Abstract
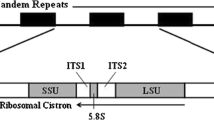
The arrangement of the 26S RNA and 18S RNA sequences of the ribosomal DNA (rDNA) from the sea urchin Lytechinus variegatus was investigated by an electron microscopic analysis of R-loops formed between the ribosomal RNA genes and the mature ribosomal RNAs. Ninety-eight percent of observed molecules contained R-loops clearly seen as a three-stranded complex. The size of DNA complementary to mature cytoplasmic 18S and 26S ribosomal RNA (rRNA) was calculated by measuring the double-strand (ds) and single-strand (ss) part of the R-loops separately. The values for the 18S R-loop are 1.75±0.24 kb1 (ss) and 1.56±0.23 kb (ds). The 26S R-loop is 3.34±0.39 kb (ss) and 3.33±0.33 kb (ds). These measurements agree fairly well with the rRNA sizes measured on denaturing sucrose density gradients: 3.23±0.22 kb for the 26S and 1.93±0.10 kb for 18S. The short spacer between the 18S and 26S R-loops is 1.03±0.24 kb and the longer spacer is 5.36±0.53 kb. In long molecules a repeating pattern was observed. The average length of an rDNA repeat unit is 11.33±0.64 kb when computed using double-strand R-loop measurements and 11.50±0.72 when computed using R-loop single-strand lengths.
Similar content being viewed by others
Abbreviations
- kb:
-
kilobases, 1000 bases of RNA or single-strand DNA, and kilobase pairs, 1000 base pairs of duplex DNA or DNA/RNA hybrid
- EDTA:
-
ethylenediaminetetraacetate
- SSC:
-
0.15 M NaCl, 0.015 M sodium citrate
- PIPES:
-
piperazine-N,N′-bis (2-ethanesulfonic acid)-Na1.4
References
Brown, D.D., Littna, E.: RNA synthesis during the development of Xenopus laevis, the South African clawed toad. J. molec. Biol. 8, 669–687 (1964)
Davis, R.W., Simon, M., Davidson, N.: Electron microscope heteroduplex methods for mapping regions of base sequence homology in nucleic acids. In: Methods in enzymology (L. Grossman and K. Moldave, eds.), Vol. XXI, pp. 413–428. New York: Academic Press 1971
Frankel, G., Cockburn, A.F., Kindle, K.L., Firtel, R.A.: Organization of the ribosomal RNA genes of Dictyostelium discoideum. Mapping of the transcribed region. J. molec. Biol. 109, 539–558 (1977)
Gillespie, D., Spiegelman, S.: A quantitative assay for DNA-RNA hybrids with DNA immobilized on a membrane. J. molec. Biol. 12, 829–842 (1965)
Joseph, D.R., Stafford, D.W.: Purification of sea urchin ribosomal RNA genes with a single-strand specific nuclease. Biochim. biophys. Acta (Amst.) 418, 167–174 (1976)
Kirby, K.S., Cook, E.A.: Isolation of deoxyribonucleic acid from mammalian tissues. Biochem. J. 104, 254–257 (1967)
Klukas, C.K., Dawid, I.E.: Characterization and mapping of mitochondrial ribosomal RNA and mitochondrial DNA in Drosophila melanogaster. Cell 9, 615–625 (1976)
Loening, U.E.: Molecular weights of ribosomal RNA in relation to evolution. J. molec. Biol. 38, 355–365 (1968)
Miller, O.L., Jr., Beatty, B.R.: Visualization of nucleolar genes. Science 164, 955–957 (1969)
Patterson, J.B., Stafford, D.W.: Characterization of sea urchin ribosomal satellite deoxyribonucleic acid. Biochemistry 10, 2775–2778 (1971)
