Qin J (2001) Differential association of products of alternative transcripts of the candidate tumor suppressor ING1 with the mSin3/HDAC1 transcriptional corepressor complex. J Biol Chem 276:8734–8739
Simpson F, Lammerts van Bueren K, Butterfield N, Bennetts JS, Bowles J, Adolphe C, Simms LA, Young J, Walsh MD, Leggett B, Fowles LF, Wicking C (2006) The PCNA-associated factor KIAA0101/p15(PAF) binds the potential tumor suppressor product p33ING1b. Exp Cell Res 312:73–85
Garkavtsev I, Grigorian IA, Ossovskaya VS, Chernov MV, Chumakov PM, Gudkov AV (1998) The candidate tumour suppressor p33ING1 cooperates with p53 in cell growth control. Nature 391:295–298
Binda O, Nassif C, Branton PE (2008) SIRT1 negatively regulates HDAC1-dependent transcriptional repression by the RBP1 family of proteins. Oncogene 27:3384–3392
Gong W, Russell M, Suzuki K, Riabowol K (2006) Subcellular targeting of p33ING1b by phosphorylation-dependent 14-3-3 binding regulates p21WAF1 expression. Mol Cell Biol 26:2947–2954
Sarker KP, Kataoka H, Chan A, Netherton SJ, Pot I, Huynh MA, Feng X, Bonni A, Riabowol K, Bonni S (2008) ING2 as a novel mediator of transforming growth factor-beta-dependent responses in epithelial cells. J Biol Chem 283:13269–13279
Smith KT, Martin-Brown SA, Florens L, Washburn MP, Workman JL (2010) Deacetylase inhibitors dissociate the histone-targeting ING2 subunit from the Sin3 complex. Chem Biol 17(1):65–74
Nozell S, Laver T, Moseley D, Nowoslawski L, De Vos M, Atkinson GP, Harrison K, Nabors LB, Benveniste EN (2008) The ING4 tumor suppressor attenuates NF-kappaB activity at the promoters of target genes. Mol Cell Biol 28:6632–6645
Champagne KS, Saksouk N, Pena PV, Johnson K, Ullah M, Yang XJ, Cote J, Kutateladze TG (2008) The crystal structure of the ING5 PHD finger in complex with an H3K4me3 histone peptide. Proteins 72:1371–1376
Saksouk N, Avvakumov N, Champagne KS, Hung T, Doyon Y, Cayrou C, Paquet E, Ullah M, Landry AJ, Cote V, Yang XJ, Gozani O, Kutateladze TG, Cote J (2009) HBO1 HAT complexes target chromatin throughout gene coding regions via multiple PHD finger interactions with histone H3 tail. Mol Cell 33:257–265
Hung T, Binda O, Champagne KS, Kuo AJ, Johnson K, Chang HY, Simon MD, Kutateladze TG, Gozani O (2009) ING4 mediates crosstalk between histone H3 K4 trimethylation and H3 acetylation to attenuate cellular transformation. Mol Cell 33:248–256
Palacios A, Munoz IG, Pantoja-Uceda D, Marcaida MJ, Torres D, Martin-Garcia JM, Luque I, Montoya G, Blanco FJ (2008) Molecular basis of histone H3K4me3 recognition by ING4. J Biol Chem 283:15956–15964
Acknowledgments
This work was supported by the Canadian Institutes of Health Research (MOP-84559 and MOP-93810) and the Canadian Dermatology Foundation to G. Li. R.P.C.W. is a recipient of a Terry Fox Foundation Research Studentship from Canadian Cancer Society Research Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aguissa-Touré, AH., Wong, R.P.C. & Li, G. The ING family tumor suppressors: from structure to function. Cell. Mol. Life Sci. 68, 45–54 (2011). https://doi.org/10.1007/s00018-010-0509-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0509-1