Summary
About 300 revertants were derived from 44 cob - mutants, mapping in the structure coding regions (exon 1, 3, 4, 5, or 6) of the mitochondrial apocytochrome b gene in Saccharomyces cerevisiae, strain 777-3A. Most of the revertants could not be distinguished from the wild-type by means of physiological properties. Twenty-two revertants different in phenotype are described here in more detail.
The suppressor mutations (sup a) that compensate the primary cob - mutations (i.e., restore growth on glycerol) are mitochondrially inherited. They were localized in the same cob exon regions as the respective primary mutations, except for one revertant with a primary mutation in exon 6 and a suppressor, 4.2 map units distant, which may be located either in intron 5 or downstream in exon 6.
Of 21 suppressors 17 are closely coupled to the primary mutation with recombination frequencies of ≦0.1%–0.3%. An estimate predicts that in more than 80% of these revertants only one amino acid is altered at that point of the polypeptide corresponding to the cob - site in the gene.
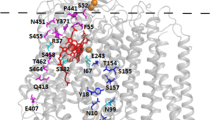
The most interesting revertant phenotypes are: (1) reduced growth rate on glycerol. The respective cob -/supa mutations are scattered over the whole cob region and cannot be correlated exclusively with special gene regions. (2) decreased cytochrome b content. The most extreme reductions (28% and 30% of wild-type level) were observed to be due to mutations located in the 5′ proximal part of exon 1. The highest percentage of revertants with decreased cytochrome b content was predominantly found mapping in exon 3. Complications in protoporphyrin attachment or the chelatase reaction were assumed to be the basic lesion causing reduced cytochrome b content, since in 10 out of 11 revertants examined the polypeptide is produced at wild-type level. (3) shifted maximum absorption wavelength of cytochrome b. The double mutations of the respective revertants map in the middle part of exon 1, in exon 4 and exom 5. The corresponding regions in the polypeptide presumably surround the heme group.
Similar content being viewed by others
References
Anziano PQ, Hanson DK, Mahler HR, Perlman PS (1982) Functional domains in introns: Trans-acting and cis-acting regions of intron 4 of the cob gene. Cell 30:925–932
Bechmann H, Haid A, Schweyen RJ, Mathews S, Kaudewitz F (1981) Expression of the “split gene” cob in yeast mtDNA. J Biol Chem 256:3525–3531
Burger G, Hofner E (1984) Cytochrome b of cob revertants in yeast. 2. Bioenergetic characterization of revertants with reduced content and shifted maximum absorption wavelength of cytochrome b. Eur J Biochem, in the press
Burger G, Wolf K (1981) Mitochondrially inherited resistance to Antimycin and Diuron in the petite negative yeast Schizosaccharomyces pombe. Mol Gen Genet 181:134–139
Burger G, Lang B, Bandlow W, Schweyen RJ, Backhaus B, Kaudewitz F (1976) Antimycin resistance in Saccharomyces cerevisiae. A new mutational site on the mitDNA conferring Antimycin resistance on the mitochondrial respiratory chain. Biochem Biophys Res Commun 72:1201–1208
Chance B (1958) The kinetics and inhibition of cytochrome components of the succinic oxidase system. III. Cytochrome b. J Biol Chem 233:1223–1229
Chevillotte-Brivet P, Meunier-Lemesle D (1980) Cytochrome b-565 in Saccharomyces cerevisiae: Use of mutants in the cob-box region of the mitochondrial DNA to study the functional role of this spectral species of cytochrome b. 2. Relation between energetic data and cytochrome b-565 content. Eur J Biochem 111:161–169
Claisse ML, Spyridakis A, Wambier-Kluppel ML, Pajot P, Slonimski PP (1978) Mosaic organisation and expression of the mitochondrial DNA region controlling cytochrome c reductase and oxidase. II. Analysis of proteins translated from the box region. In: Bacila M, Horecker BL, Stoppani AOH (eds) Biochemistry and Genetics of yeast. Pure and applied aspects. Academic Press, New York, p 369
Davies RW, Waring RB, Ray JA, Brown TA, Scazzocchio C (1982) Making ends meet: a model for RNA splicing in fungal mitochondria. Nature 300:719–724
De la Salle H, Jacq C, Slonimski PP (1982) Critical sequences within mitochondrial introns: pleiotropic mRNA maturase and cis-dominant signals of the box intron controlling reductase and oxidase. Cell 28:721–732
Dujardin G, Pajot P, Groudinsky O, Slonimski PP (1980) Long range control circuits within mitochondria and between nucleus and mitochondria. I. Methodology and phenomenology of suppressors. Mol Gen Genet 179:469–482
Groudinsky O, Dujardin G, Slonimski PP (1981) Long range control circuits within mitochondria and between nucleus and mitochondria. II. Genetic and biochemical analyses of suppressors which selectively alleviate the mitochondrial intron mutations. Mol Gen Genet 184:493–503
