Abstract.
Major parts of amino-acid-coding regions of elongation factor (EF)-1α and EF-2 in Trichomonas tenax were amplified by PCR from total genomic DNA and the products were cloned into a plasmid vector, pGEM-T. The three clones from each of the products of the EF-1α and EF-2 were isolated and sequenced. The insert DNAs of the clones containing EF-1α coding regions were each 1,185 bp long with the same nucleotide sequence and contained 53.1% of G + C nucleotides. Those of the clones containing EF-2 coding regions had two different sequences; one was 2,283 bp long and the other was 2,286 bp long, and their G + C contents were 52.5 and 52.9%, respectively. The copy numbers of the EF-1α and EF-2 gene per chromosome were estimated as four and two, respectively.
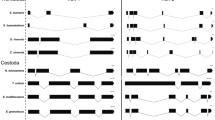
The deduced amino acid sequences obtained by the conceptual translation were 395 residues from EF-1α and 761 and 762 residues from the EF-2s. The sequences were aligned with the other eukaryotic and archaebacterial EF-1αs and EF-2s, respectively.
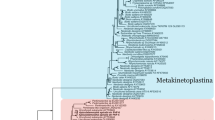
The phylogenetic position of T. tenax was inferred by the maximum likelihood (ML) method using the EF-1α and EF-2 data sets. The EF-1α analysis suggested that three mitochondrion-lacking protozoa, Glugea plecoglossi, Giardia lamblia, and T. tenax, respectively, diverge in this order in the very early phase of eukaryotic evolution. The EF-2 analysis also supported the divergence of T. tenax to be immediately next to G. lamblia.
Similar content being viewed by others
References
Adachi J (1995) Modeling of molecular evolution and maximum likelihood inference on molecular phylogeny. PhD thesis of the graduate University for advanced studies. Institute of Statistical Mathematics, Tokyo
Adachi J, Hasegawa M (1996) Computer science monographs, no. 28, MOLPHY version 2.3: programs in molecular phylogenetics based on maximum likelihood. The Institute of Statistical Mathematics, Tokyo
Baroin A, Perasso RL, Qu GB, Bachellerie J, Adoutte A (1988) Partial phylogeny of the unicellular eukaryotes based on rapid sequencing of a portion of 28S ribosomal RNA. Proc Natl Acad Sci USA 85:3474–3478
Bozner P, Demes P (1991a) Cell-associated and extracellular proteolytic activity of an oral flagellate, Trichomonas tenax. Arch Oral Biol 36:77–83
Bozner P, Demes P (1991 b) Degradation of collagen type I, III, IV and V by extracellular proteinases of an oral flagellate Trichomonas tenax. Arch Oral Biol 36:765–770
Brown JR, Doolittle WF (1995) Root of the universal tree of life based on ancient aminoacyl-tRNA synthetase gene duplications. Proc Natl Acad Sci USA 92:2441–2445
Cao Y, Adachi J, Janke A, Pääbo S, Hasegawa M (1994a) Phylogenetic relationships among eutherian orders estimated from inferred sequences of mitichondrial proteins: instability of a tree based on a single gene. J Mol Evol 39:519–527
Cao Y, Adachi J, Yano T, Hasegawa M (1994b) Phylogenetic place of guinea pigs: no support of the rodent-polyphyly hypothesis from maximum-likelihood analyses of multiple protein sequences. Mol Biol Evol 11:593–604
Cavalier-Smith T (1987) Eukaryotes with no mitochondria. Nature 326: 332–333
Cavalier-Smith T (1993) Kingdom protozoa and it’s 18 phyla. Microbiol Rev 57:953–994
Clark CG, Roger AK (1995) Direct evidence of secondary loss of mitochondria in Entamoeba histolytica. Proc Natl Acad Sci USA 92:6518–6521
Fukura K, Yamamoto A, Hashimoto T, Goto N (1996) Nucleotide sequence of the SrRNA gene and phylogenetic analysis of Trichomonas tenax. Microbiol Immunol 40:183–188
Hasegawa M, Hashimoto T (1993) Ribosomal RNA trees misleading? Nature 361:23
Hasegawa M, Kishino H (1994) Accuracies of the simple methods for estimating the bootstrap probability of a maximum-likelihood tree. Mol Biol Evol 11:142–145
Hasegawa M, Cao Y, Adachi J, Yano T (1992) Rodent polyphyly? Nature 355:595
Hasegawa M, Hashimoto T, Adachi J, Iwabe N, Miyata T (1993) Early branchings in the evolution of eukaryotes: ancient divergence of Entamoeba that lacks mitochondria revealed by protein sequence data. J Mol Evol 36:380–388
Hashimoto T, Adachi J, Hasegawa M (1992) Phylogenetic place of Giardia lamblia, a protozoan that lacks mitochondria. Endocytobiosis Cell Res 9:59–69
Hashimoto T, Nakamura Y, Nakamura F, Shirakura T, Adachi J, Goto N, Okamoto K, Hasegawa M (1994) Protein phylogeny gives a robust estimation for early divergences of eukaryotes: phylogenetic place of a mitochondria-lacking protozoan, Giardia lamblia. Mol Biol Evol 11:65–71
Hashimoto T, Nakamura Y, Kamaishi T, Adachi J, Nakamura F, Okamoto K, Hasegawa M (I 995a) Phylogenetic place of kinetoplastid protozoa inferred from a protein phylogeny of elongation factor 1α. Mol Biolchem Parasitol 70:181–185
