Abstract
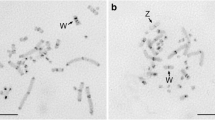
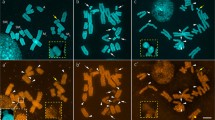
The two closely related species Apodemus sylvaticus and Apodemus flavicollis (Muridae) differ in the distribution of their heterochromatin. Two major repetitive sequences known to occur in both species were isolated from A. flavicollis after digestion of total nuclear DNA with the restriction enzymes HindIII and EcoRI respectively and characterized in both species by filter hybridisation and in situ hybridisation to metaphase chromosomes. The EcoRI clone detects a dispersed repetitive sequence family in the genome of both species. Southern blot hybridisation with the HindIII satellite DNA probe reveals major similarities and minor differences in the two species. In situ hybridisation with the HindIII probe labels all chromosomes of A. flavicollis exclusively in the centromeric heterochromatin, whereas in A. sylvaticus several autosomes are also labelled distally. The labelling patterns correspond to the distribution of heterochromatin in the two species. It is concluded that the additional distal heterochromatin of A. sylvaticus contains similar sequences to those of the centromeric heterochromatin of both species. The distal heterochromatin in A. sylvaticus most likely evolved by transposition and amplification of centromeric satellite DNA elements, after the separation of the two species.
Similar content being viewed by others
References
Adolph S, Bartram CR, Hameister H (1987) Mapping of the oncogenes Myc, Sis, and int-1 to the distal part of mouse chromosome 15. Cytogenet Cell Genet 44:65–68
Ambros PF, Matzke MA, Matzke AJM (1986) Detection of a 17 kb unique sequence (T-DNA) in plant chromosomes by in situ hybridization. Chromosoma 94:11–18
Arrighi FE, Hsu TC (1971) Localization of heterochromatin in human chromosomes. Cytogenetics 10:81–86
Bekasova TS, Vorontsov NN, Korobitsyna KV, Korablev VK (1980) B-Chromosomes and comparative karyology of the genus Apodemus. Genetica 52/53:33–43
Brown SDM, Dover G (1981) Organization and evolutionary progress of a dispersed repetitive family of sequences in widely separated rodent genomes. J Mol Biol 150:441–466
Caspersson T, Farber S, Foley GE, Kudynowski J, Modest EJ, Simonsson E, Wagh U, Zech L (1968) Chemical differentiation along metaphase chromosomes. Exp Cell Res 49:219–222
Cooke HJ (1975) Evolution of the long range structure of satellite DNAs in the genus Apodemus. J Mol Biol 94:87–99
De la Chapelle A, Schröder J, Selander RK, Stenstrand K (1973) Differences in DNA composition along mammalian metaphase chromosomes. Chromosoma 42:365–382
Engel W, Vogel W, Voiculescu I, Ropers H-H, Zenzes M-T, Bender K (1973) Cytogenetic and biochemical differences between Apodemus sylvaticus and Apodemus flavicollis, possibly responsible for the failure to interbreed. Comp Biochem Physiol 44B:1165–1173
Gamperl R, Ehmann Ch, Bachmann K (1982) Genome size and heterochromatin variation in rodents. Genetica 58(3):199–212
Habenicht H (1978) Cytogenetischer Vergleich der drei Apodemus-Arten A. sylvaticus, A. flavicollis und A. agrarius unter morphologischen und ökologischen Aspekten. Dissertation, Graz
Howell WM, Black DA (1980) Controlled silverstaining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia 36:1014–1015
Jewell PA, Fullagar PI (1965) Fertility among races of field mouse (Apodemus sylvaticus) and their failure to form hybrids with yellow necked mouse (Apodemus flavicollis). Evolution 19:175–181
Koop BF, Baker RJ, Haiduk MW, Engstrom MD (1984) Cladistical analysis of primitive G-band sequences for the karyotype of the ancestor of the Cricetidae complex of rodents. Genetica 64:199–208
Kral B (1970) Chromosome studies in two subgenera of the genus Apodemus. Zool Listy 19(2):119–134
Manuelidis L, Langer-Safer PR, Ward DC (1982) High-resolution mapping of satellite DNA using biotin-labeled DNA probes. J Cell Biol 95:619–625
Saint-Girons MCh (1962) Apropos de l'hybriditation eventuelle entre mulot gris, Apodemus sylvaticus (Linnaeus 1758) et mulot fauve, Apodemus flavicollis (Melchior 1834). Säugetierkd Mitt 10:25
Schulz WA, Gais G (1989) Constitutive c-myc expression enhances proliferation of differentiating F9 teratocarcinoma cells. Biochim Biophys Acta 1013:125–132
Schulz WA, Crawford N, Locker J (1988) Albumin and a-fetoprotein gene expression and DNA methylation in rat hepatoma cell lines. Exp Cell Res 174:433–447
Schweizer D, Tohidast-Akrad M, Strehl D, Dann O (1987) Diverse fluorescent staining of human heterochromatin by the isomeric DAPI derivates D 288/45 and D288/48. Ann Univ Sarav Med-Suppl 7:285–290
Seabright M (1971) A rapid banding technique for human chromosomes. Lancet 2:971–972
Soldatovic B, Savic I, Seth P, Reichenstein H, Tolksdorf M (1975) Comparative karyological study of the genus Apodemus (Kaup 1829). Acta Vet (Beograd) 25(1):1–10
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306
Vogel W, Autenrieth M, Speit G (1986) Detection of bromdeoxyuridine-incorporation in mammalian chromosomes by a bromdeoxyuridine-antibody. Hum Genet 72:129–132
Vujosevic M, Rimsa D, Zivcovic S (1984) Patterns of G- and C- bands distribution on chromosomes of three Apodemus species. Z Säugetierkd 49:234–238
Walker PMB (1968) How different are the DNAs from related animals? Nature 219:228–232
Zimmermann K (1957) Sind Gelbhalsmaus und Waldmaus miteinander kreuzbar? Z Säugetierkd 22:214–217
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hirning, U., Schulz, W.A., Just, W. et al. A comparative study of the heterochromatin of Apodemus sylvaticus and Apodemus flavicollis . Chromosoma 98, 450–455 (1989). https://doi.org/10.1007/BF00292791
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00292791