Summary
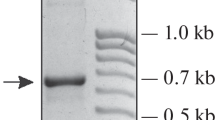
Ribosomal DNA (rDNA) repeats of the plant-parasitic nematode Meloidogyne arenaria are heterogeneous in size and appear to contain 5S rRNA gene sequences. Moreover, in a recA + bacterial host, plasmid clones of a 9 kb rDNA repeat show deletion events within a 2 kb intergenic spacer (IGS), between 28S and 5S DNA sequences. These deletions appear to result from a reduction in the number of tandem 129 by repeats in the IGS. The loss of such repeats might explain how rDNA length heterogeneity, observed in the Meloidogyne genome, could have arisen. Each 129 by repeat also contains three copies of an 8 by subrepeat, which has sequence similarity to an element found in the IGS repeats of some plant rDNAs.
Similar content being viewed by others
References
Arnheim N, Kuehn M (1979) The genetic behaviour of a cloned mouse ribosomal DNA segment mimics mouse ribosomal gene evolution. J Mol Biol 134:743–765
Barker RF, Harberd NP, Jarvis MG, Flavell RB (1988) Structure and evolution of the intergenic region in a ribosomal DNA repeat unit of wheat. J Mol Biol 201:1–17
Boseley P, Moss T, Machler M, Portmann R, Birnstiel M (1979) Sequence organization of the spacer DNA in a ribosomal gene unit of Xenopus laevis. Cell 17:19–31
Dover GA (1982) Molecular Drive: a cohesive mode of species evolution. Nature 299:111–117
Gerbi SA (1985) Evolution of ribosomal DNA. In: MacIntyre R (ed) Molecular evolutionary genetics. Plenum Press, New York, pp 419–517
Hattori M, Sakaki Y (1986) Dideoxysequencing method using denatured plasmid templates. Anal Biochem 152:232–238
Hayward DC, Glover DM (1989) The promoters and spacers in the rDNAs of the melanogaster species subgroup of Drosophila. Gene 77:271–285
Henikoff S (1984) Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene 28:351–359
La Volpe A, Simeone A, D'Esposito M, Scotto L, Fidanza V, de Falco A, Bocinelli E (1985) Molecular analysis of the heterogeneity region of the human ribosomal spacer. J Mol Biol 183:213–223
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Morgan GT, McMahon H (1986) Alterations in cloned Xenopus ribosomal spacers generated by high-frequency plasmid recombination. Gene 49:389–394
Piller KJ, Baerson SR, Polaris NO, Kaufman LS (1990) Structural analysis of the short length ribosomal DNA variant from Pisum sativum L. cv. Alaska. Nucleic Acids Res 18:3135–3145
Smith HO, Birnstiel ML (1976) A simple method for DNA restriction site mapping. Nucleic Acids Res 3:2387
Vahidi H, Curran J, Nelson DW, Webster JM, McClure MA, Honda BM (1988) Unusual sequences, homologous to 5S RNA, in ribosomal DNA repeats of the nematode Meloidogyne arenaria. J Mol Evol 27:222–227
Vieira J, Messing J (1982) The pUC plasmds, an m13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19:259–268
Yakura K, Kato A, Tanifuji S (1984) Length heterogeneity of the large spacer of Vicia faba rDNA is due to the differing number of a 325 by repetitive sequence elements. Mol Gen Genet 193:400–405
Yavachev LP, Georgiev OJ, Braga EA, Avdonina TA, Bogomolova AE, Zhurkin VB, Nosikov VV, Hadjiolov AA (1986) Nucleotide sequence analysis of the spacer regions flanking the rat rRNA transcription unit and identification of repetitive elements. Nucleic Acids Res 14:2799–2810
Author information
Authors and Affiliations
Additional information
Communicated by D.Y. Thomas
Rights and permissions
About this article
Cite this article
Vahidi, H., Honda, B.M. Repeats and subrepeats in the intergenic spacer of rDNA from the nematode Meloidogyne arenaria . Molec. Gen. Genet. 227, 334–336 (1991). https://doi.org/10.1007/BF00259687
Issue Date:
DOI: https://doi.org/10.1007/BF00259687