Abstract
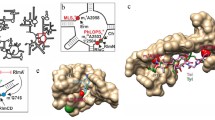
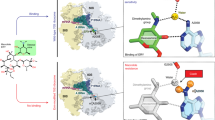
Erm methyltransferases mediate the resistance to the macrolide-lincosamide-streptogramin B antibioticsvia dimethylation of a specific adenine residue in 23S rRNA. The role of positively charged N-terminal residues of the ErmC′ methyltransferase in RNA binding and/or catalysis was determined. Mutational analysis of amino acids K4 and K7 was performed and the mutants were characterized inin vivo andin vitro experiments. The K4 and K7 residues were suggested not to be essential for the enzyme activity but to provide a considerable support for the catalytic step of the reaction, probably by maintaining the optimum conformation of the transition state through interactions with the phosphate backbone of RNA.
Access this article
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.
Similar content being viewed by others
References
Bujnicki J.M.: Comparison of protein structures reveals monophyletic origin of the AdoMet-dependent methyltransferase family and mechanistic convergence rather than recent differentiation of N4-cytosine and N6-adenine DNA methylation.In Silico Biol. 1, 175–182 (1999).
Bussiere D.E., Muchmore S.W., Dealwis C.G., Schluckebier G., Nienaber V.L., Edalji R.P., Walter K.A., Ladror U.S., Holzman T.F., Abad-Zapatero C.: Crystal structure of ErmC′, an rRNA methyltransferase which mediates antibiotic resistance in bacteria.Biochemistry 37, 7103–7112 (1998).
Cheng X., Roberts R.J.: AdoMet-dependent methylation, DNA methyltransferases and base flipping.Nucl.Acids Res. 29, 3784–3795 (2001).
Farrow K.A., Lyras D., Polekhina G., Koutsis K., Parker M.W., Rood J.I.: Identification of essential residues in the Erm(B) rRNA methyltransferase ofClostridium perfringens.Antimicrob.Agents Chemother. 46, 1253–1261 (2002).
Fauman E.B., Blumenthal R.M., Cheng X.: Structure and evolution of AdoMet-dependent methyltransferases, pp. 1–38 in X. Cheng, R.M. Blumenthal (Eds):S-Adenosylmethionine Dependent Methyltransferases: Structures and Function. World Scientific, Singapore 1999.
Monod M., Denoya C., Dubnau D.: Sequence and properties of pIM13, a macrolide-lincosamide-streptogramin B resistance plasmid fromBacillus subtilis.J.Bacteriol. 167, 138–147 (1986).
Schluckebier G., Zhong P., Stewart K.D., Kavanaugh T.J., Abad-Zapatero C.: The 2.2Å structure of the rRNA methyltransferase ErmC′ and its complexes with cofactor and cofactor analogs: implications for the reaction mechanism.J.Mol.Biol. 289, 277–291 (1999).
Schlunzen F., Zarivach R., Harms J., Bashan A., Tocilj A., Albrecht R., Yonath A., Franceschi F.: Structural basis for the interaction of antibiotics with the peptidyltransferase centre in eubacteria.Nature 413, 814–821 (2001).
Su S. L., Dubnau D.: Binding ofBacillus subtilis ErmC′ methyltransferase to 23S rRNA.Biochemistry 29, 6033–6042 (1990).
Weisblum B.: Erythromycin resistance by ribosome modification.Antimicrob.Agents Chemother. 39, 577–585 (1995).
Yu L.A., Petros M., Schnuchel A., Zhong P., Severin J.M., Walter K., Holzman T.F., Fesik S.W.: Solution structure of an rRNA methyltransferase (ErmAM) that confers macrolide-lincosamide-streptogramin antibiotic resistance.Nature Struct.Biol. 4, 483–489 (1997).
Zhong P., Pratt S.D., Edalji R.P., Walter K.A., Holzman T.F., Shivakumar A.G., Katz L.: Substrate requirements for ErmC′ methyltransferase activity.J.Bacteriol. 177, 4327–4332 (1995).
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by theMinistry of Science of Republic Croatia (no. 006320 and 006611). The second author is theEuropean Molecular Biology Organization andHoward Hughes Medical Institute young investigator and theFoundation for Polish Science fellow.
Rights and permissions
About this article
Cite this article
Maravić, G., Bujnicki, J.M. & Flögel, M. Mutational analysis of basic residues in the N-terminus of the rRNA:m6A methyltransferase ErmC′. Folia Microbiol 49, 3–7 (2004). https://doi.org/10.1007/BF02931637
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02931637