Abstract
Homing genes encode endonucleases that make a double stranded break in the DNA, destroying a target site on the homologous chromosome. When the cell repairs the break the homing allele is copied, converting a heterozygote into a homozygote. This results in gene drive (GD), an overrepresentation of the homing allele in the next generation. GD may propel CRISPR-Cas9 genes and new genes physically coupled to the GD through natural populations. I revisit the population genetic models of GD with the aim of making these models more understandable to non-specialists. What can we learn about risk evaluation from the models? A GD with no or a small effect on fitness (viability) always spreads in the population and goes to fixation. That is provided that no resistance mechanism evolves, for instance due to a mutation in the target site. However, when GDs have a large negative effect on fitness, their spread depends on a threshold or they may not spread at all. The chance of GDs increasing until fixation is much higher in systems with meiotic drive than in systems with embryo conversion. The presence or absence of a meiotic promoter is therefore relevant to take into account in the environmental risk assessment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Gene drives (GD) based on CRISPR-Cas9 currently receive much attention because of their potential ability to propel genes through natural populations (Esvelt et al. 2014). At the same time researchers are worried that a single release or escape can have consequences at a worldwide scale. In general GD is the transmission of an allele to the offspring at a rate higher than the 50% expected under Mendelian inheritance (Burt and Trivers 2006). The oldest known GD system is probably the autosomal t-complex in mice that was discovered in the 1920s (reviewed in Stefanova and Chubykin 2013). In an influential paper Hamilton (1967) discussed GDs in insects, mosquitos included, which were coupled to sex determining genes. Such drives become apparent from variable and strongly biased sex ratios in the offspring. Curtis and co-workers subsequently used these GDs for controlling insect populations, especially mosquitos (Suguna and Curtis 1974). Most naturally occurring GD systems are segregation distorters that use an active mechanism to damage gametes or embryos carrying a wild-type allele (reviewed in Lindholm et al. 2016). This paper focuses only on one class of GDs, that includes CRISPR-Cas9, namely those that use homing in diploid organisms. The homing genes encode endonucleases that create a double stranded break at precisely their own locus in the complementary chromosome. When the cell then uses the GD allele as a template for repairing the destroyed target site, gene conversion occurs and a heterozygous organism becomes homozygous for the GD allele. Homing Endonuclease Genes (HEGs) occur naturally in yeast (Goddard et al. 2005). CRISPR-Cas9 uses the same mechanism and Gantz and Bier (2015) referred to this as the ‘Mutagenic Chain Reaction’ (MCR). With modern biotechnological techniques it is now possible to insert homing alleles into different organisms. The efficiency of the GD depends on the cell using homology-directed repair. Homology-directed repair is a precise method that uses the complementary chromosome as template for repairing the destroyed target site. Alternatively, the cell can use non-homologous end joining, in which case the ends of the broken DNA strings are stitched together and there is no homing (Alberts et al. 2014). This ‘quick and dirty’ repair method usually results in some errors and loss of base pairs where the target site was destroyed. Depending on the specific repair mechanism that the cell uses and the efficiency of homology-directed repair, the transmission of the homing allele may increase from 50% to even close to 100% (Hammond et al. 2016) in a sexual species.
GDs based on homing can be applied in three ways, each with specific consequences for population genetics. First, when a new HEG is introduced in the DNA of a species it will always influence the population in a negative way. HEGs are located in genes and disturb their function. Homozygotes for the HEG have reduced fitness (viability) or may even die immediately (Goddard et al. 2005). When the frequency of the HEG increases, this will lead to a lower population density and possibly even to extinction when females produce too little offspring to replace themselves. CRISPR-Cas9 can be used in the same way when you insert the DNA inside an essential gene (Esvelt et al. 2014). The first upcoming application of these negative GDs is the control of mosquito populations to reduce malaria in humans (www.targetmalaria.org).
