Summary
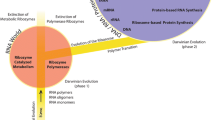
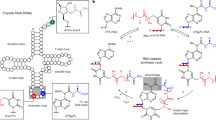
A model is presented for the evolution of metabolism and protein synthesis in a primitive, acellular RNA world. It has been argued previously that the ability to perform metabolic functions logically must have preceded the evolution of a message-dependent protein synthetic machinery and that considerable metabolic complexity was achieved by ribo-organisms (i.e., organisms in which both genome and enzymes are comprised of RNA). The model proposed here offers a mechanism to account for the gradual development of sophisticated metabolic activities by ribo-organisms and explains how such metabolic complexity would lead subsequently to the synthesis of genetically encoded polypeptides. RNA structures ancestral to modern ribosomes, here termed metabolosomes, are proposed to have functioned as organizing centers that coordinated, using base-pairing interactions, the order and nature of adaptor-mounted substrate/catalyst interactions in primitive metabolic pathways. In this way an ancient genetic code for metabolism is envisaged to have predated the specialized modern genetic code for protein synthesis. Thus, encoded amino acids initially would have been used, in conjunction with other encoded metabolites, as building blocks for biosynthetic pathways, a role that they retain in the metabolism of contemporary organisms. At a later stage the encoded amino acids would have been condensed together on similar RNA metabolosome structures to form the first genetically determined, and therefore biologically meaningful, polypeptides. On the basis of codon distributions in the modern genetic code it is argued that the first proteins to have been synthesized and used by ribo-organisms were predominantly hydrophobic and likely to have performed membrane-related functions (such as forming simple pore structures), activities essential for the evolution of membrane-enclosed cells.
Similar content being viewed by others
References
Benner SA, Allemann RK, Ellington AD, Ge L, Glasfeld A, Leanz GF, Krauch T, MacPherson LJ, Moroney S, Piccirilli JA, Weinhold E (1987) Natural selection, protein engineering, and the last riboorganism: rational model building in biochemistry. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 53–63
Brack A, Orgel L (1975) β-structures of alternating polypeptides and their possible prebiotic significance. Nature 256:383–387
Cavalier-Smith T (1987) The origin of cells: a symbiosis between genes, catalysts and membranes. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 805–824
Cech TR, Bass BL (1986) Biological catalysis by RNA. Annu Rev Biochem 55:599–629
Crick FHC (1968) The origin of the genetic code. J Mol Biol 38:367–379
Dyson F (1985) Origins of life. Cambridge University Press, Cambridge, United Kingdom
Eigen M, Gardiner W, Schuster P, Winkler-Oswatitisch R (1981) The origin of genetic information. Sci Am 244:88–118
Epps DE, Sherwood E, Eichberg J, Oro J (1978) Cyanamide mediated synthesis under plausible primitive Earth conditions. V. The synthesis of phosphatic acids. J Mol Evol 11: 279–292
Fox S, Dose K (1972) Molecular evolution and the origin of life. WH Freeman, San Francisco
Gilbert W (1986) The RNA world. Nature 319:618
Gilbert W (1987) The exon theory of genes. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 901–905
Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S (1983) The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35:849–857
Joyce GF (1989) RNA evolution and the origins of life. Nature 338:217–224
Joyce GF, Schwartz AW, Miller SL, Orgel LE (1987) The case for an ancestral genetic system involving simple analogues of the nucleotides. Proc Natl Acad Sci USA 84:4398–4402
Koshland DE (1987) Evolution of catalytic function. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 1–7
Lear JD, Wasserman ZR, DeGrado WF (1988) Synthetic amphiphilic peptide models for protein ion channels. Science 240:1177–1181
Maizels N, Weiner AM (1987) Peptide-specific ribosomes, genomic tags, and the origin of the genetic code. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 743–749
Miller SL, Orgel LE (1974) The origins of life on the Earth. Prentice-Hall, Englewood Cliffs NJ
Moore PB (1987) On the modus operandi of the ribosome. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 721–728
Noller HF (1984) Structure of ribosomal RNA. Annu Rev Biochem 53:119–162
Nomura N (1987) The role of RNA and protein in ribosome function: a review of early reconstitution studies and prospects of future studies. In: Symposia on quantitative biology, vol 52. Cold Spring Harbor Laboratory, Cold Spring Harbor NY, pp 653–663
Orgel LE (1986) RNA catalysis and the origins of life. J Theor Biol 123:127–149
Saenger W (1984) Principles of nucleic acid structure. Springer-Verlag, New York, p 401
Sharp PA (1985) On the origin of RNA splicing and introns. Cell 42:397–400
Shvedova TA, Korneeva GA, Otroshchenko VA, Venkstern TV (1987) Catalytic activity of the nucleic acid component of the 1,4-α-glucan branching enzyme from rabbit muscles. Nucleic Acids Res 15:1745–1752
Visser CM, Kellogg RM (1978) Biotin. Its place in evolution. J Mol Evol 11:171–187
Weiner AM, Maizels N (1987) tRNA-like structures tag the 3′ ends of genomic RNA molecules for replication: implications for the origin of protein synthesis. Proc Natl Acad Sci USA 84:7383–7387
Westheimer FH (1987) Why nature chose phosphates. Science 235:1173–1178
White HB III (1976) Coenzymes as fossils of an earlier metabolic state. J Mol Evol 7:101–104
Yarus M (1988) A specific amino acid binding site composed of RNA. Science 240:1751–1758
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Gibson, T.J., Lamond, A.I. Metabolic complexity in the RNA world and implications for the origin of protein synthesis. J Mol Evol 30, 7–15 (1990). https://doi.org/10.1007/BF02102448
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02102448