Abstract
Background: Currently, prostate cancer (CaP) cytogenetics is not well defined, largely because of technical difficulties in obtaining primary tumor metaphases.
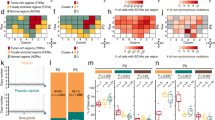
Methods and Results: We examined three CaP cell lines (LNCaP, DU145, PC-3) using sequential Giemsa banding and spectral karyotyping (SKY) to search for a common structural aberration or translocation breakpoint. No consistent rearrangement common to all three cell lines was detected. A clustering of centromeric translocation breakpoints was detected in chromosomes 4, 5, 6, 8, 11, 12, 14, and 15 in DU145 and PC-3. Both these lines were found to have karyotypes with a greater level of complexity than LNCaP.
Conclusion: The large number of structural aberrations present in DU145 and PC-3 implicate an underlying chromosomal instability and subsequent accumulation of cytogenetic alterations that confer a selective growth advantage. The high frequency of centromeric rearrangements in these lines indicates a potential role for mitotic irregularities associated with the centromere in CaP tumorigenesis.
Similar content being viewed by others
References
Haas GP, Sakr WA: Epidemiology of prostate cancer. CA Cancer J Clin 1997;47:273–287
Weinberg R: How cancer arises. Sci Am 1996;275: 62-70
Rowley JD: The critical role of chromosome translocations in human leukemias. Annu Rev Genet 1998;32: 495–519
Ladanyi M: The emerging molecular genetics of sarcoma translocations. Diagn Mol Pathol 1995;4: 162–173
Webb HD, Hawkins AL, Griffin CA: Cytogenetic abnormalities are frequent in uncultured prostate cancer cells. Cancer Genet Cytogenet 1996,88: 126–132
Konig JJ, Teubel W, van Dongen JW, Hagemeijer A, Romijn JC, Schroder FH: Tissue culture loss of aneuploid cells from carcinomas of the prostate. Genes Chromosomes Cancer 1993;8: 22–27
Ketter R, Zwergel T, Romanakis K, et al.: Selection toward diploid cells in prostatic carcinoma-derived cell cultures. Prostate 1996;28: 364–371
Chopra DP, Sarkar FH, Grignon DJ, Sakr WA, Mohammed A, Waghray A: Growth of human nondiploid primary prostate tumor epithelial cells in vitro. Cancer Res 1997;57: 3688–3692
Limon J, Lundgren R, Elfving P, et al.: An improved technique for short-term culturing of human prostatic adenocarcinoma tissue for cytogenetic analysis. Cancer Genet Cytogenet 1990;46: 191–199
Peehl DM, Sellers RG, Wong ST: Defined medium for normal adult human prostatic stromal cells. In Vitro Cell Dev Biol Anim 1998;34: 555–560
Szucs S, Zitzelsberger H, Breul J, Bauchinger M, Hofler H: Two-phase short-term culture method for cytogenetic investigations from human prostate carcinoma. Prostate 1994;25: 225–235
Zwergel T, Kakirman H, Schorr H, Wullich B, Unteregger G: A new serial transfer expiant cell culture system for human prostatic cancer tissues preventing selection toward diploid cells. Cancer Genet Cytogenet 1998;101: 16–23
Arps S, Rodewald A, Schmalenberger B, Carl P, Bressel M, Kastendieck H: Cytogenetic survey of 32 cancers of the prostate. Cancer Genet Cytogenet 1993;66: 93–99
Micale MA, Sanford JS, Powell IJ, Sakr WA, Wolman SR: Defining the extent and nature of cytogenetic events in prostatic adenocarcinoma: Paraffin FISH vs metaphase analysis. Cancer Genet Cytogenet 1993;69: 7–12
Zitzelsberger H, Szucs S, Robens E, Weier HU, Hofler H, Bauchinger M: Combined cytogenetic and molecular genetic analyses of fifty-nine untreated human prostate carcinomas. Cancer Genet Cytogenet 1996;90: 37–44
Horoszewicz JS, Leong SS, Kawinski E, et al.: LNCaP model of human prostatic carcinoma. Cancer Res 1983;43: 1809–1818
Gibas Z, Becher R, Kawinski E, Horoszewicz J, Sandberg AA: A high-resolution study of chromosome changes in a human prostatic carcinoma cell line (LNCaP). Cancer Genet Cytogenet 1984;11: 399-404
Stone KR, Mickey DD, Wunderli H, Mickey GH, Paulson DF: Isolation of a human prostate carcinoma cell line (DU 145). Int J Cancer 1978;21: 274–281
Kaighn ME, Narayan KS, Ohnuki Y, Lechner JF, Jones LW: Establishment and characterization of a human prostatic carcinoma cell line (PC-3). Invest Urol 1979;17: 16–23
