Abstract
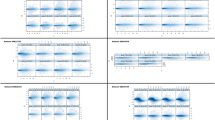
Background. The development and progression of cancer are accompanied by complex changes in patterns of gene expression. The purpose of this study was to clarify the relevance of macroarray analysis of human colorectal cancer tissues. Methods. Hybridization of cDNA macroarray filters on which 550 genes had been spotted was performed with biotin-labeled cDNA targets that were prepared from mRNA extracted from 20 pairs of colorectal cancer and corresponding noncancerous tissues. Expression of differentially expressed genes was further studied by semiquantitative RT-PCR. Results. Fourteen (2.5%) of the 550 genes were differentially expressed and up- or downregulated in cancer tissues by at least threefold compared with matched noncancerous tissues in 10 or more of the 20 patients. The genes that were upregulated in cancer tissues were associated with transcription, cell cycle, growth factor receptor, cell adhesion, extracellular matrix-degrading enzymes, and angiogenesis, and the downregulated genes were those involved in apoptosis and immune recognition. Semiquantitative RT-PCR analysis of these differentially expressed genes gave results consistent with those by cDNA array analysis. Conclusions. Although the macroarray used in this study contained only a small number of genes, our results support the feasibility and usefulness of this approach to study variation in gene expression patterns in human colorectal cancer tissues. The results also suggest the possibility of a diagnostic application of cDNA macroarrays in daily clinical settings.
Similar content being viewed by others
References
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 1999;286:531–7.
van Berkum NL, Holstege FC. DNA microarrays: raising the profile. Curr Opin Biotechnol 2001;12:48–52.
Su AI, Welsh JB, Sapinoso LM, Kern SG, Dimitrov P, Lapp H, et al. Molecular classification of human carcinomas by use of gene expression signatures. Cancer Res 2001;61:7388–93.
Lu J, Liu Z, Xiong M, Wang Q, Wang X, Yang G, et al. Gene expression profile changes in initiation and progression of squamous cell carcinoma of esophagus. Int J Cancer 2001;91:288–94.
Kan T, Shimada Y, Sato F, Maeda M, Kawabe A, Kaganoi J, et al. Gene expression profiling in human esophageal cancers using cDNA microarray. Biochem Biophys Res Commun 2001;286: 792–801.
Hippo Y, Taniguchi H, Tsutsumi S, Machida N, Chong JM, Fukayama M, et al. Global gene expression analysis of gastric cancer by oligonucleotide microarrays. Cancer Res 2002;62:233–40.
Alon U, Barkai N, Notterman DA, Gish K, Ybarra S, Mack D, et al. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sei USA 1999;96:6745–50.
Okuno K, Yasutomi M, Nishimura N, Arakawa T, Shiomi M, Hida J, et al. Gene expression analysis in colorectal cancer using practical DNA array filter. Dis Colon Rectum 2001;44:295–9.
Notterman DA, Alon U, Sierk AJ, Levine AJ. Transcriptional gene expression profiles of colorectal adenoma, adenocarcinoma, and normal tissue examined by oligonucleotide arrays. Cancer Res 2001;61:3124–30.
Kitahara O, Furukawa Y, Tanaka T, Kihara C, Ono K, Yanagawa R, et al. Alterations of gene expression during colorectal carcinogenesis revealed by cDNA microarrays after laser-capture microdissection of tumor tissues and normal epithelia. Cancer Res 2001;61:3544–9.
Zhang H, Yu CY, Singer B, Xiong M. Recursive partitioning for tumor classification with gene expression microarray data. Proc Natl Acad Sei USA 2001;98:6730–5.
Takemasa I, Higuchi H, Yamamoto H, Sekimoto M, Tomita N, Nakamori S, et al. Construction of preferential cDNA microarray specialized for human colorectal carcinoma: molecular sketch of colorectal cancer. Biochem Biophys Res Commun 2001;285:1244–9.
Okabe H, Satoh S, Kato T, Kitahara O, Yanagawa R, Yamaoka Y, et al. Genome-wide analysis of gene expression in human hepato- cellular carcinomas using cDNA microarray identification of genes involved in viral carcinogenesis and tumor progression. Cancer Res 2001;61:2129–37.
Shirota Y, Kaneko S, Honda M, Kawai HF, Kobayashi K. Identification of differentially expressed genes in hepatocellular carcinoma with cDNA microarrays. Hepatology 2001;33:832–40.
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature (Lond) 2000;403:503–ll.
Matrisian LM, Cunha GR, Mohla S. Epithelial-stromal interactions and tumor progression: meeting summary and future directions. Cancer Res 2001;61:3844–6.
Arias AM. Epithelial mesenchymal interactions in cancer and development. Cell 2001;105:425–31.
Chaib H, Cockrell EK, Rubin MA, Macoska JA. Profiling and verification of gene expression patterns in normal and malignant human prostate tissues by cDNA microarray analysis. Neoplasia 2001;3:43–52.
Fukushima H, Yamamoto H, Itoh F, Nakamura H, Min Y, Horiuchi S, et al. Association of matrilysin mRNA expression with K-ras mutations and progression in pancreatic ductal adeno- carcinomas. Carcinogenesis (Oxf) 2001;22:1049–52.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Yamamoto, H., Imsumran, A., Fukushima, H. et al. Analysis of gene expression in human colorectal cancer tissues by cDNA array. J Gastroenterol 37 (Suppl 14), 83–86 (2002). https://doi.org/10.1007/BF03326421
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03326421