Second, in the lab a GD based on CRISPR-Cas9 can also be coupled to a new gene for the organism. When the destroyed target site is repaired using the homologous chromosome as a template, the cell copies not only the GD but also the new gene next to it to the corresponding chromosome, so that they spread in tandem. In this way new functional genes that are themselves selectively neutral or have a small negative effect on fitness can be propelled through the population. These genes that are copied together with the GD are called ‘payload’ or ‘cargo’ genes (Champer et al. 2016). An example is a gene that modifies the ability for mosquitos to transmit the human malaria parasite (Gantz et al. 2015). This gene normally does not increase in frequency in the mosquito population because it gives no advantage to the mosquitos that carry it. It does spread when coupled to a GD.
Thirdly, homing GDs can be used to promote the spread of coupled alleles that increase fitness. This could potentially be of vital importance for the rapid introduction of resistance alleles in populations of species that are now threatened by exotic fungi and viruses from which they have always been isolated (Stegen et al. 2017). Application of GD in breeding programs is based on the same idea (Gonen et al. 2017). One might expect that these new alleles that increase fitness are picked up by selection anyway and increase in frequency. However linkage with other genes that reduce fitness slows this process down (de Jong and Rong 2013). Introducing the payload gene alongside a GD speeds up the increase in frequency.
Fitness-neutral driving alleles always go to fixation into the population (Hartl and Clark 1989). However, when a GD is costly and reduces fitness, it may fail. The public misconception that GDs always increase in frequency may partly be due to terminology used. When it is stated that a GD ‘drives through the population’, the term drive is synonymous to increase. The novel definition of GD (Wikipedia.org) refers to “a technique that promotes the inheritance of a particular gene to increase its prevalence in a population”. This also links GD automatically to increase. However, biologists have used the definition as GD as biased transmission for at least 60 years (Lindholm et al. 2016) and it is unwise to dismiss this information. In summary, GDs that are costly may increase in frequency or they may not, in which case they disappear from the population.
There are four possible outcomes of the genetic model of a costly GD-allele in a large well-mixed, outbreeding population (Hartl and Clark 1989; Deredec et al. 2008). (A) The driving allele always increases in frequency and goes to fixation, even when it is initially rare. (B) The driving allele goes to fixation only when introduced at a minimum frequency, but goes extinct when it is below this threshold. This is similar to the outcome of a genetic model of a ‘normal’ non-driving gene with two alleles and underdominance (fitness of the heterozygote is less than that of the two homozygotes). (C) The GD will become extinct if fitness costs are too high. (D) Finally a driving allele will coexist with a non-driving allele. This is similar to the equilibrium of two alleles with overdominance (fitness of the heterozygote is higher than that of the two homozygotes).
The rest of this paper is concerned with GDs that have negative effects on fitness. When will these GDs sweep through the population? For biological control it is important that this process goes quickly and that the wild-type allele is completely eliminated. The longer it takes for the GD allele to go to fixation, the greater the chance that resistant alleles arise along the way. The success of the homing GD depends on three factors: (i) The effect of the GD on fitness. (ii) The degree of dominance of the fitness cost of the driving allele in heterozygotes. (iii) The life stage (meiosis or embryo) during which gene conversion takes place. I revisit the genetic models of GD with the aim of outlining in a graphical way when GDs will and will not increases in the population. Then the relevance of these results for risk evaluation is discussed.