Veronese ML, Bullrich F, Negrini M, Croce CM: The t(6;16)(p21;q22) chromosome translocation in the LNCaP prostate carcinoma cell line results in a tpc/hpr fusion gene. Cancer Res 1996;56: 728–732
Bernardino J, Bourgeois CA, Muleris M, Dutrillaux AM, Malfoy B, Dutrillaux B: Characterization of chromosome changes in two human prostatic carcinoma cell lines (PC-3 and DU145) using chromosome painting and comparative genomic hybridization. Cancer Genet Cytogenet 1997;96: 123–128
Schrock E, du Manoir S, Veldman T, et al.: Multicolor spectral karyotyping of human chromosomes. Science 1996;273: 494–497
Horoszewicz JS, Leong SS, Chu TM, et al.: The LNCaP cell line—A new model for studies on human prostatic carcinoma. Prog Clin Biol Res 1980; 37: 115–132
Gustashaw KM: Chromosome stains. In Barch MJ, Knutsen T, Spurbeck JL: The AGT cytogenetics laboratory manual, 3. Lippincott-Raven, Philadelphia, 1997, pp. 259-324
Veldman T, Vignon C, Schrock E, Rowley JD, Ried T: Hidden chromosome abnormalities in haematological malignancies detected by multicolour spectral karyotyping. Nat Genet 1997; 15: 406–410
Garini Y, Macville M, Manoir S, et al.: Spectral karyotyping. Bioimaging 1996;4: 65–72
Mitelman F: ISCN (1995): International system for human cytogenetic nomenclature. Karger, New York, 1995
Padilla-Nash HM, Nash WG, Padilla GM, et al.: Molecular cytogenetic analysis of the bladder carcinoma cell line BK-10 by spectral karyotyping. Genes Chromosomes Cancer 1999;25: 53–59
Allen RJ, Smith SD, Moldwin RL, et al.: Establishment and characterization of a megakaryoblast cell line with amplification of MLL.Leukemia 1998; 12: 1119-1127
Ford S, Gray IC, Spurr NK: Rearrangement of the long arm of chromosome 10 in the prostate adeno-carcinoma cell line LNCaP. Cancer Genet Cytogenet 1998;102: 6–11
Brothman AR, Maxwell TM, Cui J, Deubler DA, Zhu XL: Chromosomal clues to the development of prostate tumors. Prostate 1999;38: 303–312
Camby I, Etievant C, Petein M, et al.: Influence of culture media on the morphological differentiation of the PC-3 and DU145 prostatic neoplastic cell lines. Prostate 1994;24: 187–196
Nupponen NN, Hyytinen ER, Kallioniemi AH, Visakorpi T: Genetic alterations in prostate cancer cell lines detected by comparative genomic hybridization. Cancer Genet Cytogenet 1998;101: 53–57
Vig BK, Sternes KL, Paweletz N: Centromere structure and function in neoplasia. Cancer Genet Cytogenet 1989;43: 151–178
Broccoli D, Paweletz N, Vig BK: Sequence of centromere separation: Characterization of multicentric chromosomes in a rat cell line. Chromosoma 1989; 98: 13–22
Kaiser-Rogers K, Rao K: Structural chromosome rearrangements. In Gersen SL, Keagle MB: The principles of clinical cytogenetics, Humana Press, Totowa, 1999, pp. 191-228
Chan GK, Schaar BT, Yen TJ: Characterization of the kinetochore binding domain of CENP-E reveals interactions with the kinetochore proteins CENP-F and hBUBR 1. J Cell Biol 1998; 143: 49–63
Faulkner NE, Vig B, Echeverri CJ, Wordeman L, Vallee RB: Localization of motor-related proteins and associated complexes to active, but not inactive, centromeres. Hum Mol Genet 1998;7: 671–677
Cahill DP, da Costa LT, Carson-Walter EB, Kinzler KW, Vogelstein B, Lengauer C: Characterization of MAD2B and other mitotic spindle checkpoint genes. Genomics 1999;58: 181–187
Sawyer JR, Tricot G, Mattox S, Jagannath S, Barlogie B: Jumping translocations of chromosome 1q in multiple myeloma: Evidence for a mechanism involving decondensation of pericentromeric heterochromatin. Blood 1998;91: 1732–1741
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported in part by the US Army Medical Research and Materiel Command Prostate Cancer Research Program; and a grant from the American Foundation for Urologic Diseases/ American Urologic Association Research Scholar Program and Imclone Systems, Inc (P.C.P.)
Rights and permissions
About this article
Cite this article
Beheshti, B., Karaskova, J., Park, P.C. et al. Identification of a High Frequency of Chromosomal Rearrangements in the Centromeric Regions of Prostate Cancer Cell Lines by Sequential Giemsa Banding and Spectral Karyotyping. Molecular Diagnosis 5, 23–32 (2000). https://doi.org/10.1007/BF03262019
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03262019