Summary
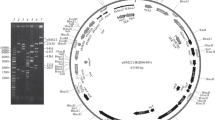
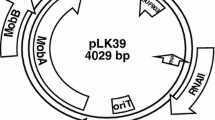
By hybridizing the IncFVII haemolytic plasmid pSU233 with a probe containing the origin of transfer of the IncFII plasmid R1, we isolated a 1.9 kb BglII fragment containing at least the origin of transfer (oriT), and the genes traM and finP. Functional complementation analysis of deletion derivatives was used to map the origin of transfer. We also determined the nucleotide sequence of traM and finP. Comparison with similar regions of several plasmids, also belonging to the RepFIIA family, revelaed that pSU233 resembles the F plasmid by very close. The homology is not evenly distributed along this region, but clustered into homologous regions (TraZb-oriT, TraMb-oriT and traM separated by non-homologous regions (TraYb-oriT, finP). This organization resembles that reported for the replication region and also suggests evolution by exchange of modules. In addition, the nucleotide sequence of finP is different from those previously described for other IncF plasmids and constitutes a new allele, which we have denominated allele VI.
Similar content being viewed by others
References
Achtman M, Willetts N, Clark AJ (1971) Beginning a genetic analysis of conjugational transfer determined by the F factor in E. coli by isolation and characterization of transfer-deficient mutants. J Bacteriol 106:529–538
Alfaro G, Willetts N (1972) The relationship between the transfer systems of some bacterial plasmids. Genet Res 20:229–289
Andrés I, Rodriguez JC, Barbadillo J, Ortiz JM (1987) Identification and expression of a copy number control gene in the IncFIII hemolytic plasmid pSU316. J Bacteriol 169:2405–2409
Bergquist PL, Saadi S, Maas WK (1986) Distribution of basic replicons having homology with RepFIA, RepFIB and RepFIC among IncF group plasmids. Plasmid 15:19–34
Bradley DE (1980) Morphological and serological relationships of conjugative pili. Plasmid 4:155–169
De la Cruz F, Gristed J (1982) Genetic and molecular characterization of Tn ZI, a multiple resistance transposon of R100. I. J Bacteriol 151:222–228
De la Cruz F, Zabala JC, Ortiz JM (1979) Incompatibility among α-hemolytic plasmids studied after inactivation of the hemolysin gene by transposition of Tn802. Plasmid 2:507–519
Dempsey WB (1987) Transcript analysis of the plasmid R100 traJ and finP genes. Mol Gen Genet 209:533–544
Fee BE, Dempsey WB (1986) Cloning, mapping and sequencing of plasmid R100 traM and finP genes. J Bacteriol 167:336–345
Feinberg AP, Vogelstein B (1983). A technique for radiolabelling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Finlay BB, Frost LS, Paranchych W (1986a) Origin of transfer of IncF plasmids and nucleotide sequence of the type II oriT, traM and traY alleles from Co1B4-K98 and the type IV traY allele from R100-1. J Bacteriol 168:132–139
Finlay BB, Frost LS, Paranchich W, Willetts N (1986b) Nucleotide sequences of five IncF plasmid finP alleles. J Bacteriol 167:754–757
Finlay BB, Frost LS, Paranchich W (1986c) Nucleotide sequences of the R1-19 plasmid transfer genes traM, finP, traJ and traY and traYZ promoter. J Bacteriol 166:368–374
Finnegan D, Willetts N (1971) Two classes of Flac mutants insensitive to transfer inhibition by an F-like R factor. Mol gen Genet 111:256–264
Henikoff S (1984) Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene 28:351–359
Ish-Horowicz D, Burke JF (1981) Rapid and efficient cosmid cloning. Nucleic Acids Res 9:2989–2998
Korneluk RG, Quan F, Gravel RA (1985) Rapid and reliable dideoxy sequencing method of double-stranded DNA. Gene 40:317–323
Koraimann G, Koraimann C, Koronakis V, Schlager S, Hogenauer G (1991) Repression and derepression of conjugation of plasmid R1 by wild-type and mutated finP antisense RNA. Mol Microbiol 5:77–87
Koronakis VE, Bauer E, Hogenauer G (1985) The traM gene of the resistance plasmid R1: comparison with the corresponding sequence of the E. coli F-factor. Gene 36:79–86
