Summary
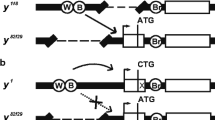
We undertook a deletional analysis of the gypsy retrotransposon in order to determine which sequences of the element are required for its mutagenic effect. We show that a phenotype indistinguishable from that ofy 2 flies can be generated by transformingy − flies with a construct containing theyellow gene and a gypsy element located at the same insertion site inyellow as found iny 2 flies. When flies are transformed with similar constructs in which increasing amounts of the 5′ transcribed untranslated region of gypsy have been removed, either a partialy 2 revertant or a completely revertant phenotype is obtained. These results yield direct proof that the region of gypsy to which thesu(Hw) protein binds is required for the generation of mutant phenotypes by this retrotransposon.
Similar content being viewed by others
References
Biessmann H (1985) Molecular analysis of theyellow gene (y) region ofDrosophila melanogaster. Proc Nat Acad Sci USA 82:7369–7373
Boeke JD, Corces VG (1989) Transcription and reverse transcription of retrotransposons. Annu Rev Microbiol 43:403–434
Bohmann D, Keller W, Dale T, Scholer HR, Tebb G, Mattaj IW (1987) A transcription factor which binds to the enhancers of SV40, immunoglobulin heavy chain and U2 snRNA genes. Nature 325:268–272
Chia W, Howes G, Martin M, Meng YB, Moses K, Tsubota S (1986) Molecular analysis of theyellow locus ofDrosophila. EMBO J 5:3597–3605
Flavell AJ, Alphey LS, Ross SJ, Leigh-Brown AJ (1990) Complete reversions of a gypsy retrotransposon-inducedcut locus mutation inDrosophila melanogaster involving jockey transposon insertions and flanking gypsy sequence deletions. Mol Gen Genet 220:181–185
Geyer PK, Corces VG (1987) Separate regultory elements are responsible for the complex pattern of tissue-specific and developmental transcription of theyellow locus inDrosophila melanogaster. Genes Dev 1:996–1004
Geyer PK, Green MM, Corces VG (1988) Mutant gene phenotypes mediated by aDrosophila melanogaster retrotransposon require sequences homologous to mammalian enhancers. Proc Nat Acad Sci USA 85:8593–8597
Geyer PK, Green MM, Corces VG (1990) Tissue-specific transcriptional enhancers may act in trans on the gene located in the homologous chromosome: the molecular basis of transvection inDrosophila. EMBO J 9:2247–2256
Geyer PK, Spana C, Corces VG (1986) On the molecular mechanism of gypsy-induced mutations at theyellow locus ofDrosophila melanogaster. EMBO J 5:2657–2662
Holdridge C, Dorsett D (1991) Repression ofhsp70 heat shock gene transcription by thesuppressor ofHairy-wing protein ofDrosophila melanogaster. Mol Cell Biol 11:1894–1900
Hope IA, Mahadevan S, Struhl K (1988) Structural and functional characterization of the short acidic transcriptional activation region of yeast GCN4 protein. Nature 333:635–640
Karess RE, Rubin GM (1984) Analysis of P transposable element functions in Drosophila. Cell 38:135–146
Klug A, Rhodes D (1987) “Zinc fingers”: a novel protein motif for nucleic acid recognition. Trends Biochem Sci 12:464–469
Landschulz WH, Johnson PF, McKnight SL (1988) The leucine zipper: a hypothetical structure common to a new class of DNA binding proteins. Science 240:1759–1764
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: A laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Mazo AM, Mizrokhi LJ, Karavanov AA, Sedkov YA, Krichevskaya AA, Ilyin YV (1989) Suppression inDrosophila: su(Hw) andsu(f) gene products interact with a region of mdg4 (gypsy) regulating its transcriptional activity. EMBO J 8:903–911
Mizrokhi LJ, Obolenkova LA, Priimagi AF, Ilyin YV, Gerasimova TI, Georgiev GP (1985) The nature of unstable insertion mutations and reversion in the locuscut ofDrosophila melanogaster: molecular mechanisms of transposition memory. EMBO J 4:3781–3787
Modolell J, Bender W, Meselson M (1983).D. melanogaster mutations suppressible by thesuppressor ofHairy-wing are insertions of a 7.3 kb mobile element. Proc Nat Acad Sci USA 80:1678–1682
Parkhurst SM, Corces VG (1986) Interactions among the gypsy transposable element and theyellow and thesuppressor ofHairy-wing loci inDrosophila melanogaster. Moll Cell Biol 6:47–53
Parkhurst SM, Harrison DA, Remington MP, Spana C, Kelley RL, Coyne RS, Corces VG (1988) TheDrosophila su(Hw) gene, which controls the phenotypic effect of the gypsy transposable element, encodes a putative DNA binding protein. Genes Dev 2:1205–1215
Peifer M, Bender W (1988) Sequences of the gypsy transposon ofDrosophila necessary for its effects on adjacent genes. Proc Nat Acad Sci USA 85:9650–9654
Rosales R, Vigneron M, Macchi M, Davidson I, Xiao JH, Chambon P (1987) In vitro binding of cell-specific and ubiquitous nuclear proteins to the octamer motif of the SV40 enhancer and related motifs present in other promoters and enhancers. EMBO J 6:3015–3025
Rubin G, Spradling AC (1982) Genetic transformation ofDrosophila with transposable element vectors. Science 218:348–353
Rutledge BJ, Mortin MA, Schwarz E, Thierry-Mieg D, Meselson M (1988) Genetic interactions of modifier genes and modifiable alleles inDrosophila melanogaster. Genetics 119:391–397
Spana C, Corces VG (1990) DNA bending is a determinant of binding specificity for aDrosophila zinc finger protein. Genes Dev 4:1505–1515
Spana C, Harrison DA, Corces VG (1988) TheDrosophila melanogaster suppressor of Hairy-wing protein binds to specific sequences of the gypsy retrotransposon. Genes Dev 2:1414–1423
Author information
Authors and Affiliations
Additional information
Communicated by B.H. Judd
Rights and permissions
About this article
Cite this article
Smith, P.A., Corces, V.G. The suppressor of Hairy-wing binding region is required for gypsy mutagenesis. Molec. Gen. Genet. 233, 65–70 (1992). https://doi.org/10.1007/BF00587562
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00587562