2 Results
2.1 Model 1: Gametic drive
This is the first model that Deredec et al. (2008) dealt with under the heading ‘HEG active after gene expression’. In the heterozygote organism the homing allele cuts the wild-type allele after it has been functional, i.e. during meiosis at the end of the life cycle of the organism. This is model 1 or meiotic drive. The probability of successful conversion of a gamete is e (0 ≤ e ≤ 1) and a heterozygote Dd then produces 1 + e gametes with driving allele D and 1-e gametes with the wild-type allele d (Table 1). Parameter e is also referred to as the homing rate. The fraction D alleles in the gametes is (1 + e)/2. Mendelian inheritance occurs when e = 0 and with e = 1 all gametes are of type D. The fitness (viability) of homozygote DD is reduced by a factor 1-s, in which s denotes the selection coefficient. The fitness cost of the driving allele D in the heterozygote can be zero, when it is recessive. The effect can also be as large as in the homozygote DD, equal to s, when D is dominant. The parameter h describes the degree of dominance of the fitness effect and h ranges from 0 (recessive) to 1 (dominant). The fitness of the heterozygote is 1-hs. The recurrence equations for model 1 can be derived from Fig. 1 and were given by Deredec et al. (2008, their Eq. 1).
Flow diagram of gene drive (homing). The driving allele is denoted as D and the wild-type allele as d. Gene conversion may occur in the embryo, as measured by parameter c, or in during meiosis (parameter e). The fitness (viability or number of gametes produced) of individuals homozygous for D is reduced by a factor 1-s (s is the selection coefficient). The fitness of heterozygotes is reduced by 1-hs, in which h measures the dominance of the fitness effect of the driving allele D. The text refers to embryo conversion as model 1 and meiotic drive as model 2 and keeps these processes separate. However, they could both occur in the same organism
2.2 Model 2: Embryo conversion
Unckless et al. (2015) described GD caused by a CRISPR-Cas9 gene in the following way: “The wild-type allele in heterozygous individuals is converted to the MCR allele at rate c. We assume this happens in the embryo so that individuals that experienced conversion essentially become homozygous and have fitness 1-s.” This is what one would expect if the GD is always active and fully in line with the description of the process in Gantz and Bier (2015). However, if one would combine the GD with a meiosis specific promoter, then homing would only occur during meiosis and this is model 1. This application already exists (for instance, the paper by Hammond et al. 2016 that is discussed later). So CRISPR-Cas9 gene drive is not necessarily the model of embryo conversion. The model of embryo conversion developed by Unckless et al. (2015) is identical to the second model analyzed by Deredec et al. (2008, their Eq. 8) under the heading ‘HEG active before gene expression’. This is model 2 or embryo conversion. Since conversion of gametes and embryos are two distinct processes it is most clear to denote them by different symbols and use c for the rate of embryo conversion (0 ≤ c ≤ 1). One could argue that gene conversion can take place at any time during the life cycle. However, the cell is most likely to use the precise, costly homology-directed repair in critical stages, notably the embryo stage and meiosis, and use non-homologous end joining in between. If this is true, homing in gamete and embryo are two distinct events. Figure 1 shows the full model, including both c and e.
The iconic scheme of GD (Esvelt et al. 2014) shows a fly homozygous for the driving allele (shown as blue) that is crossed with a wild-type fly (black) to produce blue offspring, all with the GD. But this scheme and extensions (Collins and Heitman 2016, their Fig. 1.2) are quite confusing if one wants to understand the genetics. All versions of model 1 will need to include heterozygote flies next to the homozygotes; all offspring from a cross between DD and dd will be Dd for their entire life. Homing occurs only in the gametes. Also model 2 will generally include heterozygotes. Only with 100% embryo conversion Dd individuals are never produced. Figure 2 illustrates that the iconic scheme of GD assumes model 2 and 100% homing in embryos (c = 1). Figure 1 is more complex than the figure from Esvelt et al. (2014) but is clear about underlying genetics and the differences between the two models.
