Abstract
Background
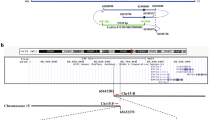
Mutations in the mouse formin (Fmn) gene result in limb deformities and incompletely penetrant renal aplasia. A molecular genetic approach was taken to characterize novel circular RNAs from the Fmn gene and to understand the developmental effects of gene-targeted mutations.
Materials and Methods
RT-PCR and ribonuclease protection analyses were done to characterize the circular RNA transcripts arising from the Fmn gene. Two lines of mice with gene-targeted deletions of specific Fmn exons, namely exon 4 or exon 5, were generated and analyzed.
Results
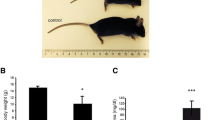
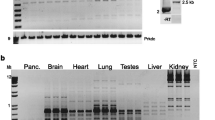
In our analysis of formin cDNAs, we discovered a class of transcripts in which the exon order is reversed such that downstream exons are joined to the acceptor end of a specific exon that lies 5′ to them in the genome. RT-PCR and ribonuclease protection analyses indicate that these transcripts are circular and are the major transcripts arising from this locus in adult brain and kidney. To gain insight into the biological function of these transcripts, we have systematically deleted the relevant exons using gene-targeted homologous recombination. The resulting mice fail to produce circular transcripts, but appear to produce normal amounts of the linear RNA isoforms from this locus. While these deficient mice have normal limbs, they display variably penetrant renal aplasia characteristic of other mutant formin alleles.
Conclusions
Our results demonstrate novel circular transcripts arising from the Fmn gene. Moreover, their high levels of expression suggest that they are not products of aberrant splicing events, but instead, may play important biological roles. Mice with gene-targeted deletions of Fmn exons 4 or 5 lack these circular transcripts and have an incompletely penetrant renal agenesis phenotype. While the biologic function of circular Fmn RNA transcripts is not entirely known, our work suggests their possible involvement in kidney development.







Similar content being viewed by others
References
Wang CC, Chan DC, Leder P (1997) The mouse formin (Fmn) gene: genomic structure, novel exons, and genetic mapping. Genomics 39: 303–311.
Woychik RP, Maas RL, Zeller R, Vogt TF, Leder P (1990) ‘Formins’: proteins deduced from the alternative transcripts of the limb deformity gene. Nature 346: 850–853.
Cupp MB (1960) Limb-deformity, ld. Mouse News Lett. 22: 50.
Green MC, Masiak C (1962) Limb deformity. Mouse News Lett. 26: 34–35.
Rhodes R (1994) Remutation at limb deformity locus. Mouse Genome 92: 355.
Woychik RP, Stewart TA, Davis LG, D’Eustachio P, Leder P (1985) An inherited limb deformity created by insertional mutagenesis in a transgenic mouse. Nature 318: 36–40.
Woychik RP, Generoso WM, Russel LB, et al. (1990) Molecular and genetic characterization of a radiation-induced structural rearrangement in mouse chromosome 2 causing mutations at the limb deformity and agouti loci. Proc. Natl. Acad. Sci. U.S.A. 87: 2588–2592.
Messing A, Behringer RR, Slapak JR, Lemke G, Palmiter RD, Brinster RL (1990) Insertional mutation at the ld locus (again!) in a line of transgenic mice. Mouse Genome 1990: 107.
Zeller R, Jackson-Grusby L, Leder P (1989) The limb deformity gene is required for apical ectodermal ridge differentiation and anteroposterior limb pattern formation. Genes Dev. 3: 1481–1492.
Chan DC, Wynshaw-Boris A, Leder P (1995) Formin isoforms are differentially expressed in the mouse embryo and are required for normal expression of fgf-4 and shh in the limb bud. Development 121: 3151–3162.
Haramis AG, Brown JM, Zeller R (1995) The limb deformity mutation disrupts the SHH/FGF-4 feedback loop and regulation of 5′ HoxD genes during limb pattern formation. Development 121: 4237–4245.
Maas R, Elfering S, Glaser T, Jepeal L (1994) Deficient outgrowth of the ureteric bud underlies the renal agenesis phenotype in mice manifesting the limb deformity (ld) mutation. Dev. Dyn. 199: 214–228.
Wynshaw-Boris A, Ryan G, Deng C-X, et al. (1997) The role of a single formin isoform in the limb and renal phenotypes of limb deformity. Mol. Med. 3: 372–384.
Nigro JM, Cho KR, Fearon ER, et al. (1991) Scrambled exons. Cell 64: 607–613.
Cocquerelle C, Daubersies P, Majérus M-A, Kerckaert J-P, Bailleul B (1992) Splicing with inverted order of exons occurs proximal to large introns. EMBO J. 11: 1095–1098.
Cocquerelle C, Mascrez B, Hétuin D, Bailleul B (1993) Mis-splicing yields circular RNA molecules. FASEB J. 7: 155–160.
Bailleul B (1996) During in vivo maturation of eukaryotic nuclear mRNA, splicing yields excised exon circles. Nucl Acids Res. 24: 1015–1019.
Zaphiropoulos PG (1996) Circular RNAs from transcripts of the rat cytochrome P450 2C24 gene: correlation with exon skipping. Proc. Natl Acad. Sci. U.S.A. 93: 6536–6541.
Capel B, Swain A, Nicolis S, et al. (1993) Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73: 1019–1030.
Jackson-Grusby L, Kuo A, Leder P (1992) A variant limb deformity transcript expressed in the embryonic mouse limb defines a novel formin. Genes Dev. 6: 29–37.
Sambrook J, Fritsch EF, Maniatis T (eds). (1989) Molecular Cloning: A Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Melton DA, Krieg PA, Rebagliati MR, Maniatis T, Zinn K, Green MR (1984) Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucl Acids Res. 12: 7035–7056.
Tybulewicz VL, Crawford CE, Jackson PK, Bronson RT, Mulligan RC (1991) Neonatal lethality and lymphopenia in mice with a homozygous disruption of the c-abl proto-oncogene. Cell 65: 1153–1163.
Deng C, Wynshaw-Boris A, Zhou F, Kuo A, Leder P (1996) Fibroblast growth factor receptor 3 is a negative regulator of bone growth. Cell 84: 911–921.
Deng C-X, Wynshaw-Boris A, Shen MM, Daugherty C, Ornitz DM, Leder P (1994) Murine FGFR-1 is required for early postimplantation growth and axial organization. Genes Dev. 8: 3045–3057.
Deng C, Zhang P, Harper JW, Elledge SJ, Leder P (1995) Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell 82: 675–684.
Chan DC, Leder P (1996) Genetic evidence that formins function within the nucleus. J. Biol Chem. 271: 23472–23477.
Harlow E, Lane D (1988) Antibodies: A Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Acknowledgments
We thank Mark Bedford and Chuxia Deng for their technical expertise and helpful discussions. We gratefully acknowledge technical help from Cathie Daugherty, Judy Dunmore, Anne Harrington, and Montserrat Michelman. We are grateful to Hsiuchen Chen, Ian Krane, Ben Stanger, and Christoph Westphal for careful reading of the manuscript. C. W. Chao and D. C. Chan were supported in part by the NIH Medical Scientist Training Program.
Author information
Authors and Affiliations
Additional information
Communicated by P. Leder.
Rights and permissions
About this article
Cite this article
Chao, C.W., Chan, D.C., Kuo, A. et al. The Mouse formin (Fmn) Gene: Abundant Circular RNA Transcripts and Gene-Targeted Deletion Analysis. Mol Med 4, 614–628 (1998). https://doi.org/10.1007/BF03401761
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03401761