Summary
-
1.
The measurement of cellular mRNA content by quantitativein situ hybridization is a valuable approach to the study of gene expression in brain since this tissue exhibits a high degree of phenotypic heterogeneity.
-
2.
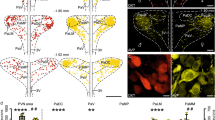
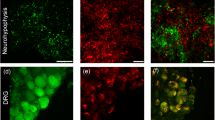
The cellular content of vasopressin and oxytocin mRNA in hypothalamo-neurohypophysial system neurons was altered by maintaining rats for 24 hr on 2% sodium chloride water.
-
3.
Statistical and graphical techniques were then used to analyze cell by cell how mRNA levels were altered as a result of osmotic stimulation. We propose that the negative binomial probability distribution is a suitable model to describe how mRNA content varies across a defined cell population. For both measures of oxytocin and vasopressin mRNA levels, maximum-likelihood estimation indicated that this model adequately described empirical findings obtained from rats drinking tap water or salt water.
-
4.
Both graphical and statistical analyses suggested how the defined neural system responds to osmotic stimulation: mRNA content was altered as a multiplicative function of “initial state.” The utility and limitations of the quantitative approach are discussed.
Similar content being viewed by others
References
Chambers, J. M., Cleveland, W. S., Kleiner, B., and Tukey, P. A. (1983).Graphical Methods for Data Analysis, Wadsworth, Belmont, CA.
Defendini, R., and Zimmerman, E. A. (1978). The magnocellular neuro-secretory system of the mammalian hypothalamus. InThe Hypothalamus (S. Reichlin, R. J. Baldessarini, and J. B. Martin, Eds.), Raven Press, New York, pp. 137–152.
England, J. M., and Rogers, A. W. (1970). The statistical analysis of autoradiographs. I. Grain count distributions over uniformly labelled sources.J. Microsc. 92159–165.
Hou-Yu, A., Lamme, A. T., Zimmerman, E. A., and Silverman, A.-J. (1986). Comparative distribution of vasopressin and oxytocin neurons in the rat brain using a double-label procedure.Neuroendocrinology 44235–246.
Hyodo, S., Fujiwara, M., Sato, M., and Urano, A. (1988). Molecular- and immuno-histochemical study on expression of vasopressin and oxytocin genes following sodium loading.Zoo. Sci. 51033–1042.
Ivell, R., and Richter, D. (1984). Structure and comparison of the oxytocin and vasopressin genes from rat.Proc. Natl. Acad. Sci. USA 812006–2010.
Kawata, M. (1983). Immunohistochemistry of oxytocin and vasopressin neurons in dog and rat under normal and experimental conditions. InStructure and Function of Peptidergic and Aminergic Neurons (Y. Sano, Y. Ibata, and E. A. Zimmerman, Eds.), JSSP-VNU Science Press, Tokyo, pp. 33–53.
Kawata, M., McCabe, J. T., Harrington, C., Chikaraishi, D., and Pfaff, D. W. (1988a).In situ hybridization analysis of osmotic stimulus-induced changes in ribosomal RNA in rat supraoptic nucleus.J. Comp. Neurol. 270528–536.
Kawata, M., McCabe, J. T., and Pfaff, D. W. (1988b).In situ hybridization histochemistry with oxytocin synthetic oligonucleotide: Strategy for making the probe and its application.Brain Res. Bull. 20693–697.
Kawata, M., McCabe, J. T., Pfaff, D. W., and Sano, Y. (1988c). Gene expression for posterior pituitary hormones studied byin situ hybridization histochemistry. InRecent Progress in Posterior Pituitary Hormones 1988 (S. Yoshida and L. Share, Eds.), Elsevier Science, Amsterdam, pp. 249–255.
Kendall, M., and Stuart, A. (1977).The Advanced Theory of Statistics, Vol. 1, 4th ed., Griffin, London.
Lewis, M. E., Rogers, W. T., Krause, R. G., II, and Schwaber, J. S. (1989). Quantitation and digital representation ofin situ hybridization histochemistry.Met. Enzymol. 168808–821.
