Abstract
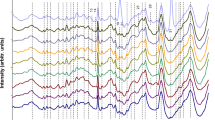
Correspondence analysis was used to classify the pattern-like FT-IR spectra of intact bacteria. The analysis was performed on a data set of approximately 80 normalized spectral derivatives of a selection of pathogenic bacteria. The correspondence analysis proved that the various different bacterial species were clustering in distinct regions of the correspondence maps suggesting that there do exist correlations between spectral data and biochemical/microbiological classification.
Similar content being viewed by others
References
J. E. S. Greenstreet, K. P. Norris,Spectrochim. Acta 1957,9,177.
K. P. Norris,J. Hyg. 1959,57, 326.
J. W. Riddle, P. W. Kabler, B. A. Kenner, R. H. Bordner, S. W. Rockwood, H. J. R. Stevenson,J. Bact. 1956,72, 593.
P. Giesbrecht, D. Naumann, H. Labischinski, G. Barnickel, in:Rapid Methods and Automation in Microbiology and Immunology (K.-O. Habermehl, ed.), Springer-Verlag, Berlin-Heidelberg-New York-Tokyo, 1985, p. 198.
D. Naumann,SPIE 1985,553, 268.
M. van Heel, W. Keegstra,Ultramicroscopy 1981,7, 113.
J. Frank, M. van Heel,J. Mol. Biol. 1982,161, 134.
M. van Heel,Ultramicroscopy 1984,13, 165.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Naumann, D., Fijala, V. & Labischinski, H. The differentiation and identification of pathogenic bacteria using FT-IR and multivariate statistical analysis. Mikrochim Acta 94, 373–377 (1988). https://doi.org/10.1007/BF01205910
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF01205910