Summary
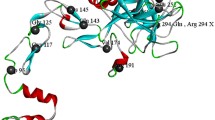
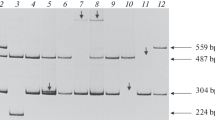
The molecular characterization of two haemophilia B defects, Calgary 1 and Calgary 2, was carried out using polymerase chain reaction (PCR) amplification and direct dideoxy sequencing. It had been previously shown that the Calgary 1 mutation affects the 5′ TaqI restriction site of exon VIII, whereas Calgary 2 involves the loss of the 3′ TaqI site of exon VIII of the factor IX gene. Sequencing data has now revealed that each of these alterations involves a C-to-T transition within a CpG dinucleotide. In each instance an arginine residue is replaced by a stop codon. These cases represent the recurrence of each particular alteration, both of which are predicted to result in the production of a truncated protein lacking a significant part of the catalytic region. A recently developed technique that reveals base substitutions as single-strand conformation polymorphisms (SSCP) was adapted for modelling in the detection of point mutations. Referred to here as single-strand conformation (SSC) analysis, this procedure, used in association with PCR, provided a reliable and sensitive system for molecular diagnosis in each of the cases presented. Computer-generated secondary structure predictions demonstrated a strong correlation with experimental results and the technique was used to screen 11 additional patients in the same region. A change detected by SSC analysis in one patient was localized to 55 base pairs, sequenced, and identified as a conservative amino acid substitution. This patient is now referred to as Calgary 3.
Similar content being viewed by others
References
Bird AP (1980) DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res 8:1499–1504
Brownlee GG (1987) The molecular pathology of haemophilia B, Biochem Soc Trans 15:1–8
Chen S-H, Scott CR, Lovrien EW (1989) Factor IX inPortland: a nonsense mutation (CGA to TGA) resulting in hemophilia B. Am J Hum Genet 44:567–569
Dimnik LS, Fraser BM, Poon M-C, Hoar DI (1990) Molecular diagnosis of single strand DNA conformational variation (abstract). Am J Hum Genet 47:A214
Driscoll MC, Bouhassira E, Aledort LM (1989) A codon 338 nonsense mutation in the factor IX gene in unrelated hemophilia B patients: factor IX338 New York. Blood 74:737–742
Fraser BM (1990) Molecular characterization of exon VIII mutation in the factor IX gene of two haemophilia B patients. Masters thesis, University of Calgary, Calgary, Alberta
Freedenberg DL, Chen S-H, Scott R (1989) A C to T mutation in the second TaqI site of exon VIII in the factor IX gene: detection by PCR amplification and direct sequencing (abstract). Am J Hum Genet 45 [Suppl]:A186
Gianelli F, Green PM, High KA, Lozier JN, Lillicrap DP, Ludwig M, Olek K, Reitsma PH, Goossens M, Yoshioka A, Sommers S, Brownlee GG (1990) Haemophilia B: database of point mutations and short additions and deletions. Nucleic Acids Res 18:4053–4059
Green PM, Bentley DR, Mibashan RS, Nilsson IM, Gianelli F (1990) The incidence and distribution of CpG and TpG transitions in the coagulation factor IX gene: a fresh look at CpG mutational hotspots. Nucleic Acids Res 18:3227–3231
Innis MA, Myambo KB, Gelfand DH, Brow MAD (1988) DNA sequencing with Thermus aquaticus DNA polymerase and direct dideoxy sequencing of polymerase chain reaction-amplified material DNA. Proc Natl Acad Sci USA 85:9436–9440
Ludwig M, Schwaab R, Eigel A, Horst J, Egli H, Brackman H-H, Olek K (1989) Identification of a single nucleotide C-to-T transition and five different deletions in patients with severe hemophilia B. Am J Hum Genet 45:115–122
Madisen L, Hoar DI, Holroyd CD, Crisp M, Hodes ME (1987) DNA banking: the effects of storage of blood and isolated DNA on the integrity of DNA. Am J Med Genet 27:379–390
Maniatis T, Fritsch EF, Sambrock J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Orita M, Iwahana H, Kanazawa H, Hayashi K, Sekiya T (1989a) Detection of polymorphisms of human DNA by gel electrophoresis as single-strand conformation polymorphisms. Proc Natl Acad Sci USA 86:2766–2770
Orita M, Suzuki Y, Sikiya T, Hayashi K, (1989b) A rapid and sensitive detection of point mutations and DNA polymorphisms using the polymerase chain reaction. Genomics 5:874–879
Osterberg G, Sommer R (1981) Computer support of DNA sequence analysis. Comput Prog Biomed 13:101–109
Poon M-C, Chui DHK, Patterson M, Starozik DM, Dimnik LS, Hoar DI (1987) Haemophilia B (Christmas disease) variants and carrier detection analyzed by DNA probes. J Clin Invest 79:1204–1209
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich HA (1988) Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239:487–491
Siguret V, Amselem S, Vidaud M, Assouline Z, Kerbiriou-Nabias D, Pietu G, Goossens M, Larrieu MJ, Bahnak B, Meyer D, Lavergne JM (1988) Identification of a CpG mutation in the coagulation factor IX gene by analysis of amplified DNA sequences. Br J Haematol 70:411–416
Taylor SAM (1990) Molecular analysis of mutations within the human factor IX gene. PhD thesis, Queen's University, Kingston, Ontario
Yoshitake S, Schach BG, Foster DC, Davie EW, Kurachi K (1985) Nucleotide sequence of the gene for human factor IX. Biochemistry 24:3736–3750
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics. Nucleic Acids Res 9:133–148
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fraser, B.M., Poon, MC. & Hoar, D.I. Identification of factor IX mutations in haemophilia B: application of polymerase chain reaction and single strand conformation analysis. Hum Genet 88, 426–430 (1992). https://doi.org/10.1007/BF00215677
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00215677