The full model of homing gene drive from Fig. 1 corresponds to the iconic picture of gene drive only when one assumes embryo conversion (model 2) and 100% efficiency of gene conversion
3 Discussion of model outcomes
In model 2 the heterozygote is transformed into a DD homozygote at the start of embryonic life and then makes 1-s zygotes at the end of its life. In model 1 the individual remains in the heterozygote state until the end of its life and then produces 1-hs zygotes. The models are identical in two cases. First, when the driving allele is fully dominant (h = 1) the heterozygote passes on 1-s zygotes with D in both model 1 and model 2. Second, when there are no fitness costs of having this driving allele (s = 0) the heterozygote passes on one zygote with the D allele in both models. In all other cases it is much easier in model 1 for a driving allele to invade the population. For instance, assume the wild-type homozygote dd produces 10 gametes. With s = 0.2, h = 0.5 and e = 1, in model 1 the heterozygote will make 9 gametes, all of type D. In model 2 the Dd heterozygote is converted soon after embryo formation into a homozygote DD that suffers the full fitness cost s and makes only 8 gametes of type D.
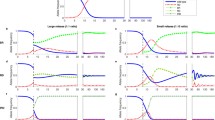
Figure 3a shows the results for model 1 plotted using the graphical method of Alphey and Bonsall (2014). The two lines I and II divide the four areas where the outcome of the model differs. (A) The driving allele increases from rare to 100% in area A, i.e. when selection against the driving allele is weak and when the effects of the driving allele on fitness are recessive. (B) With higher selection coefficients and more dominance of the driving allele, there is a threshold that the driving allele must reach before it will increase in frequency (area B). (C) At the highest values of s and h the driving allele goes extinct (area C). (D) Finally, with high s and low h coexistence of the driving and non-driving allele is possible (area D). Line II crosses the X-axis at point (e, 0), in the example in Fig. 3a e = 0.8 (for further details on the math, see Deredec et al. 2008). Higher conversion rates (e.g. e = 0.9) shift line II to the right. When e is very high, it is it is possible for initially rare recessive genes (h = 0) with large negative effects on fitness, to spread in the population. Theoretically this makes it possible in model 1 to reduce fitness to such an extent that the density of the target organism is compromised, possibly even to the point where the species can no longer replace itself.
a Model 1 of gametic drive. The fate of the driving allele depends on its fitness cost (s) and the degree of dominance (h) of this fitness cost in heterozygotes. The rate of successful conversion during meiosis is set at e = 0.8. Four possible outcomes of the model are shown. The equation of the separating line I is h = e/((1 + e)s) and for line II it is h = (e − s)/(s(e − 1)) (Deredec et al. 2008). Starting clockwise in the top left corner, the points on the edge are (e/(1 + e), 1), (1, 1), (1, e/(1 + e) and (e, 0). b Model 2 of embryo conversion. The rate of successful gene conversion in the embryonic stage is set at c = 0.8. With this value of c there are only three possible outcomes of the model. The equation of the separating line I is h = (c − 2sc)/(s(1 − c)) and for line II it is h = (c − s)/(s(c − 1)), which is the same as in a) (Deredec et al. 2008). Starting clockwise in the top left corner, the points on the edge are (c/(1 + c), 1), (1, 1), (c, 0) and (0.5, 0). The circle at (1, 0) indicates a fully recessive, lethal driving allele. Such an allele will always go extinct, regardless of the frequency of introduction
For model 2 line II stays in place (Fig. 3b). Line I stays attached to the point c/(1 + c) at the top. However, compared to model 1 the line now swings to the left, as though a curtain is closed at the end of the show. In model 2 line I crosses the X-axis always at (0.5, 0). For c = 0.8 there are only three possible outcomes. (A) At low values of s and h, the driving allele goes to fixation (area A). (B) With higher fitness cost the driving allele no longer spreads when it is rare, but it does spread and goes to fixation when introduced at a frequency that is above a certain threshold (area B). (C) Finally with the highest fitness cost the driving allele goes extinct (area C). For c = 0.8 coexistence of the two alleles, outcome D, is not possible. Coexistence is possible in model 2 with other parameter values (e.g. with low values for embryo conversion, c = 0.2). This is not relevant in the context of current GD applications, which appear to be quite efficient (c = 0.90–0.98). In model 2 rare driving alleles can only spread if they reduce fitness with less than 40%. This result depends only weakly on the gene conversion rate (c). Line I is almost horizontal for c = 0.8 and becomes a fully horizontal line at s = 0.5 at c = 1. The main difference between models 1 and 2 is that the area separated by line I, as it swings down, changes from ‘GD increases from rare’ (A) to ‘GD only increases above a certain threshold’ (B).
