Summary
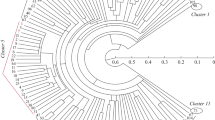
The introduction of molecular biology methodologies to plant improvement programs offers an invaluable opportunity for extensive germplasm characterization. However, the detection of adequate DNA polymorphism in self-pollinating species remains on obstacle. We have optimized a denaturing-gradient-gel electrophoresis (DGGE) system which, when used in combination with random amplified polymorphic DNA (RAPD) analysis, greatly facilitates the detection of reproducible DNA polymorphism among closely related plant lines. We have used this approach to estimate pedigree relationships among a spectrum of plant materials in wheat, barley and oat. Based on analysis with one or two primers, we were able to distinguish soft from hard winter wheat, and 2-rowed from 6-rowed barley. Further analysis with additional primers allowed resolution of polymorpisms even among closely related lines in highly selected populations. We placed 17 cultivars of oat into two distinct clusters that differed significantly from previous oat pedigree assessments. We believe that DGGE-RAPD is a superior method for detecting DNA polymorphism when compared to RFLP, agarose-RAPD, or polyacrylamide-RAPD methods.
Similar content being viewed by others
References
Bonierbale MW, Plaisted RL, Tanksley SD (1988) RFLP maps based on a common set of clones reveal modes of chromosomal evolution in potato and tomato. Genetics 120:1095–1103
Burmeister M, DiSibio G, Cox DR, Myers RM (1991) Identification of polymorphisms by genomic denaturing-gradient-gel electrophoresis: application to the proximal region of human chromosome 21. Nucleic Acids Res 19:1475–1481
Caetano-Anolles G, Bassam BJ, Gresshoff PM (1991) DNA amplification fingerprinting using very short arbitrary oligonucleotide primers. Biotechnology 9:553–557
Cox TS, Kiang YT, Gorman MB, Rodgers DM (1985a) Relationship between coefficient of parentage and genetic similarity indices in the soybean. Crop Sci 25:529–532
Cox TS, Lookhart GL, Walker DE, Harrell LG, Albers LD, Rodgers DM (1985b) Genetic relationships among hard red winter wheat cultivars as evaluated by pedigree analysis and gliadin polyacrylamide-gel electrophoretic patterns. Crop Sci 25:1058–1063
Delannay X, Rodgers DM, Palmer RG (1983) Relative genetic contributions among ancestral lines to North American soybean cultivars. Crop Sci 23:944–949
Efron B, Gong G (1983) A leisurely look at the bootstrap, the jackknife, and crossvalidation. Am Stat 37:36–48
Fisher SG, Lerman LS (1983) DNA fragments differing by single base-pair substitutions are separated in denaturing-gradient gels: correspondence with melting theory. Proc Natl Acad Sci USA 80:1579–1583
Gawel NJ, Jarrett RL (1991) A modified CTAB DNA-extraction procedure for Musa and Ipomoea. Plant Mol Biol Rep 91:262–266
Gray M, Charpentier A, Walsh K, Wu P, Bender W (1991) Mapping point mutations in the Drosophila rosy locus using denaturing-gradient-gel blots. Genetics 127:139–149
Halward T, Stalker T, LaRue E, Kochert G (1992) Use of singleprimer DNA amplification in genetic studies of peanut (Arachis hypogaea L). Plant Mol Biol 18:315–325
He S, Ohm H, Mackenzie S (1992) Detection of sequence polymorphism among wheat varieties. Theor Appl Genet (in press)
Hu J, Quiros C (1991) Identification of broccoli and cauliflower cultivars with RAPD markers. Plant Cell Rep 10:505–511
Landry BS, Kesseli RV, Farrara B, Michelmore RW (1987) A genetic map of lettuce (Lactuca sativa L) with restriction fragment length polymorphisms, isozyme, disease resistance and morphological markers. Genetics 116:331–337
Lessa EP (1992) Rapid surveying of DNA sequence variation in natural populations. Mol Biol Evol 9:323–330