Lightman, S. L., and Young, W. S., III (1987). Vasopressin, oxytocin, dynorphin, enkephalin and corticotrophin-releasing factor mRNA stimulation in the rat.J. Physiol. 39423–39.
McCabe, J. T., and Pfaff, D. W. (1989).In situ hybridization: A methodological guide.Meth. Neurosci. 198–126.
McCabe, J. T., Morrell, J. I., Ivell, R., Schmale, H., Richter, D., and Pfaff, D. W. (1986a).In situ hybridization to localize rRNA and mRNA in mammalian neurons.J. Histochem. Cytochem. 3445–50.
McCabe, J. T., Morrell, J. I., and Pfaff, D. W. (1986b).In situ hybridization as a quantitative autoradiographic method: An example from vasopressin and oxytocin gene transcription in the Brattleboro rat. InIn Situ Hybridization in Brain (G. R. Uhl, Ed.), Plenum Press, New York, pp. 73–95.
McCabe, J. T., Morrell, J. I., and Pfaff, D. W. (1986c). Measurement of vasopressin and oxytocin genes in single neurons byin situ hybridization. InNeuroendocrine Molecular Biology (G. Fink, A. J. Harmar, and K. W. McKerns, Eds.), Plenum Press, New York, pp. 219–229.
McCabe, J. T., Desharnais, R. A., and Pfaff, D. W. (1989). Graphical and statistical approaches to data analysis forin situ hybridization.Meth. Enzymol. 168822–848.
Przybylski, R. J. (1970). Principles of quantitative autoradiography. InIntroduction to Quantitative Cytochemistry—II (G. L. Wied and G. F. Bahr, Eds.), Academic Press, New York, pp. 477–505.
Rhodes, C. H., Morrell, J. I., and Pfaff, D. W. (1981). Immunohistochemical analysis of magnocellular elements in rat hypothalamus: Distribution and numbers of neurophysin, oxytocin and vasopressin containing cells.J. Comp. Neurol. 19845–64.
Schmale, H., Ivell, R., Briendl, M., Darmer, D., and Richter, D. (1984). The mutant vasopressin gene from diabetes insipidus (Brattleboro) rats is transcribed but the message is not efficiently translated.EMBO J. 33289–3293.
Sherman, T. G., McKelvey, J. F., and Watson, S. J. (1986). Vasopressin mRNA regulation in individual hypothalamic nuclei: A Northern andin situ hybridization analysis.J. Neurosci. 61685–1694.
Takamatsu, T., Kitamura, T., and Fujita, S. (1986). Quantitative fluorescence image analysis.Acta Histochem. Cytochem. 1961–71.
Tukey, J. W. (1977).Exploratory Data Analysis, Addison-Wesley, Reading, MA.
Uhl, G. R. (1989).In situ hybridization: Quantitation using radiolabled hybridization probes.Meth. Enzymol. 168741–752.
Vandesande, F., and Dierickx, K. (1975). Identification of the vasopressin producing and oxytocin producing neurons in the hypothalamic magnocellular neurosecretory system of the rat.Cell Tissue Res. 164153–162.
Van Tol, H. H. M., Voorhuis, D. Th. A. M., and Burbach, J. P. H. (1987). Oxytocin gene expression in discrete hypothalamic magnocellular cell groups is stimulated by prolonged salt loading.Endocrinology 12071–76.
Wilk, M. B., and Gnanadesikan, R. (1968). Probability plotting methods for the analysis of data.Biometrika 551–17.
Young, W. S., III (1989).In situ hybridization histochemical detection of neuropeptide mRNAs using DNA and RNA probes.Meth. Enzymol. 168702–710.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
McCabe, J.T., Kawata, M., Sano, Y. et al. Quantitativein situ hybridization to measure single-cell changes in vasopressin and oxytocin mRNA levels after osmotic stimulation. Cell Mol Neurobiol 10, 59–71 (1990). https://doi.org/10.1007/BF00733636
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00733636