Lahuc E, Matson SW (1990) Purified E. coli F-factor TraY protein binds oriT. J Bacteriol 172:1385–1391
López J, Crespo P, Rodríguez JC, Andrés I, Ortiz JM (1989a) Analysis of IncF plasmids evolution: nucleotide sequence of an IncFIlI plasmid replication region. Gene 78:183–187
Lopéz, J, Rodríguez JC, Andrés I, Ortiz JM (1989b) Characterization of the RepFVII replicon of the haemolytic plasmid pSU233: nucleotide sequence of an IncFVII determinant J Gen Microbiol 135:1763–1768
López J, Delgado D, Andrés I, Ortiz JM, Rodríguez JC (1991a) Isolation and evolutionary analysis of a RepFVIB replicon of the plasmid pSU212. J Gen Microbiol 137:1093–1099
López J, Salazar L, Andrés I, Ortiz JM, Rodríguez JC (1991b) Nucleotide sequence of the oriT-traM-finP region of the haemolytic plasmid pSU316: Comparison to F. Nucleic Acids Res 19:3451
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning. Cold Spring Harbor Laboratory Press Cold Spring Harbor, New York
McIntire SA, Dempsey W (1987) oriT sequence of the antibiotic resistance plasmid R100. J Bacteriol 169:3829–3832
Meinkoth J, Wahl G (1984) Hybridization of nucleic acids inmobilized on solid supports. Anal Biochem 138:267–284
Meynell E, Datta N (1967) Mutant drug-resistant factors of high transmissibility. Nature 214:885–887
Mullineaux P, Willetts N (1985) Promoters in the transfer region of plasmid F. In: Helinski DR, Cohen SN, Clewell DB, Jackson DA, Hollander A (eds) Plasmds in bacteria. Plenum Press, New York, London, pp 605–614
Novick RP (1987) Plasmid incompatibility. Microbiol Rev 51:381–395
Ostermann E, Kricek F, Hogenauer G (1984) Cloning the origin of transfer region of the resistance plasmid R1. EMBO J 3:1731–1735
Queen C, Korn LJ (1984) A comprehensive sequence analysis program for the IBM personal computer. Nucleic Acids Res 12:581–599
Saadi S, Maas WK, Hill DF, Bergquist PL (1987) Nucleotide sequence analysis of RepFIC, a basic replicon present in IncFI plasmids P307 and F, and its relation to the RepA replicon of IncFII plasmids. J Bacteriol 169:1836–1846
Schwab M, Gruber H, Hogenauer G (1991) The TraM protein of plasmid R1 is a DNA-binding protein. Mol Microbiol 5:439–446
Takeshita S, Sato M, Toba M, Masahashi W, Hashimotoh-Gotoh T (1987) High copy number and low copy number plasmid vectors for lacZ complementation and chloramphenicol or kanamycin resistance selection. Gene 61:63–74
Thompson R, Taylor L (1982) Promoter mapping and DNA sequencing of the F plasmid transfer genes traM and traJ. Mol Gen Genet 188:513–518
Thompson R, Taylor R, Kelly K, Everett R, Willetts N (1984) The F-plasmid origin of transfer: DNA sequence of wild-type and mutant origins and location of origin specific nicks. EMBO J 3:1175–1180
Thompson TL, Centola MB, Deonier RC (1989) Location of the nick at oriT of the F plasmid. J Mol Biol 207:505–512
Willetts N (1977) The transcriptional control of fertility in F-like plasmids. J Mol Biol 112:141–148
Willetts N, Maule J (1985) Specifities of IncF plasmid conjugation genes. Genet Res 47:1–11
Willetts N, Skurray R (1980) The conjugation systems of F-like plasmids. Annu Rev Genet 14:41–76
Willetts N, Skurray R (1987) Structure and function of the F factor mechanism of conjugation in E. coli and Salmonella typhimurium. Cellular and molecular biology. Neidhardt NC (ed) American Society for Microbiology, Washington DC, pp 1110–1133
Zabala JC, De la Cruz F, Ortiz JM (1983) Relationship of the replication and incompatibility genes of an IncFVI hemolysin plasmid with other F-like hemolysin plasmids. J Gen Microbiol 129:801–807
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Salazar, L., Lopéz, J., Andrés, I. et al. Characterization and nucleotide sequence of the orif-traM-finP region of the IncFVII plasmid pSU233. Molec. Gen. Genet. 234, 442–448 (1992). https://doi.org/10.1007/BF00538704
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00538704