Summary
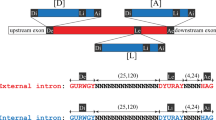
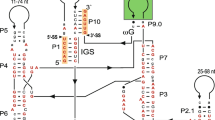
The overlapping ND4L and ND5 genes of Neurospora crassa mitochondria are interrupted by one and two intervening sequences, respectively, of about 1,490, 1,408 and 1,135 bp in length. All three intervening sequences are class I introns and as such have the potential to fold into the conserved secondary structure that has been proposed for the majority of fungal mitochondrial introns. They contain long open reading frames (ORFs; from 306 to 425 codons long) that are continuous and in frame with the upstream exon sequences. These ORFs contain the conserved decapeptide-encoding sequences that are characteristic of the ORFs present in most class I intron. Extensive homology exists among the ORFs encoded by the ND4L intron, ND5 intron 1, and the second intron of the N. crassa oli2 gene. Also, internal repeats of about 130 amino acid residues are present twice in each of these three ORFs, suggesting that a duplication event may have occurred in the formation of these ORFs. The ND4L intron shares extensive homology (at the levels of both primary and proposed secondary structures) with the self-splicing intervening sequence present in the Tetrahymena nuclear rRNA gene. This homology includes but is not limited to the core secondary structure, as peripheral structural elements are also conserved in the two introns.
Similar content being viewed by others
Abbreviations
- bp:
-
base pairs
- rRNA:
-
ribosomal RNA
- ND:
-
NADH dehydrogenase
- ORF:
-
open reading frame
- IGS:
-
internal guide sequence
References
Burke JM, Irvine KD, Kaneko KJ, Kerker BJ, Oettgen AB, Tierney WM, Williamson CL, Zaug AJ, Cech TR (1986) Role of conserved sequence elements 9L and 2 in self-splicing of the Tetrahymena ribosomal RNA precursor. Cell 45:167–176
Cech TR, Tanner NK, Tinoco I, Weir BR, Zuker M, Perlman PS (1983) Secondary structure of the Tetrahymena ribosomal RNA intervening sequence: structural homology with fungal mitochondrial intervening sequences. Proc Natl Acad Sci USA 80:3903–3907
Colleaux L, d'Auriol L, Betermier M, Cottarel G, Jacquier A, Galibert F, Dujon B (1986) Universal code equivalent of a yeast mitochondrial intron reading frame is expressed into E. coli as a specific double strand endonuclease. Cell 44:521–533
Davies RW, Waring RB, Ray JA, Brown TA, Scazzocchio C (1982) Making ends meet: a model for RNA splicing in fungal mitochondria. Nature 300:719–724
de Vries H, de Jonge JC, Schrage C (1985) The Neurospora, mitochondrial stopper mutants, E35, lacks two protein genes indispensable for the formation of complexes I, III and IV. In: Quagliarello E, Slater EC, Palmieri F, Saccone C, Kroon AM (eds) Achievements and perspectives of mitochondrial research, vol. II: Biogenesis. Elsevier Science Publishers, Amsterdam, New York, Oxford, pp 285–292
Dujon B (1980) Sequence of the intron and flanking exons of the mitochondrial 21S rRNA gene of yeast strains having different alleles at the ω and rib-1 loci. Cell 20:185–197
Friedman EY, Rosbach M (1977) The synthesis of high yields of full-length reverse transcripts of globin mRNA. Nucleic Acids Res 4:3455–3471
Garriga G, Lambowitz AM (1984) RNA splicing in Neurospora mitochondria: self-splicing of a mitochondrial intron in vitro. Cell 38:631–641
Grivell LA, Borst P (1982) Mitochondrial mosaics — maturases on the move. Nature 298:703–704
Hensgens LAM, Bonen L, de Haan M, van der Horst G, Grivell LA (1983) Two intron sequences in yeast mitochondrial COX1 gene: homology among URF-containing introns and strain-dependent variation in flanking exons. Cell 32:379–389