Marsan PA, Livini C, Messmer MM, Mechinger E, Franceschini P, Monfredini G, Motto M (1992) Comparison between cluster analyses from RFLP and pedigree data of inbred lines related to stiff stalk synthetic heterotic group. Maize Genet Coop Newslett 66:18–20
Martin JM, Black TK, Hockett EA (1991a) Diversity among North American spring barley cultivars based on coefficients of parentage. Crop Sci 31:1131–1137
Martin GB, Williams JG, Tanksley SD (1991b) Rapid identification of markers near a Pseudomonas resistance gene in tomato using random primers and near-isogenic lines. Proc Natl Acad Sci USA 80:2336–2340
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis. A rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Murphy JP, Cox TS, Rodgers DM (1986) Cluster analysis of red winter wheat cultivars based upon coefficients of parentage. Crop Sci 26:672–676
Myers RM, Maniatis T, Lerman LS (1987) Detection and localization of single base changes by denaturing-gradient-gel electrophoresis. Methods Enzymol 155:501–527
Nei M, Li W-H (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Nei M, Shephens J, Saitou N (1985) Methods for computing the standard errors of branching points in an evolutionary tree and their application to molecular data from humans and apes. Mol Biol Evol 2:66–85
Price S, Shumaker KM, Kahler AL, Allard RW, Hill JE (1984) Estimates of population differentiation obtained from enzyme polymorphism and quantitative characters. J. Hered 72:141–142
Reiter RS, Williams J, Feldmann K, Rafalski JA, Tingey SV, Scolnik P (1992) Global and local genome mapping in Arabidopsis thaliana by using recombinant inbred lines and random amplified polymorphic DNAs. Proc Natl Acad Sci USA 89:1477–1481
Riedel GE, Swanberg SL, Kuranda KD, Marquette K, LaPan P, Bledsoe P, Kennedy A, Lin B-Y (1990) Denaturing-gradient-gel electrophoresis identifies genomic DNA polymorphism with high frequency in maize. Theor Appl Genet 80:1–10
Rodgers DM, Murphy JP, Frey KJ (1983) Impact of plant breeding on the grain yield and genetic diversity of spring oats. Crop Sci 23:737–740
Saiki RK, Gelfand DH, Stoffel S, Scharf S, Higuchi RH, Horn GT, Mullis KB, Elrich HA (1988) Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239:487–491
Souza E, Sorrells M (1988) Coefficients-of-parentage for North American oat cultivars released from 1951 to 1985. Search: agriculture no. 33. Cornell Univ Agric Exp Stn, Ithaca, New York
Souza E, Sorrells M (1989) Coefficients-of-parentage for North American oat cultivars released from 1951 to 1985. Crop Sci 29:595–601
St. Martin SK (1982) Effective population size for the soybean improvement program in maturity groups 00 to IV. Crop Sci 22:151–152
Tanksley SD, Bernatsky R, Lapitan NL, Prince JP (1988) Conservation of gene repertoire but not gene order in pepper and tomato. Proc Natl Acad Sci USA 85:6419–6423
Van Heusden AW, Bachmann K (1992) Genotype relationships in Microseris-Elegans (Asteraceae, Lactuceae) revealed by DNA amplification from arbitrary primers RAPDs. Plant Syst Evol 179:221–233
Welsh J, McClelland M (1990) Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res 189:7213–7218
Wilde J, Waugh R, Powell W (1992) Genetic fingerprinting of Theobroma clones using randomly amplified polymorphic DNA markers. Theor Appl Genet 83:871–887
Williams JG, Kublik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Wrigley CW (1970) Protein mapping by combined gel electrofocusing and electrophoresis: application to the study of genotypic variation in wheat gliadins. Biochem. Genet 4:509–516
Author information
Authors and Affiliations
Additional information
Communicated by P. L. Pfahler
Rights and permissions
About this article
Cite this article
Dweikat, I., Mackenzie, S., Levy, M. et al. Pedigree assessment using RAPD-DGGE in cereal crop species. Theoret. Appl. Genetics 85, 497–505 (1993). https://doi.org/10.1007/BF00220905
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00220905