Abstract
The noncoding DNA region of the chloroplast genome, flanked by the genes rbcL and psaI (ORF36), has been sequenced for seven species of the grass family (Poaceae). This region had previously been observed as a hotspot area for length mutations. Sequence comparison reveals that short duplications, probably resulting from slipped-strand mispairing, account for many small length differences between sequences but that major mutational hotspots are localized in three small areas, two of which show potential secondary structure. Mutation in one of these hotspots appears to be a result of more complex recombination events. All seven species contain a pseudogene for rpl23 and evidence is presented that this pseudogene is being maintained by gene conversion with the functional gene. Different transition/transversion biases and AT contents between the pseudogene and the surrounding noncoding sequences are noted. In the subfamily Panicoideae there is a deletion in which almost 1 kb of ancestral sequence, including the 3′ end of the rpl23 pseudogene, has been replaced by a non-homologous 60-base sequence of unknown origin. Two other deletions of almost the same region have occurred in the grass family. The deleted noncoding region has mutational and compositional properties similar to the rbcL coding sequence and the rpl23 pseudogene. The three independent deletions, as well as the pattern of mutation in the localized hotspots, indicate that such noncoding DNA may be misleading for studies of phylogenetic inference.
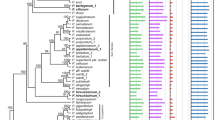
Similar content being viewed by others
References
Bowman CM, Barker RF, Dyer T (1988) Curr Genet 14:127–136
Clegg MT (1993) Proc Natl Acad Sci USA 90:363–367
Devereux J, Haeberli H, Smithies O (1984) Nucleic Acids Res 12:387–395
Doebley J, Durbin M, Golenberg EM, Clegg MT, Ma DP (1990) Evolution 44:1097–1108
Downie SR, Palmer JD (1992) Use of chloroplast DNA rearrangements in reconstructing plant phylogeny. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematics of Plants. Chapman and Hall, New York
Doyle JJ, Doyle JL (1987) Phytochem Bull 19:11–15
Felsenstein J (1985) Evolution 39:783–791
Gaut BS, Muse SV, Clark WD, Clegg MT (1992) J Mol Evol 35:292–303
Golenberg EM, Clegg MT, Durbin ML, Doebley J, Ma DP (1993) Mol Phylogenetics Evol (in press)
Hiratsuka J, Shimada H, Whittier R, Ishibashi T, Sakamoto M, Mori M, Kondo C, Honji Y, Sun C-R, Meng B-Y, Li Y-Q, Kanno A, Nishizawa Y, Hirai A, Shinozaki K, Sugiura M (1989) Mol Gen Genet 217:185–194
Levinson G, Gutman GA (1987) Mol Biol Evol 4:203–221
McIntosh L, Poulsen C, Bogorad L (1980) Nature 288:556–560
Ogihara Y, Terachi T, Sasakuma T (1988) Proc Natl Acad Sci USA 85:8573–8577
Ogihara Y, Terachi T, Sasakuma T (1991) Genetics 129:873–884
Ogihara Y, Terachi T, Sasakuma T (1992) Curr Genet 22:251–258
Palmer JD, Jansen RK, Michaels HJ, Chase MW, Manhart JR (1988) Ann Mo Bot Gard 75:1180–1206
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning. Cold Spring Harbor Laboratory, Cold Spring Harbour, New York.
Schaaper RM, Danforth BN, Glickman BW (1986) J Mol Biol 189:273–284
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchi-Shinozaki K, Ohto C, Torazawa K, Meng BY, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Suguira M (1986) EMBO J 5:2043–2049
Stern DB, Gruissem W (1987) Cell 51:1145–1157
Swofford DL (1991) PAUP: Phylogenetic Analysis Using Parsimony, version 3.0s. Computer program distributed by the Illinois Natural History Survey, Champaign, Illinois
Terachi T, Ogihara Y, Tsunewaki K (1987) Jpn J Genet 62:375–387
Thomas KM, Wood BJ, Bassett CL, Rawson JRY (1984) Curr Genet 8:291–297
Umesono K, Yasuhiko S, Takeuchi M, Chang Z, Aota S-i, Inokuchi H, Ozeki H (1988) J Mol Biol 203:281–298
Wolfe KH, Li W-H, Sharp PM (1987) Proc Natl Acad Sci USA 84:9054–9058
Zurawski G, Clegg MT (1987) Annu Rev Plant Physiol 38:391–418
Zurawski G, Clegg MT, Brown AHD (1984) Genetics 106:735–749
Author information
Authors and Affiliations
Additional information
Communicated by C. W. Birky, Jr.
Rights and permissions
About this article
Cite this article
Morton, B.R., Clegg, M.T. A chloroplast DNA mutational hotspot and gene conversion in a noncoding region near rbcL in the grass family (Poaceae). Curr Genet 24, 357–365 (1993). https://doi.org/10.1007/BF00336789
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00336789