Abstract
Background: In fluorescence in situ hybridization (FISH) applications, the efficiency of probe hybridization is greatly enhanced by treating the cell or tissue preparation with a variety of reagents that make the target permeable while preserving important morphological features. Pretreatment protocols can be very labor intensive, adding cost to the test. The automation of specimen pretreatment eliminates human variation in the procedure through standardization of such variables as treatment times and temperatures.
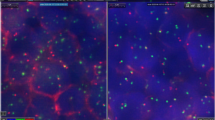
Methods and Results: We developed an instrument, the VP 2000 Processor, that can process up to 50 specimens simultaneously under computer control. In this comparative FISH study of matched specimens, one set was processed according to the manual pretreatment protocol and compared with specimens processed by the VP 2000. Processed specimen types included paraffin-embedded breast tissue, uncultured amniocytes, bone marrow, peripheral-blood lymphocytes, and uroepithelial cells recovered from urine. Data show equivalent or brighter and more specific overall signal quality compared with matched manually executed controls.
Conclusions: The automation of sample pretreatment for FISH provides a superior, more consistent level of performance than the manual format.
Similar content being viewed by others
References
Hopman AHN, Poddighe P, Moesker O, Ramaekers FCS: Interphase cytogenetics: An approach to the detection of genetic aberrations in tumors. In McGee JO, Herrington CS: Diagnostic molecular pathology: A practical approach. New York, NY, IRL Press, 1991, pp. 141-167
Hopman AH, Voorter CE, Ramaekers FC: Detection of genomic changes in cancer by in situ hybridization. Mol Biol Rep 1994;19:31–44
Pinkel D, Straume T, Gray JW: Cytogenetic analysis using quantitative, high sensitivity fluorescence hybridization. Proc Natl Acad Sci U S A 1986;83: 2934–2938
Hyytinen E, Visakorpi T, Kallioniemi A, Kallioniemi OP, Isola JJ: Improved technique for analysis of formalin-fixed, paraffin-embedded tumors by fluorescence in situ hybridization. Cytometry 1994;16: 93–99
Kapranos N, Kontogergos G, Frangia K, Kokka E: Effect of fixation on interphase cytogenetic analysis by direct fluorescence in situ hybridization on cell imprints. Biotechnol Histochem 1997;72:148–151
Persons DL, Bui MM, Lowery MC, et al.: Fluorescence in situ hybridization (FISH) for detection of Her-2/neu amplification in breast cancer: A multicenter portability study. Ann Clin Lab Sci 2000;1: 41–48
Abati A, Sanford JS, Fetsch P, Marincola FM, Wolman SR: Fluorescence in situ hybridization (FISH): A user’s guide to optimal preparation of cytologic specimens. Diagn Cytopathol 1995;13:486–492
Grogan TM: Automated immunohistochemical analysis. Am J Clin Pathol 1992;98(Suppl 1):S35–S38
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jacobson, K., Thompson, A., Browne, G. et al. Automation of Fluorescence In Situ Hybridization Pretreatment: A Comparative Study of Different Sample Types. Molecular Diagnosis 5, 209–220 (2000). https://doi.org/10.1007/BF03262078
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03262078