Hashimoto T, Nakamura Y, Kamaishi T, Nakamura F, Adachi J, Okamoto K, Hasegawa M (1995b) Phylogenetic place of mitochondrion-lacking protozoan, Giardia lamblia, inferred from amino acid sequence of elongation factor 2. Mol Biol Evol 12:782–793
Hayawan IAE, Bayoumy MM (1992) The prevalence of Entamoeba gingivalis and Trichomonas tenax in periodontal disease. J Egypt Soc Parasitol 22:101–105
Hersh SM (1985) Pulmonary trichomonads and Trichomonas tenax. J Med Microbiol 20:1–10
Kamaishi T, Hashimoto T, Nakamura Y, Nakamura F, Murata S, Okada N, Okamoto K, Shimizu M, Hasegawa M (1996) Protein phylogeny of EF-1 α suggests microsporidians are extremely ancient eukaryotes. J Mol Evol 42:257–263
Kishino H, Miyata T, Hasegawa M (1990) Maximum likelihood inference of protein phylogeny and the origin of chloroplasts. J Mol Evol 30:151–160
Länge S, Rozario C, Muller M (1994) Primary structure of the hydrogenosomal adenylate kinase of Trichomonas vaginalis and its phylogenetic relationships. Mol Biochem Parasitol 66:297–308
Lahti JC, Bradley JP, Johnson JP (1994) Molecular characterization of the α-subunit of Trichomonas vaginalis hydrogenosomal succinyl CoA synthetase. Mol Biochem Parasitol 66:309–318
Leipe DD, Gunderson JH, Nerad TA, Sogin ML (1993) Small subunit ribosomal RNA’ of Hexamita inflata and the quest for the first branch in the eukaryotic tree. Mol Biochem Parasitol 59:41–48
Loomis WF, Smith DW (1990) Molecular phylogeny of Dictyostelium discoideum by protein sequence comparison. Proc Natl Acad Sci USA 87:9093–9097
Markoŝ A, Miretsky A, Muller M (1993) A glyceraldehyde-3-phosphate dehydrogenase with eubacterial features in the amitochondriate eukaryote, Trichomonas vaginalis. J Mol Evol 37:631–643
Miyata T, Iwabe N, Kuma K, Kawanishi Y, Hasegawa M, Kishino H, Mukohata Y, Ihara K, Osawa S (eds) (1991) Evolution of archaebacteria: phylogenetic relationships among archaebacteria, eubacteria, and eukaryotes. In: Osawa S, Honjo T (eds) Evolution of life: fossils, molecules, and culture. Springer-Verlag, Tokyo, pp 337–351
Muffler M (1993) The hydrogenosome. J Gen Microbiol 139:2879–2889
Paces J, Urbánková V, Urbánek P (1992) Cloning and characterization of a repetitive DNA sequence specific for Trichomonas vaginalis. Mol Biochem Parasitol 54:247–256
Qu LH, Perasso R, Baroin A, Brugerolle G, Bachellerie JP, Afoutte A (1988) Molecular evolution of the 5’-terminal domain of largesubunit rRNA from lower eukaryotes. A broad phylogeny covering photosynthetic and non-photosynthetic protists. Biosystems 21: 203–208
Rivera MC, Lake JA (1992) Evidence that eukaryotes and eocyte prokaryotes are immediate relatives. Science 257:74–76
Sakamoto Y, Ishiguro M, Kitagawa G (1986) Akaike information criterion statistics. In: Reidel D (ed) Dordrecht
Shirakura T, Hashimoto T, Nakamura Y, Kamaishi T, Cao Y, Adachi J, Hasegawa M, Yamamoto A, Goto N (1994) Phylogenetic place of a mitochondria-lacking protozoan, Entamoeba histolytica, inferred from amino acid sequences of elongation factor 2. Jpn J Genet 69:119–135
Sogin ML (1989) Evolution of eukaryotic microorganisms and their small subunit ribosomal RNAs. Am Zool 29:487–499
Sogin ML (1991) Early evolution and the origin of eukaryotes. Curr Opin Genet Dev 1:457–463
Sogin ML, Gunderson JH, Elwood HJ, Alonso RA, Peattie DA (1989) Phylogenetic meaning of the kingdom concept: an unusual ribosomal RNA from Giardia lamblia. Science 24:75–77
Thorne JL, Churchill GA (1995) Estimation and reliability of molecular sequence alignments. Biometrics 51:100–113
Thorne JL, Kishino H, Felsenstein J (1991) An evolutionary model for maximum likelihood alignment of DNA sequences. J Mol Evol 33:114–124
Thorne JL, Kishino H, Felsenstein J (1992) Inching toward reality: an improved likelihood model of sequence evolution. J Mol Evol 33: 3–16
Viscogliosi E, Philippe H, Baroin A, Perasso R, Brugerolie G (1993) Phylogeny of trichomonads based on partial sequences of large subunit rRNA and on cladistic analysis of morphological data. J Eukar Microbiol 40:411–421
Vossbrinck CR, Maddox JV, Friedman S, Debrunner-Vossbrinck BA, Woese CR (1987) Ribosomal RNA sequence suggests microsporidia are extremely ancient eukaryotes. Nature 326:411–414
Yamamoto A, Kikuta N, Hashimoto T, Oyaizu H, Goto N (1995) Nucleotide sequence of the SrRNA gene of Entamoeba gingivalis: applications for construction of a species-specific DNA probe and phylogenetic analysis. Microbiol Immunol 39:185–192
Author information
Authors and Affiliations
Additional information
Received: 15 February 1996 / Accepted: 28 June 1996
Rights and permissions
About this article
Cite this article
Yamamoto, A., Hashimoto, T., Asaga, E. et al. Phylogenetic Position of the Mitochondrion-Lacking Protozoan Trichomonas tenax, Based on Amino Acid Sequences of Elongation Factors 1α and 2. J Mol Evol 44, 98–105 (1997). https://doi.org/10.1007/PL00006127
Published:
Issue Date:
DOI: https://doi.org/10.1007/PL00006127