Summary
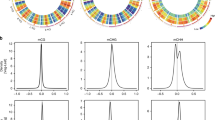
The differentiation processes of the metaxylem cell line in the root ofAllium cepa are characterized by amplification phenomena of repetitive DNA sequences mainly localized in heterochromatic regions of metaphase chromosomes. Moreover, these sequences are heavily methylated. This paper presents additional results on variation in endogenous DNA methylation in different developing root segments. The results show that methylation is higher in apical meristematic cells than the differentiating segments; contrastingly, total RNA synthesis seems to be correlated with undermethylation. Addition of labelled methyl groups to DNA by eukaryotic methylase, DNA digestions with different restriction enzymes specific for methylated sites and HPLC analysis confirmed the above results. Moreover, variation in methylation levels during differentiation occur not only at the internal cytosine of the-CCG-sites, but also at external cytosine. Furthermore, methylation affects other sites containing the trinucleotides-CXG-. In conclusion, root differentiation inAllium cepa seems to be correlated with gene activation modulated by the methylation/demethylation of particular DNA sequences.
Similar content being viewed by others
References
Avanzi S, Maggini F, Innocenti AM (1973) Amplification of ribosomal cistrons during the maturation of metaxylem in the root ofAllium cepa. Protoplasma 76: 197–210
Barnes SR, James AM, Jamieson G (1985) The organization, nucleotide sequences and chromosomal distribution of a satellite DNA fromAllium cepa. Chromosoma 92: 185–192
Bassi P, Cionini PG, Cremonini R, Seghizzi P (1984) Underrepresentation of nuclear DNA sequences in differentiating root cells ofVicia faba. Protoplasma 127: 70–77
Cremonini R, Loiero M, Durante M (1981) Differentiation of metaxylem cell line in the root ofAllium cepa. II. Changes in nucleotide sequence redundancy. Protoplasma 108: 1–7
Delsney M, Laroche M, Penon P (1984) Methylation pattern ofRaphanus sativus nuclear rRNA genes. Plant Physiol 76: 627–632
Deumling B (1981) Sequence arrangement of a highly methylated satellite DNA of a plant,Scilla: a tandemly repeated inverted repeat. Proc Natl Acad Sci USA 78: 338–342
Doerfler W (1983) DNA methylation and gene acticity. Annu Rev Biochem 52: 93–124
Durante M, Cremonini R, Brunori A, Avanzi S, Innocenti AM (1977) Differentiation of metaxylem cell line in the root ofAllium cepa. I. DNA heterogeneity and ribosomal cistrons of two different stages of differentiation. Protoplasma 93: 289–303
—, Geri C, Ciomei M (1981) DNA methylation in differentiating plant pith tissue. Experientia 38: 451–452
—, Tagliasacchi AM, Avanzi S (1985) Fast reannealing sequences of DNA inAllium cepa: characterization and chromosomal localization. Cytobios 44: 263–271
—, Cecchini E, Natali L, Citti L, Geri C, Parenti R, Nuti Ronchi V (1989) 5-Azacytidine-induced tumorous transformation and DNA hypomethylation inNicotiana tissues cultures. Dev Genet 10: 298–303
Ehrlich M, Wang RYH (1981) 5-methylcytosine in eukaryotic DNA. Science 212: 1350–1357
—, Gama-Sosa MA, Huang LH, Midgett RM, Kuo KC, McCune RA, Gehrke CW (1982) Amount and distribution of 5-methylcytosine in human DNA from different types of tissues or cells. Nucleic Acids Res 10: 2709–2722
Felsenfeld G, McGhee J (1982) Methylation and gene control. Nature 296: 602–603
Frediani M, Forino LMC, Tagliasacchi AM, Cionini PG, Durante M, Avanzi S (1986 a) Functional heterogeneity, during early embryogenesis, ofPhaseolus coccineus ribosomal cistrons in polytene chromosomes of embryo suspensor. Protoplasma 132: 51–57
