Abstract
Background: Quantitation of human papillomavirus (HPV) DNA in clinical samples may yield important clinical information.
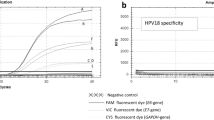
Methods and Results: We developed a 5′ exonuclease fluorescent probe assay for HPV quantitation that uses real-time PCR. The assay was optimized for HPV types 6 (HPV-6), -11, -16, and -18. A multiplex format was developed to quantify a cellular target of known iteration simultaneously with HPV quantitation, which controls for the amount of input DNA. Dilution series of target and heterologous templates were used to verify the assay. The assay was successfully used on fresh and PreservCyt-fixed cell lines, as well as cervical samples. The linear range of the assay is from 10 to 10 million copies. Intraclass correlations for HPV, actin, and globin assays ranged from 0.95 to 0.99, indicating the analytic precision of repeated measures.
Conclusion: The method is accurate over a large copy number range, reproducible, type specific, normalized for input DNA quantity, and applicable to PreservCytfixed material.
Similar content being viewed by others
References
Josefsson AM, Magnusson PK, Ylitalo N, et al.: Viral load of human papilloma virus 16 as a determinant for development of cervical carcinoma in situ: A nested case-control study. Lancet 2000;355:2189–2193
Swan DC, Tucker RA, Tortolero-Luna G, et al.: Human papillomavirus (HPV) DNA copy number is dependent on grade of cervical disease and HPV type. J Clin Microbiol 1999;37:1030–1034
Ylitalo N, Sorensen P, Josefsson AM, et al.: Consistent high viral load of human papillomavirus 16 and risk of cervical carcinoma in situ: A nested casecontrol study. Lancet 2000;355:2194–2198
Jacobs MV, Walboomers JM, van Beek J, et al.: A quantitative polymerase chain reaction-enzyme immunoassay for accurate measurements of human papillomavirus type 16 DNA levels in cervical scrapings. Br J Cancer 1999;81:114–121
Josefsson A, Livak K, Gyllensten U: Detection and quantitation of human papillomavirus by using the fluorescent 5′ exonuclease assay. J Clin Microbiol 1999;37:490–96
Serth J, Panitz F, Herrmann H, Alves J: Single-tube nested competitive PCR with homologous competitor for quantitation of DNA target sequences: Theoretical description of heteroduplex formation, evaluation of sensitivity, precision and linear range of the method. Nucleic Acids Res 1998;26:4401–4408
Swan DC, Tucker RA, Holloway BP, Icenogle JP: A sensitive, type-specific, fluorogenic probe assay for detection of human papillomavirus DNA. J Clin Microbiol 1997;35:886–891
Livak KJ, Flood SJ, Marmaro J, Giusti W, Deetz K: Oligonucleotides with fluorescent dyes at opposite ends provide a quenched probe system useful for detecting PCR product and nucleic acid hybridization. PCR Methods Appl 1995;4:357–362
Förster VT: Zwischenmolekkulare Energiewanderung and Fluorezenz. Ann Phys 1948;2:55–75
Holland PM, Abramson RD, Watson R, Gelfand DH: Detection of specific polymerase chain reaction product by utilizing the 5′-3′ exonuclease activity of Thermus aquaticus DNA polymerase. Proc NatlAcad Sci U S A 1991;88:7276–7280
Heid CA, Stevens J, Livak KJ, Williams PM: Real time quantitative PCR. Genome Res 1996;6:986–994
Lie YS, Petropoulos CJ: Advances in quantitative PCR technology: 5’ Nuclease assays. Curr Opin Biotechnol 1998;9:43–8
Orlando C, Pinzani P, Pazzagli M: Developments in quantitative PCR. Clin Chem Lab Med 1998;36:255–269
Higuchi R, Fockler C, Dollinger G, Watson R: Kinetic PCR analysis: Real-time monitoring of DNA amplification reactions. Biotechnology (NY) 1993; 11:1026–1030
Fleiss J: The design and analysis of clinical experiments. Wiley, New York, 1986
Lin LI: A concordance correlation coefficient to evaluate reproducibility. Biometrics 1989;45:255–268
Cross NC: Quantitative PCR techniques and applications. Br J Haematol 1995,89:693–697
Diaco R: Practical considerations for the design of quantitative PCR assays. In Innis MA, Gelfand DH, Sninsky JJ: PCR strategies. Academic, San Diego, 1995, pp 84–108
Leavitt J, Gunning P, Porreca P, Ng SY, Lin CS, Kedes L: Molecular cloning and characterization of mutant and wild-type human beta-actin genes. Mol Cell Biol 1984;4:1961–1969
Delassus S: Quantification of cytokine transcripts using polymerase chain reaction. Eur Cytokine Netw 1997;8:239–244
Kreuzer KA, Lass U, Landt O, et al.: Highly sensitive and specific fluorescence reverse transcription-PCR assay for the pseudogene-free detection of beta-actin transcripts as quantitative reference. Clin Chem 1999;45:297–300
Raff T, van der Giet M, Endemann D, Wiederholt T, Paul M: Design and testing of beta-actin primers for RT-PCR that do not co-amplify processed pseudogenes. Biotechniques 1997;23:456–460
Wettstein FO, Sninsky JJ: Nucleic acid hybridization and unconventional bases. In Innis MA, Gelfand DH, Sninsky JJ: PCR Strategies. Academic, San Diego, 1995, pp 69–83
Berumen J, Casas L, Segura E, Amezcua JL, Garcia-Carranca A: Genome amplification of human papillomavirus types 16 and 18 in cervical carcinomas is related to the retention of E1/E2 genes. Int J Cancer 1994;56:640–645
Kalantari M, Karlsen F, Kristensen G, Holm R, Hagmar B, Johansson B: Disruption of the E1 and E2 reading frames of HPV 16 in cervical carcinoma is associated with poor prognosis. Int J Gynecol Pathol 1998;17:146–153
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tucker, R.A., Unger, E.R., Holloway, B.P. et al. Real-time PCR-based Fluorescent Assay for Quantitation of Human Papillomavirus Types 6, 11, 16, and 18. Molecular Diagnosis 6, 39–47 (2001). https://doi.org/10.1007/BF03262102
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03262102