Summary
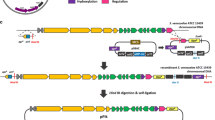
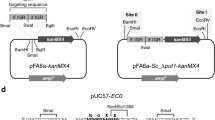
All five tryptophan biosynthetic genes of Saccharomyces cerevisiae were unified on plasmid pME554, which is based on 2 μm DNA and pBR322 sequences allowing for autonomous replication in yeast and E. coli. Homologous and heterologous expression of this artificial yeast TRP-gene cluster was studied. Plasmid pME554 allowed for nearly normal growth of a yeast strain bearing auxotrophic mutations in all five TRP-genes. The plasmid-borne genes TRP2 to TRP5 were expressed and regulated normally in the frame of the general control. Gene TRP1, carried on an EcoRI/BglII fragment lacking the ARS1 function, was expressed poorly and did not respond to the general control like the chromosomally-borne TRP1 gene.
Plasmid pME554 allowed for poor growth of E. coli strain W3110 tna − ΔtrpEA2 on minimal medium. Marked stimulation was observed, however, when anthranilic acid or indole were added. Accordingly, poor expression of the first Trp-enzyme anthranilate synthase and the last enzyme tryptophan synthase was found, whereas the other three genes were moderately well expressed in E. coli.
Similar content being viewed by others
Abbreviations
- bp:
-
basepair
- kb:
-
kilobase
References
Aebi M, Niederberger P, Hütter R (1982) Isolation of the TRP2 and the TRP3 genes of Saccharomyces cerevisiae by functional complementation in yeast. Curr Genet 5:39–46
Aebi M, Furter R, Prantl F, Niederberger P, Hütter R (1984) Structure and function of the TRP3 gene of Saccharomyces cerevisiae: Analysis of transcription, promoter sequence, and sequence coding for a glutamine amidotransferase. Curr Genet, in press
Beggs JD (1978) Transformation of yeast by a replicating hybrid plasmid. Nature 283:216–218
Broach JR, Guarascio VR, Jayaram M (1982) Recombination within the yeast plasmid 2 μm circle is site-specific. Cell 29:227–234
Crawford JP (1975) Gene rearrangements in the evolution of the tryptophan biosynthetic pathway. Bacteriol Rev 39:87–120
Dettwiler M, Kirschner K (1979) Tryptophan synthase from Saccharomyces cerevisiae is a dimer of two polypeptide chains of MJ 76000 each. Eur J Biochem 102:159–165
Dobson MJ, Futcher AB, Cox BS (1980) Loss of 2 μ DNA from Saccharomyces cerevisiae transformed with the chimaeric plasmid pJDB219. Curr Genet 2:201–205
Dobson MJ, Tuite MF, Mellor J, Roberts NA, King RM, Burke DC, Kingsman AJ, Kingsman SM (1983) Expression in Saccharomyces cerevisiae of human interferon-alpha directed by the TRP1 5′ region. Nucl Acid Res 11:2287–2302
Erhart E, Hollenberg CP (1981) Curing of Saccharomyces cerevisiae 2 μm DNA by transformation. Curr Genet 3:83–89
Falco SC, Rose M, Botstein D (1983) Homologous recombination between episomal plasmids and chromosomes in yeast. Genetics 105:843–856
Gerbaud C, Guérineau M (1980) 2 μm plasmid copy number in different yeast strains and repartition of endogenous and 2 μm chimeric plasmids in transformed strains. Curr Genet 1:219–228
Herbert D, Phipps PJ, Strange RE (1971) Chemical analysis of microbial cells. In: Norris JR, Ribbons DE (eds) Methods in microbiology, vol 5B. Academic Press, New York, p 209–240
Hsiao CL, Carbon J (1979) High frequency transformation of yeast by plasmids containing the cloned ARG4 gene. Proc Natl Acad Sci USA 76:3829–3833
Humphreys GO, Willshaw GA, Anderson ES (1975) A simple method for the preparation of large scale quantities of plasmid DNA. Biochim Biophys Acta 383:457–463
Miozzari G, Niederberger P, Hütter R (1978a) Tryptophan biosynthesis in Saccharomyces cerevisiae: control of the flux through the pathway. J Bacteriol 134:48–59
Miozzari G, Niederberger P, Hütter R (1978b) Permeabilization of microorganisms by Triton X-100. Anal Biochem 90:220–223
Mortimer RK, Schild D (1980) Genetic map of Saccharomyces cerevisiae. Microbiol Rev 44:519–571
Niederberger P, Miozzari G, Hütter R (1981) Biological role of the general control of amino acid biosynthesis in Saccharomyces cerevisiae. Mol Cell Biol 1:584–593
Niederberger P, Aebi M, Hütter R (1983) Influence of the general control of amino acid biosynthesis on cell growth and cell viability in Saccharomyces cerevisiae. J Gen Microbiol 129:2571–2583
Paluh JL, Zalkin H (1983) Isolation of Saccharomyces cerevisiae TRP3. J Bacteriol 153:345–349
Schürch A, Miozzari G, Hütter R (1974) Regulation of tryptophan biosynthesis in Saccharomyces cerevisiae: mode of action of 5-methyltryptophan and 5-methyltryptophan-sensitive mutants. J Bacteriol 117:1131–1140
Sherman F, Fink GR, Lukins HB (1970) Methods in yeast genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, p 51–53
Struhl K, Cameron JR, Davis RW (1976) Functional expression of eukaryotic DNA in Escherichia coli. Proc Natl Acad Sci USA 73:1471–1475
Tabak HF, Hecht B, Menke HH, Hollenberg CP (1979) The yeast mitochondrial small rRNA gene: Physical map, direction of transcription and absence of intervening sequences. Curr Genet 1:33–43
Tschumper G, Carbon J (1980) Sequence of a yeast DNA fragment containing a chromosomal replicator and the TRP1 gene. Gene 10:157–166
Vogel H, Bonner J (1956) Acetylornithinase of Escherichia coli. Partial purification and some properties. J Biol Chem 218:97–106
Walz A, Ratzkin B, Carbon J (1978) Control of expression of a cloned yeast (Saccharomyces cerevisae) gene (trp5) by a bacterial insertion element (IS2). Proc Natl Acad Sci USA 75:6172–6176
Yanofsky C (1981) Attenuation in the control of expression of bacterial operons. Nature 289:751–758
Author information
Authors and Affiliations
Additional information
Communicated by W. Arber
Rights and permissions
About this article
Cite this article
Niederberger, P., Aebi, M., Furter, R. et al. Expression of an artificial yeast TRP-gene cluster in yeast and Escherichia coli . Mol Gen Genet 195, 481–486 (1984). https://doi.org/10.1007/BF00341450
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00341450