An additional aspect is that the speed at which the driving allele goes to fixation depends on the value of s. The spread from 0.1% to almost 100% is achieved in 13 generations without fitness costs (s = 0) and 20 generations with low fitness costs (s = 0.2) (Fig. 4). With 40% fitness costs (s = 0.4), well within region A in which the GD increases, the spread is initially very slow and is only completed after 55 generations. For practical purposes one could say that in model 2 only GDs with fitness effects of 30% or less will increase from rare to common within a suitable time span.
Spread of a gene drive in model 2 of embryo conversion as a function of time (one generation per year). Fitness costs slow down the rate of increase of the gene drive. Solid line selection coefficient s = 0, broken line s = 0.2, dashed line s = 0.4. For all simulations h = 0.5 and c = 0.8. The driving allele was introduced at a frequency of 0.001. Note that although s = 0.4 is well within the region in which the driving allele spreads in the population (Fig. 3b), the rate of increase is initially very slow
When comparing models 1 and 2, it is evident that genes with large fitness costs can go from rare to common in model 1 but not in model 2. For mosquitoes a female produces in the order of 9.2 daughters over her life (Deredec et al. 2011). To extinguish the population one would need multiplicative effects on fitness in the order of 1/9.2 or fitness reduction of almost 90% (s = 0.9). This would be theoretically possible with a single GD in model 1. It would be impossible with a single GD in model 2. Population reduction in model 2 could only be achieved by applying many independent GDs. Note that the genetic models assume a large well-mixed population. Any spatial structure in the population or local mate competition (Hamilton 1967) reduces the fraction heterozygotes in the population and makes it more difficult for the GD to invade. Homing GDs depend on sexual reproduction. A shift in life history from outcrossing to selfing slows the GD down. In a selfing species a homing GD will come to a standstill and will disappear from the population when costly. This was shown experimentally by Goddard et al. (2005) for yeast.
3.1 Differences between the sexes
So far the genetic models assumed that all parameters are the same for females and males. In general this is not the case. In natural systems GD often affects males or females only (Lindholm et al. 2016). Model parameters may also differ between the sexes. Deredec et al. (2008) gave for model 1 the coupled recurrence equations (their Eq. 15), noticing that “the general solution to these equations is too complex to be helpful”. It is therefore useful to look for simplifications in order to analyze these cases with the graphical method of Fig. 3. Using CRISPR-Cas9 Hammond et al. (2016) built three different GDs in mosquitoes with negative effects on female viability. They used the meiosis-specific promoter vasa2, which is active in the germ line of both sexes. This falls under model 1 for gametic drive. The GD ‘AGAP007280’ had a homing rate of 0.984. All homozygous females (subscript f) died (s f = 1) and the fertility of heterozygotes was 9.3% of that of normal females (h f = 0.907). Hammond et al. (2016) assumed no effect of the GD on the fitness of males (subscript m) (s m = 0), in which case the value of h m does not matter for results (h m s m is a zero anyway). Using the appropriate Eq. 15 for model 1 in Deredec et al. (2008), Hammond et al. (2016) simulated the spread of GD ‘AGAP007280’ from 5 to 100% in about 40 generations. From simulations using this equation, I noted that (with constant e and h) it does not matter whether one uses the average value of s or the separate values for each sex. For instance, s = 0.5 gives identical results as s m = 0.9 and s f = 0.1 or s m = 1 and s f = 0. Apparently for the spread of the GD it does not matter whether one targets males or females. Note though that this is a genetic model of allele frequencies. Population density would be much more affected if females were targeted instead of males. If selection coefficients for males and females in the Hammond et al. (1996) data are averaged this gives s = 0.5. With e = 0.984 line II shifts to the far right of the graph. Assuming that h is similar for males and females, Fig. 3a can now be redrawn for the specific parameter values of GD ‘AGAP007280’ (results not shown). This figure places this GD firmly in area A where it goes from rare to fixation.
