Abstract
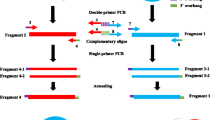
We report a simple and efficient method, which combines restriction endonuclease digestion and deoxynucleotide tailing, for cloning unknown genomic sequences adjacent to a known sequence. Total genomic DNA is partially digested with the frequent-cutting restriction enzymeNla III. A homo-oligomeric cytosine tail is added by terminal transferase. The tailed DNA fragments are used as the template for cloning flanking regions from all sequences of interest. A first round PCR amplification is performed with a gene-specific primer and the selective (modified polyguanine) anchor primer complementary to the cytosine tail and theNla III recognition site, with a universal amplification primer sequence at its 5′ end. This is followed by another PCR amplification with a nested gene-specific primer and the universal amplification primer. Finally, the amplified products are fractionated, cloned, and sequenced. Using this method, we cloned the upstream region of a salt-induced gene based upon a partial cDNA clone (RSC5-U) obtained from sunflower (Helianthus annuus L.).
Similar content being viewed by others
Abbreviations
- GSP:
-
gene specific primer
- RAGE:
-
rapid amplification of genomic ends
- RE:
-
restriction enzyme
- SAP:
-
selective anchor primer
- TdT:
-
terminal deoxynucleotide-transferase
- UAP:
-
universal amplification primer
References
Espelund M and Jakobsen KS (1992) Cloning and direct sequencing of plant promoters using primer-adaptor mediated PCR on DNA coupled to a magnetic solid phase. Bio Techniques 13: 74–81.
Frijters A, Pot J, Peleman J, Kuiper M and Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23: 4407–4414.
Iwahana H and Itakura M (1997) An end trimming method and its application to amplify adjacent cDNA and genomic DNA fragments by PCR. Methods Mol Biol 67: 247–260.
Kilstrup M and Kristiansen KN (2000) Rapid genome walking: a simplified oligo-cassette mediated polymerase chain reaction using a single genome-specific primer. Nucleic Acids Res 28: e55.
Moynihan TP, Markham AF and Robinson PA (1996) Genomic analysis of human multigene families using chromosome-specific Vectorette PCR. Nucleic Acids Res 24: 4094–4095.
Ochman H, Gerber AS and Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120: 621–623.
Qiang BQ and Schildkraut I (1986) Two unique restriction endonucleases fromNeisseria lactamica. Nucleic Acids Res 14: 1991–1999.
Parker JD, Rabinovitch PS and Burmer GC (1991) Targeted gene walking polymerase chain reaction. Nucleic Acids Res 19: 3055–3060.
Riley J, Butler R, Ogilvie D, Finniear R, Powell S, Anand R, Smith JC and Markham AF (1990) A novel, rapid method for the isolation of terminal sequences from yeast artificial chromosome (YAC) clones. Nucleic Acids Res 18: 2887–2890.
Rosenthal A and Jones DSC (1990) Genomic walking and sequencing by oligo-cassette mediated polymerase chain reaction. Nucleic Acids Res 18: 3095–3096.
Rudi K, Fossheim T and Jakobsen KS (1999) Restriction cutting independent method for cloning genomic DNA, segments outside the boundaries of known sequences. Bio Techniques 27: 1170–1174.
Sambrook J, Fritch EF and Maniatis T (1989) Molecular Cloning: A Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, X., Vance Baird, W. Rapid amplification of genomic DNA ends bynla III partial digestion and polynucleotide tailing. Plant Mol Biol Rep 19, 261–267 (2001). https://doi.org/10.1007/BF02772898
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF02772898