Summary
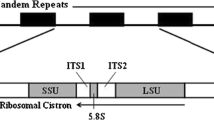
The rRNA genes are arranged in three sequential operons preceded by a fourth partial operon. Part or all of a 1462 nucleotide sequence extending from within the 3′-end of the 23S rRNA gene, across the 5S rRNA gene and a presumptive transcription terminator, to within the first structural gene (for 16S rRNA) of the rrn operon was determined for each region between operons. Homologies of the 3′-end of the 23S rRNA gene with the 4.5S rRNA genes of higher plant chloroplasts, and of the 5S rRNA gene with other 5S rRNA genes were examined. The region preceding the 16S rRNA gene, which is expected to contain sites for initiation and regulation of rrn transcription, includes a 305 base-pair sequence with substantial homology with structural genes elsewhere in the chloroplast genome. The homologies suggest that this portion of the leader evolved from copies of parts of the structural genes which had been inserted before the 16S rRNA genes. Thus the chloroplast rrn leader may provide a unique opportunity to study how a regulatory sequence evolved from well-defined structural genes.
Similar content being viewed by others
References
Adams J (1982) Gene duplication and the birthday problem. Nature 296:176–177
Albertini AM, Hofer M, Calos MP, Miller JH (1982) On the formation of spontaneous delections: the importance of short sequence homologies in the generation of large deletions. Cell 29:319–328
Bambach RK, Scotese CR, Ziegler AM (1981) Before Pangea: the geographies of the paleozoic world. Am Sci 68:26–38
Berkner KL, Folk WR (1980) Polynucleotide kinase exchange as an assay for class II restriction endonucleases. In: Grossman L, Moldave K (eds) Methods in Enzymology 651 pp. 28–36. Academic Press, New York
Broslus J, Dull TJ, Noller HF (1980) Complete nucleotide sequence of a 23S ribosomal RNA gene from Escherichia coli. Proc Natl Acad Sci USA 77:201–204
Chang SH, Hecker LI, Brum CK, Schnabel JJ, Hackman JE, Silberklang M, RajBhandary UL, Barnett WE (1981) The nucleotide sequence of Euglena cytoplasmic phenylalanine transfer RNA. Evidence for possible classification among the animal rather than the plant kingdom. Nucl Acids Res 9:3199–3204
Delihas N, Anderson J, Andresini W, Kaufman L, Lyman H (1981) The 5S ribosomal RNA of Euglena gracilis cytoplasmic ribosomes is closely homologous to the 5S RNA of the trypanosomatid protozoa. Nucl Acids Res 9:6627–6633
DeWachter R, Chen MW, Vandenberghe A (1982) Conservation of secondary structure in 5S ribosomal RNA: A uniform model for eukaryotic, eubacterial, archaebacterial and organelle sequences is energetically favorable. Biochemie 64:311–329
Drake JW (1969) Comparative spontaneous mutation rates. Nature 221:1128–1132
Edlund T, Normark S (1981) Recombination between short DNA homologies causes tandem duplication. Nature 292:269–271
Edwards K, Bedbrook J, Dyer T, Kössel H (1981) 4.5S rRNA from Zea mays chloroplasts shows structural homology with the 3′-end of prokaryotic 23S rRNA. Biochem Int 2:533–538
El-Gewely MR, Helling RB (1980) Preparative separation of DNA-ethidium bromide complexes by zonal density gradient centrifugation. Anal Biochem 102:423–428
El-Gewely MR, Lomax MI, Lau ET, Helling RB, Farmerie W, Barnett WE (1981) A map of specific cleavage sites and tRNA genes in the chloroplast genome of Euglena gracilis bacillaris. Mol Gen Genet 181:296–305
El-Gewely MR, Helling RB, Farmerie W, Barnett WE (1982) Location of a phenylalanine tRNA gene on the physical map of the Euglena gracilis chloroplast genome. Gene 17:337–339
Erdman VA, Huysmans E, Vandenberghe A, DeWachter R (1983) Collection of published 5S and 5.8S ribosomal RNA sequences. Nucl Acids Res 11:r105-r133
Gausing K (1977) Regulation of ribosome production in Escherichia coli: synthesis and stability of ribosomal RNA and of ribosomal protein messenger RNA at different growth rates. J Mol Biol 115:335–354
Gegenheimer P, Apirion D (1981) Processing of prokaryotic ribonucleic acid. Microbiol Rev 45:502–541
Girvitz S, Bacchetti S, Rainbow AJ, Graham FL (1980) A rapid and efficient procedure for the purification of DNA from agarose gels. Anal Biochem 106:492–496
Graf L, Kossel H, Stutz E (1980) Sequencing of 16S–23S spacer in a ribosomal RNA operon of Euglena gracilis reveals two tRNA genes. Nature 286:908–910
Graf L, Roux E, Stutz E, Kössel H (1982) Nucleotide sequence of a Euglena gracilis chloroplast gene coding for the 16S rRNA: homologies to E. coli and Zea mays chloroplast 16S rRNA. Nucl Acids Res 10:6369–6381
Gray PW, Hallick RB (1979) Isolation of Euglena gracilis chloroplast 5S ribosomal RNA and mapping the 5S rRNA gene on chloroplast DNA. Biochemistry 18:1820–1825
Hallick RB (1983) Chloroplast DNA. In: Buetow D (ed) The Biology of Euglena, vol 4. Academic Press, New York, in press
