Abstract
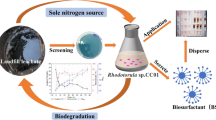
A biosurfactant-producing strain Z25 was isolated from the oil production water in Daqing Oilfield, China. The strain was identified asRhodococcus ruber by 16S rDNA sequencing.Rhodococcus ruber Z25 showed a preference of alkanes as carbon sources for biosurfactant synthesis. The optimal biosurfactant production was achieved at the NaCI concentration of 2.5% at 34 °C. In batch cultivation, R. ruber Z25 exhibited a cell-growth associated biosurfactant-pro ducing process onn-hexadecane and the maximum yield of biosurfactant production was 13.34 g/L at 44 h. The rate of biosurfactant production per unit biomass (R(biosurf/biomass)) was used to characterize the biosurfactant-producing profile. A two-stage biosurfactant-producing profile of R.ruber Z25 was established by the R(biosurf/biomass) curve, exhibiting the biosurfactant production in correlation with cellular hydrophobicity and biomass accumulation. The crude biosurfactant was extracted by methyltert-butyl ether (MTBE) method and only one glycolipid fraction at Rf value of 0.64 was detected by thin layer chromatography (TLC) analysis.
Similar content being viewed by others
References
Bell K.S., Philp J.C., Aw D.W., Christofi N. (1998). The genusRhodococcus. J. Appl. Microbiol. 85: 195–210.
Bicca F.C., Fleck L.C., Ayub M.A.Z. (1999). Production of bio-surfactant by hydrocarbon degradingRhodococcus ruber andRhodococcus erythropolis. Rev. Microbiol., 30: 231–236.
Bradford M.M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem., 72: 248–254.
Cooper D.G., Goldenberg B.G. (1987). Surface active agents fromBacillus species. Appl. Environ. Microbiol., 55: 224–229.
Desai J.D., Banat I.M. (1997). Microbial production of surfactants and their commercial potential. Microbiol. Mol. Biol., 61: 47–64.
Dubois M., Gilles K.A., Hamilton J.K., Rebers R.A., Smith F. (1956). Colorimetric method for the determination of sugars and related substances. Anal. Chem., 28: 350–353.
Fleck L.C., Bicca F.C., Ayub M.A.Z. (2000). Physiological aspects of hydrocarbon emulsification, metal resistance and DNA profile of biodegrading bacteria isolated from oil polluted sites. Biotechnol. Lett., 22: 285–289.
Ghurye G.L., Vipulanandan C. (1994). A practical approach to biosurfactant production using nonaseptic fermentation of mixed cultures. Biotechnol. Bioeng., 44: 661–666.
Ivshina I.B., Kuyukina M.S., Philp J.C., Christofi N. (1998). Oil desorption from mineral and organic materials using biosur factant complexes produced byRhodococcus species. World J. Microbiol. Biotechnol., 14: 711–717.
Kim J.S., Powalla M., Lang S., Wagner F., Lunsdorf H., Wray V. (1990). Microbial glycolipid production under nitrogen limita tion and resting cell condition. J. Bacterid., 13: 257–266.
Kitamoto D., Isoda H., Nakahara T. (2002). Functions and poten tial applications of glycolipid biosurfactants: from energy-saving materials to gene delivery carriers. J. Biosci. Bioeng., 94: 187–201.
Kuyukina M.S., Ivshina I., Philip J., Christofi N., Dunbar S., Ritchkova M. (2001). Recovery ofRhodococcus biosur factants using methyl tertiary-butyl ether extraction. J. Microbiol. 46: 149–156.
Lang S., Wullbrandt D. (1999). Rhamnose lipids-biosynthe-sis, microbial production and application potentials. Appl. Microbiol. Biotechnol., 51: 22–32.
Lin T.C., Young C.C., Ho M., Yeh M.S., Chou CL, Wei Y.H., Chang J.S. (2005). Characterization of floating activity of indigenous diesel-assimilating bacterial isolates. J. Biosci. Bioeng., 99: 466–472.
Mutalik S.R., Vaidya B.K., Joshi R.M., Desai K.M., Nene S. (2008). Use of response surface optimization for the produc tion of biosurfactant fromRhodococcus spp. MTCC 2574, Bioresource Tech., 99: 7875–7880.
Philp J.C., Kuyukina M.S., Ivshina I.B., Dunbar S.A., Christofi N., Lang S., Wray V. (2002). AlkanotrophicRhodococcus ruber as a biosurfactant producer. Appl. Microbiol. Biotechnol., 59: 318–324.
Pirog T.P., Shevchuk T.A., Voloshina I.N., Karpenko E.V. (2004). Production of surfactants byRhodococcus erythropolis strain EK-1, grown on hydrophilic and hydrophobic substrates. Appl. Biochem. Microbiol., 40: 470–475.
Pruthi V., Cameotra S.S. (1997). Rapid identification of biosur factant producing bacterial strains using a cell surface hydro-phobicity technique. Biotechnol. Tech., 11: 671–674.
Rahman K.S.M., Rahman T.J., Kourkoutas Y., Petsas I., Marchant R., Banat I.M. (2003). Enhanced bioremediation ofn-alkane in petroleum sludge using bacterial consortium amended with rhamnolipid and micronutrients. Bioresource Tech., 90: 159–168.
Rodrigues L., Teixeira J., Oliveira R., van der Mei H.C (2006). Response surface optimization of the medium components for the production of biosurfactants by hydrophobic bacteria. Process Biochem., 41: 1–10.
Sandrin T.R., Chech A.M., Maier R.M. (2000). A rhamnolipid biosurfactant reduces cadmium toxicity during naphtha lene biodegradation. Appl. Environ. Microbiol., 66: 4585–4588.
Wagner F., Behrendt U., Bock H., Kretschmer A., Lang S., Syldatk C (1983). Production and chemical characterisation of sur factants fromRhodococcus erythropolis andPseudomonas sp. MUB grown on hydrocarbons. In: Zajic J.E., Cooper D.G., Jack TR., Kosaric N., Eds, Microbial Enhanced Oil Recovery, Pennwell, Tulsa, Okla, pp. 55–60,
Wei Y.H., Lai H.C., Chen S.Y., Yeh M.S., Chang J.S. (2004). Biosurfactant production bySerratia marcescens SS-1 and its isogenic strain SMΔR defective in SpnR, a quorum-sensing LuxR family protein. Biotechnol. Lett., 26: 799–802.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zheng, C., Li, S., Yu, L. et al. Study of the biosurfactant-producing profile in a newly isolatedRhodococcus ruber strain. Ann. Microbiol. 59, 771–776 (2009). https://doi.org/10.1007/BF03179222
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03179222