Abstract
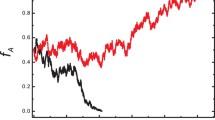
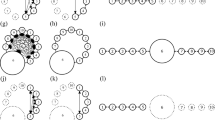
An individual-based simulation model was used to examine the effect of population subdivision, dispersal distance of offspring, and migration rates between subpopulations on genetic variability(H 1 H S andH T ) in a continuously distributed population. Some difficulties with mathematical models of a continuously distributed population have been pointed out. The individual-based model can avoid these difficulties and can be used to examine genetic variability in a population within which individuals are distributed continuously and in which the dispersal of individuals is disturbed by geographical or artificial barriers. The present simulation showed that the pattern of decrease inH 1 had three stages. During the first stage,H 1 decreased at the rates predicted by Wright’s neighborhood size. During the second stage,H 1 decreased more rapidly when the migration rate decreased, while during the third stage, it decreased less rapidly when the migration rate decreased. Increasing the number of subdivisions increased the rate of decrease after the 200th generation. The pattern of decrease inH T was classified into 2 stages. During the first stage, the rates of decrease corresponded with those of a randomly mating population. During the second stage, a decrease in the migration rates of the subpopulations slowed the rate of decrease inH T . A uniform spatial distribution and a reduced total dispersal distance of offspring causedH 1 H S , andH T to decrease more rapidly. Habitat fragmentation in a continuously distributed population usually was detrimental to the genetic variability in the early generations. Other implications of the results for conservation are discussed.
Similar content being viewed by others
References
Allendorf, F. W. (1983) Isolation, gene flow, and genetic differentiation among populations, pp. 51–65. In S. M. Schonewald-Cox, M. Chamberss, B. Macbryde, and L. Thomas (eds.)Genetics and conservation. Benjamin/Cummings, Menlo Park.
Chambers, S. M. (1983) Genetic principles for managers, pp. 15–146. In S. M. Schonewald-Cox, M. Chamberss, B. Macbryde and L. Thomas (eds.)Genetics and conservation. Benjamin/Cummings, Menlo Park.
Chambers, S. M. (1995) Spatial structure, genetic variation, and the neighborhood adjustment to effective population size.Conservation Biology 9: 1312–1315.
Chesser, R. K. (1983) Genetic principles for managers, pp. 66–77. In S. M. Schonewald-Cox, M. Chamberss, B. Macbryde and L. Thomas (eds.)Genetics and conservation. Benjamin/Cummings, Menlo Park.
Crawford, T. J. (1984) The estimation of neighborhood parameters in plant populations.Heredity 52: 272–283.
Felsenstein, J. (1975) A pain in the torus: some difficulties with models of isolation by distance.American Naturalist 109: 359–368.
Felsenstein, J. (1976) The theoretical population genetics of variable selection and migration.Annual Review of Genetics 10: 235–280.
Franklin, I. R. (1980) Evolutionary change in small populations, pp. 135–149. In S. M. Wilcox and B. Wilcoxs (eds.)Conservation biology: an evolutionary-ecological perspective. Sinauer Associates, MA, USA.
Harris, R. B. and F. W. Allendorf (1989) Genetically effective population size of large mammals: an assessment of estimators.Conservation Biology 3: 181–191.
Hedrick, P. W. and M. E. Gilpin (1997) Genetic effective size of metapopulation. pp. 165–181. In I.A. Hanski and M.E. Gilpin (eds.)Metapopulation biology: ecology, genetics, and evolution. Academic Press, California.
Kawata, M. (1995) Effective population size in a continuously distributed population.Evolution 49: 1046–1054.
Kimura, M. and T. Maruyama (1971) Patterns of neutral polymorphism in a geographically structured population.Genetical Research 18: 125–131.
Kimura, M. and G. Weiss (1964) The stepping stone model of population structure and the decrease of genetic correlation with distance.Genetics 49: 561–576.
Lacy, R. C. (1987) Loss of genetic diversity from managed populations: Interacting effects of drift, mutation, immigration, selection, and population subdivision.Conservation Biology 1: 143–158.
Levin, D. A. (1987) Local differentiation and the breeding structure of plant populations, pp. 305–325. In L. D. Gottlieb and K. Jain (eds.)Evolutionary biology. Chapman and Hall, London.
Malécot, G. (1950) Some probabilistic schemes for the variability of natural population.University of Lyon Science Section A 13: 37–60.
Maruyama, T. (1970) Stepping stone models of finite length.Advanced in Applied Probability 2: 229–258.
Maruyama, T. (1971) The rate of decrease of heterozygosity in a population occupying a circular or a linear habitat.Genetics 67: 437–454.
Nei, M. (1973) Analysis of gene diversity in subdivided populations.Proceedings of the National Academy of Sciences of the USA 70: 3321–3323.
Shield, W. M. (1982)Philoparty, inbreeding, and the evolution of sex.State University of New York Press, Albany.
Slatkin, M. (1985) Gene flow in natural populations.Annual Review of Ecology and Systematics 16: 393–430.
Slatkin, M. (1987) Gene flow and the geographic structure of natural populations.Science 236: 787–792.
Varvio, S.-L., R. Chakraborty and M. Nei (1985) Genetic variation in subdivided populations and conservation genetics.Heredity 57: 189–198.
Wright, S. (1943) Isolation by distance.Genetics 28: 139–156.
Wright, S. (1946) Isolation by distance under diverse systems of mating.Genetics 31: 39–59.
Wright, S. (1951) The genetical structure of populations.Annals of Eugenics 15: 323–354.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kawata, M. Loss of genetic variability in a fragmented continuously distributed population. Res Popul Ecol 39, 227–237 (1997). https://doi.org/10.1007/BF02765269
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02765269