Holl J, Rodel G, Schweyen RJ (1985) Suppressor mutations identify box9 as a central nucleotide sequence in the highly ordered structure of intron RNA in yeast mitochondria. EMBO J 4:2081–2085
Kan NC, Gall JG (1982) The intervening sequence of the ribosomal RNA gene is highly conserved between two Tetrahymena species. Nucleic Acids Res 10:2809–2822
Kruger K, Grabowski PJ, Zaug AJ, Sands J, Gottschling DE, Cech TR (1982) Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31:147–157
Lazowska J, Jacq C, Slonimski PP (1981) Splice points of the third intron in the yeast mitochondrial cytochrome b gen. Cell 27:12–14
Maxam AM, Gilbert W (1980) Sequencing end-labelled DNA with base-specific chemical cleavages. Methods Enzymol 65:499–560
Michel F, Cummings DJ, (1985) Analysis of class I introns in a mitochondrial plasmid associated with senescence of Podospora anserina reveals extraordinary resemblance to the Tetrahymena ribosomal intron. Curr Genet 10:69–79
Michel F, Dujon B (1983) Conservation of RNA secondary structures in two intron families including mitochondrial-, chloroplast-and nuclear-encoded members. EMBO J 2:33–38
Michel F, Jacquier A, Dujon B (1982) Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie 64:867–881
Morelli G, Macino G (1984) Two intervening sequences in the ATPase subunit 6 gene of Neurospora crassa: a short intron (93 bp) and a long intron that is stable after excision. J Mol Biol 178:491–507
Nelson MA, Macino G (1985) Gene organization and expression in Neurospora crassa mitochondria. In: Quagliariello E, Slater EC, Palmineri F, Saccone C, Kroon AM (eds) Achievements and perspectives in mitochondrial research, vol. II: Biogenesis. Elsevier, Amsterdam, New York, Oxford, pp 293–304
Nelson MA, Macino G (1987) Structure and expression of the overlapping ND4L and ND5 genes of Neurospora crassa mitochondria. Mol Gen Genet 206:307–317
Tabak HF, Van der Horst G, Osinga KA, Arnberg AC (1984) Splicing of large ribosomal precursor RNA and processing of intron RNA in yeast mitochondria. Cell 39:623–629
Van der Horst G, Tabak HF (1985) Self-splicing of yeast mitochondrial ribosomal and messenger RNA precursors. Cell 40:759–766
Waring RB, Davies RW (1984) Assessment of a model for intron RNA secondary structure relevant to RNA self-splicing — a review. Gene 28:277–291
Waring RB, Davies RW, Lee S, Grisi E, McPhail Berks M, Scazzocchio C (1981) The mosaic organization of the apocytochrome b gene of Aspergillus nidulans revealed by DNA sequencing. Cell 27:4–11
Waring RB, Davies RW, Scazzocchio C, Brown TA (1982) Internal structure of a mitochondrial intron of Aspergillus nidulans. Proc Natl Acad Sci USA 79:6332–6336
Waring RB, Ray JA, Edwards SW, Scazzocchio C, Davies RW (1985) The Tetrahymena rRNA intron self-splices in E. coli: in vivo evidence for the importance of key base-paired regions of RNA for RNA enzyme function. Cell 40:371–380
Waring RB, Towner P, Minter SJ, Davies RW (1986) Splice-site selection by a self-splicing RNA of Tetrahymena. Nature 321:133–139
Weiss-Brummer B, Holl J, Schweyen RJ, Rodel G, Kaudewitz F (1983) Processing of yeast mitochondrial RNA: Involvement of intramolecular hybrids in splicing of cob intron 4 RNA by mutation and reversion. Cell 33:195–202
Author information
Authors and Affiliations
Additional information
Communicated by R.G. Herrmann
Rights and permissions
About this article
Cite this article
Nelson, M.A., Macino, G. Three class I introns in the ND4L/ND5 transcriptional unit of Neurospora crassa mitochondria. Mol Gen Genet 206, 318–325 (1987). https://doi.org/10.1007/BF00333590
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00333590