Summary
Mutants pWS10, pWS11, and pWS12 were derived from an, IncFI group plasmid ColV-K94 by the insertion of a transposon Tn903 (Kmr). These plasmids were all approximately 130 kb in length. The plasmid pWS12 resembled the wild type ColV-K94 in transmissibility, incompatibility and stable maintenance. Cells harboring pWS11 were poor conjugal donors but resistant to the same level of kanamycin as pWS12 containing hosts. In contrast, pWS10 conferred a higher resistance to kanamycin and exhibited reduced incompatibility properties in comparison with pWS12. The higher drug resistance associated with pWS10 appeared to be a consequence of an increase in its copy number and the generation of miniplasmids of varying sizes.
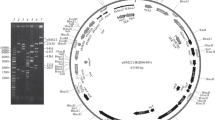
Electron microscope analysis of the copy mutant pWS10 revealed that Tn903 was located at a site adjacent to a region 32.6F to 35.3F. The latter region appears to be the primary replicon of ColV-K94 and is homologous with the secondary replicon of F. The insertional mutagenesis with Tn903 brought about an extensive DNA rearrangement including the duplication and translocation of the stems of two inverted repeat structures. The DNA alterations of pWS10 were distinguishable through comparison of its EcoRI digestion patterns with those of pWS11 and pWS12.
Similar content being viewed by others
References
Binns MM, Davis DL, Hardy KG (1979) Cloned fragments of the plasmid ColV, I-K94 specifying virulence and serum resistance. Nature 279:778–781
Cohen SN, Chang ACY, Hsu L (1972) Nonchromosomal antibiotic resistance in bacteria: Genetic transformation of Escherichia coli by R factor DNA. Proc Natl Acad Sci USA. 69:2110–2114
Davis RW, Simon M, Davidson N (1971) Electron microscope heteroduplex methods for mapping regions of base sequence homology in nucleic acids. In Methods in enzymology, vol 21D. Academic Press Inc, New York, p 413–428
Goebel W, Bonewald R (1975) Class of small multicopy, plasmids originating from the mutant antibiotic resistance factor R1 drd-19B2. J Bacteriol 123:658–665
Hirota Y (1960) The effect of acridine dyes on maying type factors in Escherichia coli. Proc Natl Acad Sci USA 46:57–64
Kahn ML, Figurski D, Ito L, Helinski DR (1979) Essential regions for replication of a stringent and a relaxed plasmid in Escherichia coli. Cold Spring Harbor Symp Quant Biol 43:99–103
Kahn P, Helinski DR (1964) Relationship between colcinogenic factors E1 and V and a F factor in Escherichia coli. J Bacteriol 88:1573–1579
Kopecko DJ, Brevet J, Cohen SN (1976) Involvement of multiple translocating DNA segments and recombinational hot spots in the structural evolution of bacterial plasmids. J Mol Biol 108:333–360
Lane HED (1981) Replication and incompatibility of F and plasmids in the IncF1 group. Plasmid 5:100–126
Lane D, Gardner RC (1979) Second EcoRI fragment of F capable of self-replication. J Bacteriol 139:141–151
Lovett MA, Helinski DR (1976) Methods for the isolation of the replication region of a bacterial replicon: Construction of a mini-F′ Km plasmid. J Bacteriol 127:982–987
Manis JJ, Kline BC (1978) F plasmid incompatibility and copy number genes. Their map location and interactions. Plasmid 1:492–507
Mickel S, Bauer W (1976) Isolation by tetracycline, selection of small plasmids derived from R-factor R12 in Escherichia coli K-12. J Bacteriol 127:644–655
Molin S, Nordström K (1980) Control of plasmid R1 replication: Functions involved in replication, copy number control, incompatibility and switch-off of replication. J Bacteriol 141:111–120
Molin S, Staugaard P, Uhlin BE, Gustafsson P, Nordström K (1979) Clustering of genes involved in replication, copy number control, incompatibility of the plasmid, R1drd-19. J Bacteriol 138:70–79
Monk M, Kinross J (1972) Conditional lethality of recA and recB derivatives of a strain of E. coli K-12 with a temperature-sensitive DNA polymerase I. J Bacteriol 109:971–978
Morris CF, Hashimoto H, Mickel S, Rownd R (1974) Round of replication mutant of a drug resistance factor. J Bacteriol 118:855–866
Ohstubo E, Rosenbloom M, Schrempf H, Goebel W, Rosel J (1978) Site specific recombination involved in the generation of small plasmids Mol Gen Genet 159:131–141
Palchaudhuri S, Maas WK (1977) Physical mapping of a DNA sequence common to plasmids of incompatibility group F1. Proc Natl Acad Sci USA 74:1190–1194
Palchaudhuri S, Maas WK, Ohtsubo E (1976) Fusion of two F-prime factors in Escherichia coli studied by electron microscope heteroduplex analysis. Mol Gen Genet 146:215–231
Palchaudhuri SR, Iyer VN (1971) Compatibility between two F prime factors in an Escherichia coli strain bearing a chromosomal mutation affection DNA synthesis. J Mol Biol 57:319–333
Palchaudhuri S, Mitra G (1981) Replication, incompatibility and acridine orange curing. In: Levy S, Clowes R, Koenig E (eds) Molecular biology, pathogencity and ecology of bacterial plasmids. Plenum publishing Corp, New York, p 631–632
Sharp PA, Cohen SN, Davidson N (1973) Electron microscope heteroduplex studies of sequence relations among plasmid of Escherichia coli. II. Structure of drug resistance (R) factors and F factors. J Mol Biol 75:235–255
So M, Heffron, F, Falkow S (1978) Method for the genetic labelling of cryptic plasmids. J Bacteriol 133:1520–1523
Starlinger P (1980) IS elements and transposons. Plasmid 3:241–259
Stougaard P, Molin S, Nordström K (1979) Plasmid R1 in Salmonella typhimurium: Molecular instability and gene dosage effects. Plasmid 2:589–597
Tolun A, Helinski DR (1981) Direct repeats of the F plasmid incC region express F incompatibility. Cell 24:687–694
Timmis KN, Danbara H, Brady G, Lurz R (1981) Inheritance functions of group IncFII transmissible antibiotic resistance plasmids. Plasmid 5:53–75
Uhlin BE, Nordström K (1975) Plasmid incompatibility and control of replication: Copy mutants of the R-factor R1 in Escherichia coli K12. J Bacteriol 124:641–649
Wechsler J, Kline BC (1980) Mutation and identification of the F plasmid locus determining resistance to acridine orange curing. Plasmid 4:276–280
Womble DD, Taylor DP, Rownd RH (1977) Method for obtaining more accurate covalently closed circular plasmid-to-chromosome ratios from bacterial lysates by dye-buoyant density centrifugation. J Bacteriol 130:148–153
Author information
Authors and Affiliations
Additional information
Communicated by G.A. O'Donovan
Rights and permissions
About this article
Cite this article
Mitra, G., Palchaudhuri, S. Genetic structure and stability of a copy number mutant of IncFI group plasmid ColV-K94 in Escherichia coli K-12. Molec Gen Genet 193, 349–357 (1984). https://doi.org/10.1007/BF00330692
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00330692