Hawley DK, McClure WR (1983) Compilation and analysis of Escherichia coli promoter DNA sequences. Nucl Acids Res 11:2237–2255
Helling RB, El-Gewely MR, Lomax MI, Baumgartner JE, Schwarzbach SD, Barnett WE (1979) Organization of the chloroplast ribosomal RNA genes of Euglena gracilis bacillaris. Mol Gen Genet 174:1–10
Hollingsworth MJ and Hallick RB (1982) Euglena gracilis chloroplast transfer RNA transcription units. Nucleotide sequence analysis of a tRNATyr-tRNAHis-tRNAMet-tRNATrp-tRNAGlu-tRNAGly gene cluster. J Biol Chem 257:12795–12799
Holmes WM, Platt T, Rosenberg M (1983) Termination of transcription in E. coli. Cell 32:1029–1032
Hutner SH, Bach MK, Ross GIM (1956) A sugar-containing basal medium for vitamin B12-assay with Euglena: application to body fluids. J Protozool 3:101–112
Ingraham JL, Maaløe O, Neidhardt FC (1983) Growth of the bacterial cell. Sinauer, Sunderland, Massachusetts, USA
Keller M, Burkard G, Bohnert HJ, Mubumbila M, Gordon K, Steinmetz A, Heiser D, Crouse EJ, Weil JH (1980) Transfer RNA genes associated with the 16S and 23S rRNA genes of Euglena chloroplast DNA. Biochem Biophys Res Commun 95:47–54
Keus RJA, Roovers DJ, Dekker AF, Groot GSP (1983) The nucleotide sequence of the 4.5S and 5S rRNA genes and flanking regions from Spirodela oligorhiza chloroplasts. Nucl Acids Res 11:3405–3410
Koller B, Delius H (1982) Parts of the sequence between the complete rRNA operons are repeated on either side of the extra 16S rRNA gene in chloroplast DNA of Euglena gracilis strain Z. FEBS Lett 140:198–202
Kumagai I, Pieler T, Subramanian AR, Erdman VA (1982) Nucleotide sequence and secondary structure analysis of spinach chloroplast 4.5S RNA. J Biol Chem 257:12924–12928
Kuntz M, Keller M, Crouse EJ, Burkard G, Weil JH (1982) Fractionation and identification of Euglena gracilis cytoplasmic and chloroplastic tRNAs and mapping of tRNA genes on chloroplast DNA. Curr Genet 6:63–69
Maaløe O, Kjeldgaard NO (1966) Control of macromolecular synthesis. W.A. Benjamin, New York
Machatt MA, Ebel J-P, Branlant C (1981) The 3′-terminal region of bacterial 23S ribosomal RNA: structure and homology with the 3′-terminal region of eukaryotic 28S rRNA and with chloroplast 4.5S rRNA. Nucl Acids Res 9:1533–1549
Maxam AM, Gilbert W (1980) Sequencing end-labeled DNA with base-specific chemical cleavages. In: Grossman L, Moldave K (eds) Methods in enzymology, vol 651. Academic Press, New York, pp 499–560
Miyata T, Kikuno R, Ohsima Y (1982) A pseudogene cluster in the leader region of the Euglena chloroplast 16S–23S rRNA genes. Nucl Acids Res 10:1771–1780
Orozco EM, Rushlow KE, Dodd JR, Hallick RB (1980) Euglena gracilis chloroplast ribosomal RNA transcription units. II. Nucleotide sequence homology between the 16S–23S ribosomal RNA spacer and the 16S ribosomal RNA leader regions. J Biol Chem 255:10997–11003
Pribnow D (1979) Genetic control signals in DNA. In: Goldberger RF (ed) Biological regulation and development I, Plenum Press, New York, pp 219–227
Pringsheim EG, Pringsheim O (1952) Experimental elimination of chromatophores and eye-spot in Euglena gracilis. New Phytol 51:65–76
Provasoli L, Hutner SH, Schatz A (1948) Streptomycin-induced chlorophyll-less races of Euglena. Proc Soc Exptl Biol Med 69:279–282
Roux E, Graf L, Stutz E (1983) Nucleotide sequence of ‘truncated tRNA operon’ of the Euglena gracilis chloroplast operon. Nucl Acids Res 11:1957–1969
Rubin CM, Schmid CW (1980) Pyrimidine-specific chemical reactions useful for DNA sequencing. Nucl Acids Res 8:4613–4619
Salser W (1977) Globin mRNA sequences: Analysis of base pairing and evolutionary implications. Cold Spring Harbor Symp Quant Biol 42:985–1002
Schmitt JM, Bohnert H-J, Gordon KHJ, Herrmann R, Bernardi G, Crouse EJ (1981) Compositional heterogeneity of the chloroplast DNAs from Euglena gracilis and Spinacia oleracea. Eur J Biochem 117:375–382
Steege DA, Graves MC, Spremulli LL (1982) Euglena gracilis chloroplast small subunit rRNA sequence and base pairing potential of the 3′ terminus, cleavage by colicin E3. J Biol Chem 257:10430–10439
Takaiwa F, Kusuda M, Sugiura M (1982) The nucleotide sequence of chloroplast 4.5S rRNA from a fern, Dryopteris acuminate. Nucl Acids Res 10:2257–2260
Takaiwa F, Sugiura M (1982) The complete nucleotide sequence of a 23S rRNA gene from tobaco chloroplasts. Eur J Biochem 124:13–19
Wildeman AG, Nazar RW (1980) Nucleotide sequence of wheat chloroplastid 4.5S ribonucleic acid. J Biol Chem 255: 11896–11900
Author information
Authors and Affiliations
Additional information
Communicated by G.A. O'Donovan
Rights and permissions
About this article
Cite this article
El-Gewely, M.R., Helling, R.B. & Dibbits, J.G.T. Sequence and evolution of the regions between the rrn operons in the chloroplast genome of Euglena gracilis bacillaris . Molec. Gen. Genet. 194, 432–443 (1984). https://doi.org/10.1007/BF00425555
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00425555