Summary
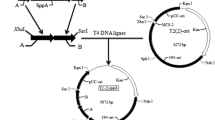
The gene coding for pectin methylesterase (PME) of Erwinia chrysanthemi B374 (pme) was cloned by a polymerase chain reaction. The pme gene was expressed in Bacillus subtilis using a secretion vector based on the promoter and signal sequence of the α-amylase gene from B. amyloliquefaciens. The cultivation of B. subtilis cells carrying the cloned pme resulted in efficient secretion of PME into the culture medium based on enzymatic and sodium dodecyl sulphate-polyacrylamide gel electrophoresis characterizations. The NH2-terminal sequence analysis of the secreted PME revealed two different NH2-termini. Heterologous processing was probably due to a second putative signal peptidase cleavage site at the joint region between the PME and α-amylase signal peptide.
Similar content being viewed by others
References
Ausubel L-Å, Brent R, Kingston RE, Moore DD, Smith JA, Seidman JG, Struhl K (1987) Current protocols in molecular biology. Harvard Medical School, Wiley Interscience, New York
Beaucage SL, Caruthers MH (1981) A new class of key intermediates for deoxypolynucleotide synthesis. Tetrahedron Lett 22:1859–1862
Gryczan TJ, Contente S, Dubnau D (1978) Characterization of Staphylococcus aureus plasmids introduced by transformation into Bacillus subtilis. J Bacteriol 134:318–329
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166:557–580
Hemilä H, Palva A, Paulin L, Arvidson S, Palva I (1990) Secretory S-complex of Bacillus subtilis: sequence analysis and identity to pyruvate dehydrogenase. J Bacteriol 172:5052–5063
Jansen EHJM, Laan CA, Reinerink EJM (1989) A simple method to improve the resolution in sodium dodecyl sulfate-polyacrylamide gels. In: Radola BJ (ed) Proceedings of the international meeting on electrophoresis, Technical University, Munich, October 23–25, 1989. Institut für Lebensmitteltechnologie und Analytische Chemie, Technische Universität München, FRG, pp 293–297
Kalkkinen N, Tilgmann C (1988) A gas-pulsed liquid-phase sequencer constructed from a Beckman 890D instrument by using Applied Biosystems delivery and cartridge blocks. J Protein Chem 7:242–243
Kotoujansky A (1987) Molecular genetics of pathogenesis by soft-rot Erwinias. Annu Rev Phytopathol 25:405–430
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head to bacteriophage T4. Nature 227:680–685
Melton DA, Krieg PA, Rebagliati MR, Maniatis T, Zinn K, Green MR (1984) Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res 12:7035–7056
Mullis KB, Faloona FA (1987) Specific synthesis of DNA in vitro via a polymerase catalysed chain reaction. Methods Enzymol 155:335–350
Palva I (1982) Molecular cloning of α-amylase gene from Bacillus amyloliquefaciens and its expression in B. subtilis. Gene 19:81–87
Plastow GS (1988) Molecular cloning and nucleotide sequence of the pectin methyl esterase gene of Erwinia chrysanthemi B374. Mol Microbiol 2:247–254
Raven PH, Ray FE, Eichhorn SE (1987) Introduction to the eucaryotic cell. In: Anderson S, Hefta G (eds) Biology of plants. Worth, New York, pp 15–44
Rexova-Benkova L, Markovic O (1976) Pectic enzymes. In: Tipson RS, Horton D (eds) Advances in carbohydrate chemistry and biochemistry. Academic Press, New York, pp 323–385
Rombouts FM, Pilnik W (1986) Pectinases and other cell-wall degrading enzymes of industrial importance. Symbiosis 2:79–90
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Saris P, Taira S, Airaksinen U, Palva A, Sarvas M, Palva I, Runeberg-Nyman K (1990) Production and secretion of pertussis toxin subunits in Bacillus subtilis. FEMS Microbiol Lett 68:143–148
Author information
Authors and Affiliations
Additional information
Offprint requests to: R. Heikinheimo
Rights and permissions
About this article
Cite this article
Heikinheimo, R., Hemilä, H., Pakkanen, R. et al. Production of pectin methylesterase from Erwinia chrysanthemi B374 in Bacillus subtilis . Appl Microbiol Biotechnol 35, 51–55 (1991). https://doi.org/10.1007/BF00180635
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00180635