4 General discussion
4.1 Ideal GD construct for biocontrol
An ideal GD for controlling a pest species should go rapidly to fixation and reduce the population density, possibly even to extinction. When the time to fixation is long or when there is coexistence with the wild-type allele, there is a higher chance for mutations to generate alleles that are resistant to GD (Unckless et al. 2017). When resistant alleles are still functional they will replace the costly GD allele. GDs in area A of Fig. 3a, b are easiest to introduce in the field, since they will increase after introducing just few individuals. GDs in area B require the release of many individuals to exceed the threshold. An advantage of a threshold may be that, in principle, releasing large numbers of non-GD individuals will revoke the GD by bringing it under its threshold. Although distinction between these two scenarios (no threshold/threshold) is clear in the models, it will be less clear in practice. If organisms die in the lab they will probably also die in the field, but for GDs with less drastic effects nobody will be able to predict in advance whether the selection coefficient will be e.g. 0.3 or 0.7. Fitness effects also typically depend on the local environment so the same GD may have s = 0.3 in one environment and s = 0.7 in a more stressful environment. The GD could then have no threshold in the first case, while there is one in the stressful environment. This genotype-environment interaction should be incorporated in risk assessment of GDs.
The models illustrate that GDs with no or mildly negative effects will always go to fixation if no resistance against the GD develops. For GDs with a strong negative effect on fitness, potentially most useful for biocontrol, the spread is not so clear-cut. This leads to a dilemma for developers. A GD with large negative effect is most effective but may not increase in frequency. A GD with a small effect will increase in frequency but is much less effective in reducing the population. Apart from the selection coefficient one could select for GDs with a low rate of dominance. This would increase the chance of success of the GD in model 1 (Fig. 3a), but much less in model 2 (Fig. 3b). Designing a GD that works only during meiosis (model 1) and not in the embryo stage (model 2) seems the most promising strategy. A GD spreads much easier and also faster (Deredec et al. 2008) in model 1 than in model 2. With homing rate e close to 1 line II becomes horizontal in Fig. 3A, starting in (1,0). In this case even GDs with large selection coefficients would take off from a very low starting frequency. If only one of the sexes is affected (s = 1) and the other is not (s = 0), as in the Hammond et al. (2016) paper, average s can be used in model 1 to predict the outcome. With s = 0.5 (and e = 1) all GDs go from rare to fixation in model 1, so targeting one of the sexes could well be a viable strategy for biocontrol.
The effect of the life stage during which GD operates has been mentioned in the literature but has not been given full attention. For instance, the report ‘Gene drives on the horizon’ from the US Academies of Sciences (Collins and Heitman 2016) reviewed the population genetic models of GD without mentioning life stage as a determining factor. On the other hand Burt (2003) commented in his landmark paper on the ideal construct for biological control: “…the HEG is under the control of a meiosis- specific promoter, so that heterozygous zygotes develop normally, but transmit the HEG to a disproportionate fraction of their gametes.” Deredec et al. (2008) commented for model 2: “Thus the strategy of creating a recessive lethal (s = 1, h = 0) will not work if homing occurs prior to gene expression.” The point they make is easy to see in Fig. 3b, in which the mentioned recessive lethal is indicated by the circle at (1,0).
4.2 Safety
There is concern that GDs with a negative effect on fitness and population density will escape from the lab and can spread globally. Abkari et al. (2015) advised that in lab experiments the two elements (single synthetic guide RNA and Cas9) necessary to create a GD should be kept separate on the genome. Due to some chromosomal rearrangement these elements could, with low probability, be combined and would then produce a GD. Abkari et al. (2015) advised using both molecular and ecological confinement strategies. While strict security measures seem a wise strategy for this new technology, it should be realized that the GDs that arise by chance are likely to be model 2 (the endonuclease becomes immediately active in the embryo) and that if the GD has large negative effects on fitness it will simply not spread or requires a threshold. Only when a meiosis-specific promoter is added it becomes theoretically possible to move genes with large negative effects on fitness through the population. The presence or absence of meiosis-specific promoters is an essential issue for developers and also for the safety evaluation.
