Summary
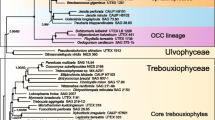
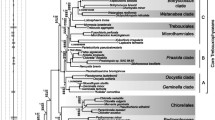
5S rRNA sequences from six additional green algae lend strong molecular support for the major outlines of higher plant and green algae phylogeny that have been proposed under varying naming conventions by several authors. In particular, the molecular evidence now available unequivocally supports the existence of at least two well-separated divisions of the Chlorobionta: the Chlorophyta and the Streptophyta (i.e., charophytes) (according to the nomenclature of Bremer). The chlamydomonad 5S rRNAs are, however, sufficiently distinct from both clusters that it may ultimately prove preferable to establish a third taxon for them. In support of these conclusions 5S rRNA sequence data now exist for members of four diverse classes of chlorophytes. These sequences all exhibit considerably more phylogenetic affinity to one another than any of them show toward members of the other cluster, the Streptophyta, or the twoChlamydomonas strains. Among the Charophyceae, new 5S rRNA sequences are provided herein for three genera,Spirogyra, Klebsormidium, andColeochate. All of these sequences and the previously publishedNitella sequence show greater resemblance among themselves and to the higher plants than they do to any of the other green algae examined to date. These results demonstrate that an appropriately named taxon that includes these green algae and the higher plants is strongly justified. The 5S rRNA data lack the resolution needed, however, to unequivocally determine which of several subdivisions of the charophytes is the sister group of the land plants. The evolutionary diversity ofChlamydomonas relative to the other green algae was recognized in earlier 5S rRNA studies but was unanticipated by ultrastructural work. These new data provide further evidence for the relative uniqueness of the chlamydomonads and are discussed further.
Similar content being viewed by others
References
Bremer K (1985) Summary of green plant phylogeny and classification. Cladistics 1:369–385
Bremer K, Humphries CJ, Mishler BD, Churchill SP (1987) On cladistic relationships in green plants. Taxon 36:339–349
Cavalier-Smith T (1981) Archaebacteria and archezoa. Nature 339:100–101
Cavalier-Smith T (1981) Eukaryotic kingdoms: seven or nine? BioSystems 14:461–481
Chapman DJ (1985) Geological factors and biochemical aspects of the origin of land plants. In: Tiffney BH (ed) Geological factors and the evolution of plants. Yale Univ Press, New Haven, pp 23–45
Darlix J-L, Rochaix JD (1981) Nucleotide sequence and structure of cytoplasmic 5S RNA and 5.8S RNA ofChlamydomonas reinhardii. Nucleic Acids Res 9:1291–1299
Donis-Keller HA, Maxam A, Gilbert W (1977) Mapping adenines, guanines and pyrimidines in RNA. Nucleic Acids Res 4:2527–2538
Felsenstein J (1984) The statistical approach to inferring evolutionary trees and what it tells us about parsimony and compatibility. In: Duncan T, Stuessy TF (eds) Cladistics: perspectives in the reconstruction of evolutionary history. Columbia University Press, New York, pp 169–191
Fox GE (1985) The structure and evolution of archaebacterial ribosomal RNA. In: Sokatch JR, Ornston LN, Woese CR, Wolfe RS (eds) The bacteria. Academic Press, New York, pp 257–310
Fox GE, Stackebrandt E (1987) The application of 16S rRNA cataloguing and 5S rRNA sequencing in bacterial systematics. Methods Microbiol 19:405–458
Graham LE (1984) The origin of the life cycle of land plants. Am Sci 73:178–186
Hinnebusch AG, Klotz LC, Blanken RI, Loeblich AR (1981) An evaluation of the phylogenetic position of the dinoflagellateCrypthecodinium cohnii based on 5S ribosomal RNA characterization. J Mol Evol 17:334–337
Hori H, Osawa S (1979) Evolutionary change in 5S RNA secondary structure and a phylogenetic tree of 54 5S rRNA species. Proc Natl Acad Sci USA 76:381–385
Hori H, Lim BK, Osawa S (1985) Evolution of green plants as deduced from 5S rRNA sequences. Proc Natl Acad Sci USA 82:820–823
Jeffrey C (1967) The origin and differentiation of the archegoniate land plants: a second contribution. Kew Bull 21:335–349
Jeffrey C (1971) Thallophytes and kingdoms—a critique. Kew Bull 25:291–299
Jukes Th, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Jupe ER, Chapman RL, Zimmer EA (1988) Nuclear ribosomal RNA genes and algal phylogeny—theChlamydomonas example. BioSystems 21:223–230
Kumazaki T, Hori H, Osawa S (1988) Phylogeny of protozoa deduced from 5S rRNA sequences. J Mol Evol 18:293–296
Luehrsen KN, Fox GE (1981) Secondary structure of eukaryotic cytoplasmic 5S ribosomal RNA. Proc Natl Acad Sci USA 78:2150–2154
Mattox KR, Stewart KD (1984) Classification of the green algae: a concept based on comparative cytology. In: Irvine DEG, John DM (eds) The systematics of green algae. Academic Press, New York, pp 29–72
Melkonian M (1984) Flagellar apparatus ultrastructure in relation to green algal classification. In: Irvine DEG, John DM (eds) The systematics of the green algae. Academic Press, New York, pp 73–210
Peattie DA (1979) Direct chemical method for sequencing RNA. Proc Natl Acad Sci USA 76:1760–1764
Perasso R, Baroin A, Qu LH, Bachellerie JP, Adoutte A (1989) Origin of the algae. Nature 339:142–144
Pickett-Heaps JD, Ott DW (1974) Cell structure and division inPedinomonas. Cytobios 11:41–58
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sluiman H (1985) A cladistic evaluation of the lower and higher green plants (Viridiplantae). Plant Syst Evol 149:217–232
Sneath PH, Sokal RR (1973) Numerical taxonomy. WH Freeman, San Francisco
Sourdis J, Krimbas C (1987) Accuracy of phylogenetic trees estimated from DNA sequence data. Mol Biol Evol 4:159–166
Starr RC (1978) Culture collection of algae at the University of Texas at Austin. J Phycol 14:46–100
Steele KP, Holsinger KE, Jansen RK, Taylor DW (1988) Phylogenetic relationships in green plants—a comment on the use of 5S ribosomal RNA sequences by Bremer et al. Taxon 37:135–138
Wolters J, Erdmann VA (1984) Comparative analyses of small ribosomal RNAs with respect to the evolution of plastids and mitochondria. Endocyt Cell Res 1:1–23
Wolters J, Erdmann VA (1986) Cladistic analysis of 5S rRNA and 16S rRNA secondary and primary structure—the evolution of eukaryotes and their relation to archaebacteria. J Mol Evol 24:152–166
Wolters J, Erdmann VA (1988) Compilation of 5S rRNA and 5S rRNA gene sequences. Nucleic Acids Res 16:r1-r70
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Devereux, R., Loeblich, A.R. & Fox, G.E. Higher plant origins and the phylogeny of green algae. J Mol Evol 31, 18–24 (1990). https://doi.org/10.1007/BF02101788
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02101788