Pellegrini, M., Manning, J., Davidson, N.: Sequence arrangement of the rDNA of Drosophila melanogaster. Cell 10, 213–224 (1977)
Perler, F.: A fine structure physical and functional map of the ribosomal RNA genes in the sea urchin, Lytechinus variegatus. Ph. D. Thesis. University of North Carolina, Chapel Hill, N.C. (1977)
Sanger, F., Air, G.M., Barrell, B.G., Brown, N.L., Coulson, A.R., Fiddes, J.C., Hutchison, C.A., III, Slocombe, P.M., Smith, M.: Nucleotide sequence of bacteriophage ΦX174 DNA. Nature (Lond.) 265, 687–695 (1977)
Stafford, D.W., Guild, W.R.: Satellite DNA from sea urchin sperm. Exp. Cell Res. 55, 347–350 (1969)
Suzuki, Y., Gage, L.P., Brown, D.D.: The genes for silk fibroin in Bombyx mori. J. molec. Biol. 70, 637–649 (1972)
Sy, J., McCarty, K.S.: Ribosomal RNA of Arbacia punctulata. Biochim. biophys. Acta (Amst.) 166, 571–574 (1968)
Thomas, M., White, R.L., Davis, R.W.: Hybridization of RNA to doublestranded DNA: Formation of R-loops. Proc. nat. Acad. Sci. (Wash.) 73, 2294–2298 (1976)
Wartell, R.M., Larson, J.E., Wells, R.D.: Netropsin. A specific probe for A-T regions of duplex deoxyribonucleic acid. J. biol. Chem. 249, 6719–6731 (1974)
Weinberg, R.A., Loening, U.E., Willems, M., Penman, S.: Acrylamide gel electrophoresis of HeLa cell nucleolar RNA. Proc. nat. Acad. Sci. (Wash.) 58, 1088–1095 (1967)
Wellauer, P.K., Dawid, I.B.: Secondary structure maps of RNA: Processing of HeLa ribosomal RNA. Proc. nat. Acad. Sci. (Wash.) 70, 2827–2831 (1973)
Wellauer, P.K., Dawid, I.E.: Secondary structure maps of ribosomal RNA and DNA. I. Processing of Xenopus laevis ribosomal RNA and structure of single-stranded ribosomal DNA. J. molec. Biol. 89, 379–395 (1974)
Wellauer, P.K., Dawid, I.B.: The structural organization of ribosomal DNA in Drosophila melanogaster. Cell 10, 193–212 (1977)
Wellauer, P.K., Dawid, I.B., Kelley, D.E., Perry, R.P.: Secondary structure maps of ribosomal RNA. II. Processing of mouse L-cell ribosomal RNA and variations in the processing pathway. J. molec. Biol. 89, 397–407 (1974a)
Wellauer, P.K., Reeder, R.H., Carroll, D., Brown, D.D., Deutch, A., Higashinakagawa, T., Dawid, I.E.: Amplified ribosomal DNA from Xenopus laevis has heterogeneous spacer lengths. Proc. nat. Acad. Sci. (Wash.) 71, 2823–2827 (1974b)
Wells, R.D., Larson, J.E.: Studies on the binding of actinomycin D to DNA and DNA model polymers. J. molec. Biol. 49, 319–342 (1970)
Wensink, P.C., Brown, D.D.: Denaturation map of the ribosomal RNA of Xenopus laevis. J. molec. Biol. 60, 235–247 (1971)
White, R.L., Hogness, D.S.: R-Loop mapping of the 18S and 28S sequences in the long and short repeating units of Drosophila melanogaster rDNA. Cell 10, 177–192 (1977)
Wilson, F.E., Blin, N., Stafford, D.W.: A denaturation map of sea urchin ribosomal DNA. Chromosoma (Berl.) 58, 247–253 (1976)
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wilson, F.E., Blin, N. & Stafford, D.W. Arrangement of the rRNA sequences in the rDNA of Lytechinus variegatus . Chromosoma 65, 373–381 (1978). https://doi.org/10.1007/BF00286416
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00286416