4.3 Risk evaluation
In the context of environmental risk assessment (ERA) of GDs I think it is useful to refer to the well-known equation: risk = hazard × probability of occurrence. The risk of a GD with large negative effect on viability escaping from the lab has a clear hazard (extinction of target population and/or that of related species) but in model 2 the GD will rapidly go extinct so it will not occur. When a GD with a small effect on viability escapes, the probability that it increases in frequency is very high. However, the hazard, the effect on the population, is low. Most organisms, mosquitos included, produce large numbers of offspring and because of density-dependence a small reduction in viability will hardly affect the number of offspring that survives to reproduce. In both cases we need a clear hypothesis of what the hazard is when a GD with its payload gene escapes from the lab or is introduced in nature. If the GD increases, goes to fixation and then slowly degrades but without any measurable effects on the population this is not per se a risk. For instance, if a GD with s = 0.3 goes to fixation in a mosquito population the number of daughters per female will go from 9.2 to 6.44. This is likely to have no effect whatsoever on population numbers. The effect of the payload gene may be more important for the hazard as the GD itself. Natural selection may eventually eliminate payload genes with negative effects on fitness from the population after the GD has run its course. Payload genes with positive fitness effects, for instance increased weediness or resistance, will be there to stay after these alleles have established in the population. The risk evaluation for GDs can be compared to that for genetically modified (GM) crops that can outcross with wild relatives (Ellstrand et al. 2013). Crop genes, transgenes included, will occasionally flow from crop to wild relatives but are now considered not to pose a risk unless the gene affects population densities. The ERA guidance (EFSA 2010) asks the following questions. “Will the GM trait affect the fitness of the organism and its relatives in its natural habitat?” “Will the GM trait affect the geographical range of the organism?” “Will the GM trait cause populations to change in size?” “Will this cause damage to the environment or human health?”
Similarly EFSA guidance for GM animals (EFSA 2013) asks the following. “Does the GM insect have the potential to persist or invade EU receiving environments?” To what extent can the GM insect species reproduce and hybridise with non-GM insects of the same or different species under EU conditions to produce viable and fertile offspring?” “Will the GM trait confer increased fitness to the resulting population that could allow it to persist or invade more than its non-GM comparator?” “Will the GM trait alter the habitat and/or the geographic range of the GM species or hybrid populations?” On the basis of these questions applicants should evaluate the persistence and invasiveness of the GM insect and any hybrid offspring and propose risk management strategies.
In combination with genetic models and models of the dynamics of populations, these questions for GM crops (EFSA 2010) and GM animals (EFSA 2013) seem a good starting point for the Environmental Risk Assessment of GDs.
