Abstract
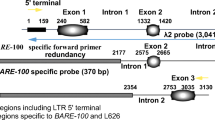
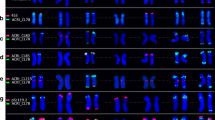
This paper describes a detailed sequence analysis of the ω-secalin gene array at theSec-1 locus on the short arm of chromosome 1 of rye. The analysis shows that the genes are separated by 8 kb of spacer sequence and that the gene/spacer units are arranged in a head to tail fashion. The boundaries of the array are identified, and a fragment containing the majority of the genes in the array is separated by PFG analysis. The sequence data of one 9.2 kb gene unit have been determined, and because of the similarity of the gene units within the array these data provide a detailed sequence analysis of 140 kb of theSec-1 locus. Fluorescence in situ hybridization, using lambda clones isolated for the structural analysis, identifies the position of the array on the rye chromosomes relative to the 5S rRNA genes.
Similar content being viewed by others

References
Appels R, Moran LB (1984) Molecular analysis of alien chromatin introduced into wheat. Stadler Symposium, pp 529–557
Baum M, Appels R (1991) The cytogenetic and molecular architecture of chromosome IR- one of the most widely utilized sources of alien chromatin in wheat varieties. Chromosoma 101:1–10
Carrillo JM, Vazquez JF, Orellana J (1990) Linkage relationships between the lociSec-1 andSec-3 in rye (Secale cereale L.). Heredity 64:125–130
Dhaliwal AS, MacRitchie F (1990) Contributions of protein fractions to dough handling properties of wheat-rye translocation cultivars. J Cereal Sci 12:113–122
Hull GA, Halford NG, Kreis M, Shewry PR (1991) Isolation and characterisation of genes encoding rye prolamins containing a highly repetitive sequence motif. Plant Molec Biol 17:1111–1115
Huynh TV, Young RA, Davis RW (1985) Construction and screening cDNA libraries in λgt10 and λgt11. In: Glover D (ed) DNA cloning: a practical approach. IRL Press, Oxford, pp 48–78
Koebner RMD, Shepherd KW (1986) Controlled introgression to wheat of genes from rye chromosome 1RS by induction of allosyndesis. 1. Isolation of recombinants. Theor Appl Genet 73:197–208
Kreis M, Ford BG, Rahman S, Miflin BJ, Shewry PR (1985) Molecular evolution of the seed storage proteins of barley, rye and wheat. J Molec Biol 183:499–502
Lawrence GT, Appels R (1984) Mapping the nucleos organiser region, seed protein and isozyme loci on chromosome 1R in rye. Theor Appl Genet 71:742–749
Lucus H, Moore G, Murphy G, Flavell RB (1992) Inverted repeats in the long-terminal repeats of the wheat retrotransposon Wis 2-1A. Molec Biol Evol 9:716–728
Maniatis T, Fritsch EF, Sambrook J (eds) (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Monte JV, Flavell RB, Gustafson JP (1995) Wis 2-1A: an ancient retrotransposon in theTriticeae tribe. Theor Appl Genet 91:367–373
Mukai Y, Endo TR, Gill BS (1991) Physical mapping of the 18S.26S rRNA multigene family in common wheat: identification of a new locus. Chromosoma 100:71–78
Reddy P, Appels R (1989) A second locus for the 5S multigene family inSecale L.: sequence divergence in two lineages of the family. Genome 32:456–467
Singh NK, Shepherd KW, McIntosh RA (1990) Linkage mapping of genes for resistance to leaf, stem and stripe rusts and ω-secalins on the short arm of 1R. Theor Appl Genet 80:609–616
Southern E (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Molec Biol 98:503
Van Deynze AE, Dubcovsky J, Gill KS, Nelson JC, Sorrells ME, Dvorak J, Gill BS, Lagudah ES, McCouch SR, Appels R (1995) Molecular-genetic maps for group 1 chromosomes of Triticeae species and their relation to chromosomes in rice and oat. Genome 38:45–59
Waters SP, Noble ER, Dalling MJ (1982) Intracellular localization of peptide hydrolases in wheat (Triticum aestivum L.) leaves. Plant Physiol 69:575–579
Author information
Authors and Affiliations
Additional information
Edited by: W. Hennig
Rights and permissions
About this article
Cite this article
Clarke, B.C., Mukai, Y. & Appels, R. TheSec-1 locus on the short arm of chromosome 1R of rye (Secale cereale). Chromosoma 105, 269–275 (1996). https://doi.org/10.1007/BF02524644
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02524644