Haid A, Schweyen RJ, Bechmann H, Kaudewitz F, Solioz M, Schatz G (1978) The mitochondrial cob region in yeast codes for apocytochrome b and is mosaic. Eur J Biochem 94:451–464
Hanson DK, Miller DH, Mahler HR, Alexander NJ, Perlman PS (1979) Regulatory interaction between mitochondrial genes. II. Detailed characterization of novel mutants mapping within one cluster in the cob2 region. J Biol Chem 254:2480–2490
Helmer-Citterich M, Morelli G, Macino G (1983) Nucleotide sequence and intron structure of the apocytochrome b gene of Neurospora crassa mitochondria. The EMBO J 2:1235–1242
Jacq C, Pajot P, Lazowska J, Dujardin G, Claisse M, Groudinsky O, de la Salle H, Grandchamp C, Labouesse M, Gargouri A, Guiard B, Spyridakis A, Dreyfus M, Slonimski PP (1982) Role of introns in the yeast cytochrome b gene: Cis- and trans-acting signals, intron manipulation, expression, and intergenic communications. In: Slonimski PP, Borst P, Attardi G (eds) Mitochondrial genes. Cold Spring Harbor Laboratory, New York, p 155–183
Kotylak Z, Slonimski PP (1977) Mitochondrial mutants isolated by a new screening method based upon the use of the nuclear mutation op1. In: Bandlow W, Schweyen RJ, Wolf K, Kaudewitz F (eds) Mitochondria 1977. Genetics and biogenesis of mitochondria. Walter de Gruyter, Berlin, New York, p 83–89
Lamb MR, Anziano PQ, Glaus KR, Hanson DK, Klapper HJ, Perlman PS, Mahler HR (1983) Functional domains in introns. A. Processing intermediates in cis- and transacting mutants in the penultimate intron of the mitochondrial gene for cytochrome b. J Biol Chem 258:1991–1999
Lancashire WE, Mattoon JR (1979) Cytoduction: A tool for mitochondrial genetic studies in yeast. Utilization of the nuclearfusion mutation kar1-1 for transfer of drug r and mit genomes in Saccharomyces cerevisiae. Mol Gen Genet 170:333–344
Lazowska J, Jacq C, Slonimski PP (1980) Sequence of introns and flanking exons in wild-type and box3 mutants of cytochrome b reveals an interlaced splicing protein coded by an intron. Cell 22:333–348
Mahler HR, Hanson DK, Lamb MR, Perlman PS, Anziano PQ, Glaus KR, Haldi ML (1982) Regulatory interactions between mitochondrial genes: Expressed introns — their function and regulation. In: Slonimski, PP, Borst P, Attardi G (eds) Mitochondrial genes. Cold Spring Harbor Laboratory, New York, p 185–200
Meunier-Lemesle D, Chevillotte-Brivet P, Pajot P (1980) Cytochrome b-565 in Saccharomyces cerevisiae: Use of mutants in the cob-box region of the mitochondrial DNA to study the functional role of this spectral species of cytochrome b. 1. Measurements of cytochromes b-562 and b-565 and selection of revertants devoid of cytochrome b-565. Eur J Biochem 111:151–159
Nobrega FG, Tzagoloff A (1980) Assembly of the mitochondrial membrane system: DNA sequence and organisation of the cytochrome b gene in Saccharomyces cerevisiae D273-10B. J Biol Chem 255:9828–9837
Pajot P, Wambier-Kluppel ML, Slonimski PP (1977) Cytochrome c reductase and cytochrome oxidase formation in mutants and revertants in the “box” region of mitochondrial DNA. In: Bandlow W, Schweyen RJ, Wolf K, Kaudewitz F (eds) Mitochondria 1977. Genetics and biogenesis of mitochondria. Walter de Gruyter, Berlin, New York, p 173–183
Sato N, Ohnishi T, Chance B (1972) Spectral study of b cytochromes in yeast mitochondria and intact cells. Biochim Biophys Acta 275:288–297
Schmelzer C, Haid A, Grosch G, Schweyen RJ, Kaudewitz F (1981) Pathways of transcript splicing in yeast mitochondria. Mutations in intervening sequences of the split gene cob reveal a requirement for intervening sequence-encoded products. J Biol Chem 256:7610–7619
von Jagow G, Sebald W (1980) b-type cytochromes. Annu Rev Biochem 49:281–314
von Jagow G, Engel WD, Schägger H, Machleidt W, Machleidt I (1981) On the mechanism of proton translocation linked to electron transfer at energy conversion site 2. In: Palmieri F, Quagliarello E, Siliprandi N, Slater EC (eds) Vectorial reactions in electron and ion transport in mitochondria and bacteria. Elsevier/North-Holland Biomedical press, Amsterdam, p 149–161
Waring RB, Davies RW, Lee S, Grisi E, McPhail Berks M, Scazzocchio C (1981) The mosaic organization of the apocytochrome b gene of Aspergillus nidulans revealed by DNA sequencing. Cell 27:4–11
Weiss-Brummer B, Holl J, Schweyen RJ, Rödel G, Kaudewitz F (1983) Processing of yeast mitochondrial RNA: Involvement of intramolecular hybrids in splicing of cob intron 4 RNA by mutation and reversion. Cell 33:195–202
Author information
Authors and Affiliations
Additional information
Communicated by W. Gajewski
Rights and permissions
About this article
Cite this article
Burger, G. Cytochrome b of cob revertants in yeast. Mol Gen Genet 196, 158–166 (1984). https://doi.org/10.1007/BF00334109
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00334109