References
Abkari OS et al (2015) Safeguarding gene drive experiments in the laboratory. Science 349:927–929
Alberts B, Johnson A, Lewis J, Morgan D, Raff M, Roberts K, Walter P (2014) Molecular biology of the cell. Garland Science, Milton Park
Alphey N, Bonsall MB (2014) Interplay of population genetics and dynamics in the genetic control of mosquitoes. J R Soc Interface 11:20131071
Burt A (2003) Site-specific selfish genes as tools for the control and genetic engineering of natural populations. Proc R Soc Lond Ser B 270:921–928
Burt A, Trivers R (2006) Genes in conflict: the biology of selfish genetic elements. Harvard University Press, Harvard
Champer J, Buchman A, Abkari S (2016) Cheating evolution: engineering gene drives to manipulate the fate of wild populations. Nat Rev Genet 17:146–159
Collins JP, Heitman E (2016) Gene drives on the horizon. Advancing science, navigating uncertainty, and aligning research with public values. The National Academies Press, Washington
de Jong TJ, Rong J (2013) Crop to wild gene flow: does more sophisticated research provide better risk assessment? Environ Sci Policy 27:135–140
Deredec A, Burt A, Godfray HCJ (2008) The population genetics of using homing endonuclease genes in vector and pest management. Genetics 179:2013–2026
Deredec A, Godfray CJ, Burt A (2011) Requirements for effective malaria control with homing endonuclease genes. Proc Nat Acad Sci 108(43):E874–E880. doi:10.1073/pnas.1110717108
EFSA (2010) Guidance of the environmental risk assessment of genetically modified plants. EFSA J 8:1879. doi:10.2903/j.efsa.2010.1879
EFSA (2013) Guidance on the environmental risk assessment of genetically modified animals. EFSA J 11:3200. doi:10.2903/j.efsa.2013.3200
Ellstrand NM et al (2013) Introgression of crop alleles into wild or weedy populations. Ann Rev Ecol Syst 44:1–17
Esvelt KM, Smidler AL, Catteruccia F, Church GM (2014) Concerning RNA-guided gene drives for the alteration of wild populations. eLife 3:e03401. doi:10.7554/elife.03401
Gantz VM, Bier E (2015) The mutagenic chain reaction: a method for converting heterozygous to homozygous mutations. Science 348:442–444
Gantz VM, Jasinskiene N, Tatarenkova O, Fazekas A, Macias VM, Bier E, James AA (2015) Highly efficient Cas9-mediated gene drive for population modification of the malaria vector Anopheles stephensi. Proc Nat Acad Sci 112(49):E6736–E6743. doi:10.1073/pnas.1521077112
Goddard MR, Godfray HCJ, Burt A (2005) Sex increases the efficacy of natural selection in experimental yeast populations. Nature 434:636–640
Gonen S, Jenko J, Gorjanc G, Mileham AJ, Whitelaw CBA, Jickey JM (2017) Potential of gene drives with genome editing to increase genetic gain in livestock breeding programs. Genet Sel Evol 49:3. doi:10.1186/s12711-016-0280-3
Hamilton WD (1967) Extraordinary sex ratios. Science 145:477–488
Hammond A et al (2016) A CRISPR-Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nat Biotech 34:78–83
Hartl DJ, Clark AG (1989) Principles of population genetics, 2nd edn. Sinauer, Sunderland
Lindholm AK et al (2016) The ecology and evolutionary dynamics of meiotic drive. Trends Ecol Evol 31:315–326
Stefanova LD, Chubykin VL (2013) Meiotic drive in mice carrying t-complex in their genome. Genetika 49:1021–1035
Stegen G et al (2017) Drivers of salamander extirpation mediated by Batrachochytrium salamandrivorans. Nature 544:353–356
Suguna SG, Curtis CF (1974) Sex ratio distorter strains in Aedes aegypti. J Commun Dis 6:102–105
Unckless RL, Messer PW, Conallon T, Clark AG (2015) Modeling the manipulation of natural populations by the mutagenic chain reaction. Genetics 201:425–431
Unckless RL, Clark AG, Messer PW (2017) Evolution of resistance against CRISPR-Cas9 gene drive. Genetics 205:827–841
Acknowledgements
This paper was inspired by the workshop ‘Challenges for the regulation of gene drive technology’ that was organized at the Lorentz Center of Leiden University from 20 to 24 March 2017. I thank Nina Alphey, Detlef Bartsch, Marjan Bovers and Boet Glandorf for discussing these ideas and for their critical comments on the MS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares he has no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
de Jong, T.J. Gene drives do not always increase in frequency: from genetic models to risk assessment. J Consum Prot Food Saf 12, 299–307 (2017). https://doi.org/10.1007/s00003-017-1131-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00003-017-1131-z