—, Tagliasacchi AM, Durante M, Avanzi S (1986 b) Distribution of 5-methylcytosine rich regions in the polytene chromosomes ofPhaseolus coccineus embryo suspensor as shown by the immunoperoxidase technique. Exp Cell Res 167: 337–342
Gama-Sosa MA, Midgett RM, Slagel CA, Gitens S, Kuo KC, Gehrke CW, Ehrlich M (1983) Tissue specific differences in DNA methylation in various mammals. Biochim Biophys Acta 740: 212–219
Gehrke CW, McCune RA, Gama-Sosa MA, Ehrlich M, Kuo KC (1984) Quantitative reversed-phase high-performance liquid chromatography of major and modified nucleosides in DNA. J Chromatogr 301: 199–219
Grisvard J (1985) Different methylation pattern of melon satellite DNA sequences in hypocotyl and callus tissues. Plant Sci 39: 189–193
Gruenbaum Y, Naveh-Many T, Cedar M, Razin A (1981) Sequence specificity of methylation in higher plant DNA. Nature 292: 860–862
Haigh LS, Owens BB, Hellewell J, Ingram VM (1982) DNA methylation in chicken α-globin gene expression. Proc Natl Acad Sci USA 79: 5332–5336
Jones PA, Taylor JM (1980) Cellular differentiation, cytidine analogs and DNA methylation. Cell 20: 85–93
McClelland M (1983) The frequency and distribution of methylable DNA sequences in leguminous plant protein coding genes. J Mol Evol 19: 346–354
Manes C, Menzel P (1981) Demethylation of CpG sites in DNA of early rabbit trophoblast. Nature 293: 589–590
Monk M, Boubelik M, Lehnert S (1987) Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ lineages during mouse embryo development. Development 99: 371–382
Natali L, Cavallini A, Cremonini R, Bassi P, Cionini PG (1986) Amplification of nuclear sequences during induced plant cell dedifferentiation. Cell Differ 18: 157–161
Nick H, Bowen B, Ferl R, Gilbert W (1986) Detection of cytosine methylation in the maize alcohol dehydrogenase gene by genomic sequences. Nature 319: 243–246
Razin A, Cedar M (1984) DNA methylation in eukaryotic cells. Int Rev Cytol 92: 159–184
—, Riggs AD (1980) DNA methylation and gene function. Science 210: 604–610
—, Feldmesser E, Kafri T, Szyf M (1985) Cell specific DNA methylation patterns: formation and a nucleosome locking model for their function. In: Cantoni G, Razin A (eds) Biochemistry and biology of DNA methylation. AR Liss, New York, pp 239–253
—, Szyf M, Kafri T, Roll M, Giloh H, Scarpa S, Carotti D, Cantoni LG (1986) Replacement of 5-methylcytosine by cytosine: a possible mechanism for transient DNA demethylation during differentiation. Proc Natl Acad Sci USA 83: 2827–2831
Scott NS, Kavanagh TA, Timmis JM (1984) Methylation of rRNA genes in some higher plants. Plant Sci Lett 35: 213–217
Singer J, Ems JR, Riggs AD (1979) Methylation of mouse liver DNA studied by means of the restriction enzymes MspI and HpaII. Science 203: 1019–1021
Spena A, Viotti A, Pirrotta V (1983) Two adjacent genomic zein sequences: structure, organization and tissue specific restriction pattern. J Mol Biol 169: 799–811
Vanyushin BF, Tkacheva SG, Belozersky AN (1970) Rare base in animal DNA. Nature 225: 948–949
Wagner I, Capesius I (1981) Determination of 5-methylcytosine from plant DNA by high-performance liquid chromatography. Biochim Biophys Acta 654: 52–56
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Durante, M., Frediani, M., Mariani, L. et al. Differentiation of metaxylem cell line in the root ofAllium cepa L.. Protoplasma 158, 149–154 (1990). https://doi.org/10.1007/BF01323127
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01323127