Summary
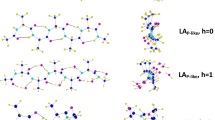
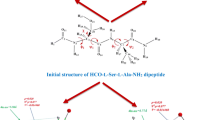
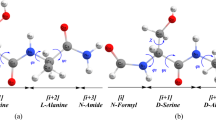
A quantum-chemical study of the chain-length dependent stability of the extended, 27-ribbon and 310-helix conformations in dehydroalanine (ΔAla) oligopeptides has been performed by using both semiempirical AM1 and ab initio 4–31G methodologies. The validity of both methods in the study of the conformational properties of ΔAla oligopeptides was tested first on the dipeptide. The results of this test showed that 4–31G and AM1 calculations are in good agreement with 6–31G* calculations and experimental data. In order to monitor the conformational conversions, ΔAla oligopeptides comprising two to six residues were constructed. Molecular geometries were fully optimized using AM1, and the final conformations were verified to be minima by analysis of the corresponding second-derivative matrices. Conformational studies revealed that the 310-helix is stabilized with respect to the 27-ribbon when the number of residues is three or four, at the AM1 and ab initio 4–31G level respectively, while the extended form is the most stable in all the calculations performed. On the other hand, if a linear behaviour is assumed for longer chains, our calculations show a trend that would predict a conversion from extended form to 310-helix in oligopeptides with around six (ab initio 4–31G) or eight (AM1) ΔAla residues. In order to explain these conformational changes, the cooperative effects for the different conformers were investigated. Large cooperative energy effects were found for the 310-helix conformation.
Similar content being viewed by others
References
Balaram, P., Proc. Ind. Acad. Sci. Chem. Sci., 93 (1984) 703.
Fasman, G.D., Trends Biochem. Sci., 14 (1989) 295.
Robson, B. and Garnier, J., Introduction to Proteins and Protein Engineering, Elsevier, Amsterdam, 1988.
Karles, I.L., Flippen-Anderson, J.L., Uma, K. and Balaram, P., Biochemistry, 28 (1989) 6696.
Paterson, Y., Rumsey, S.M., Benedetti, E., Nemethy, G. and Scheraga, H.A., J. Am. Chem. Soc., 103 (1981) 2947.
Ajò, D., Casarin, M. and Granozzi, G., J. Mol. Struct. (THEOCHEM), 86 (1982) 297.
Alagona, G., Ghio, C. and Pratesi, C., J. Comput. Chem., 8 (1991) 934.
Karle, I.L., Flippen-Anderson, J.L., Uma, K. and Balaram, P., Curr. Sci., 59 (1990) 875.
Alemán, C., Subirana, J.A. and Perez, J.J., Biopolymers, 32 (1992) 621.
Alemán, C. and Perez, J.J., Int. J. Quant. Chem., 47 (1993) 231.
Bella, J., Alemán, J., Alegre, C., Fernández-Santín, J.M. and Subirana, J.A., Macromolecules, 25 (1992) 5225.
Palmer, D.E., Pattaroni, C., Nunami, K., Chadha, R.K., Goodman, M., Wukamiya, T., Fukasi, K., Horimoto, S., Kitazawa, M., Fujita, H., Kubo, A. and Shiba, T., J. Am. Chem. Soc., 114 (1992) 5634.
Alemán, C. and Perez, J.J., Biopolymers, 33 (1993) 1811.
Gross, E. and Morell, J.L., J. Am. Chem. Soc., 89 (1967) 2791.
Gross, E., Morell, J.L. and Craig, L.C., Proc. Natl. Acad. Sci. USA, 62 (1969) 952.
Allgaier, H., Hung, G., Weiner, R.G., Schneider, U. and Zahner, H., Angew. Chem., Int. Ed. Engl., 24 (1985) 1051.
Kellner, R., Jung, G., Horner, T., Zahner, H., Schnell, N., Entian, K.D. and Gotz, F., Eur. J. Biochem., 177 (1988) 53.
Parkhurst, J.R. and Hodgins, D.S., Arch. Biochem. Biophys., 152 (1972) 597.
Wickner, R.B., J. Biol. Chem., 244 (1969) 6550.
Freund, S., Jung, G., Gutbrod, O., Folkers, G., Gibbans, W.A., Allgaier, H. and Werner, R., Biopolymers 31 (1991) 803.
Tori, K., Tokura, K., Yoshimura, Y., Okabe, K., Otsuka, H., Inagaki, F. and Miyazawa, T.M., J. Antibiot., 32 (1979) 1072.
Gupta, A. and Chauhan, V.S., Biopolymers, 30 (1990) 395.
Pascard, C., Ducruix, A., Lunel, J. and Parngé, T., J. Am. Chem. Soc., 99 (1977) 6418.
Ajò, D., Granozzi, G., Tondello, E., Del Prà, A. and Zanotti, G., J. Chem. Soc., Perkin Trans. II, (1979) 927.
Anderson, B., Hodgkin, D.C. and Viswamitra, M.A., Nature, 225 (1970) 233.
Padmanabhan, B., Dey, S., Khandelwal, B., Rao, G.S. and Singh, T.P., Biopolymers, 32 (1992) 1271.
Hariharan, P.C. and Pople, J.A., Theor. Chim. Acta, 28 (1973) 213.
Ditchfield, R., Hehre, W.J. and Pople, J.A., J. Chem. Phys., 54 (1971) 724.
Dewar, M.J.S., Zoebisch, E.G., Healy, E.F. and Stewart, J.J.P., J. Am. Chem. Soc., 107 (1985) 3902.
Alemán, C. and Orozco, M., J. Comput.-Aided Mol. Design, 6 (1992) 331.
Alemán, C. and Perez, J.J., J. Mol. Struct. (THEOCHEM), 285 (1993) 221.
Dannenberg, J.J. and Vinson, L.K., J. Phys. Chem., 92 (1988) 5635.
Vinson, L.K. and Dannenberg, J.J., J. Am. Chem. Soc., 111 (1989) 2777.
Alemán, C. and Perez, J.J., J. Comput.-Aided Mol. Design, 7 (1993) 241.
Fletcher, R. and Powell, M.J.D., Comput. J., 6 (1963) 163.
Davidson, W.C., Comput. J., 6 (1968) 406.
McIver, J.W. and Komornicki, A., J. Am. Chem. Soc., 94 (1972) 2625.
Dupuis, M., Watts, J.D., Villar, H.O. and Hurst, G.J.B., HONDO 7.0 Manual, 1987.
Olivella, S., QCPE Bull., 4 (1984) 10.
Stewart, J.J.P., QCPE Bull., 3 (1983) 431.
Schafer, L., Newton, S.Q., Cao, M., Peeters, A., Van Alsenoy, C., Wolinski, K. and Momany, F.A., J. Am. Chem. Soc., 115 (1993) 272.
Van Duijnen, P.T. and Thole, B.T., Biopolymers, 21 (1982) 1749.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Casanovas, J., Alemán, C. A quantum-mechanical study of the chain-length dependent stability of the extended and 310-helix conformations in dehydroalanine oligopeptides. J Computer-Aided Mol Des 8, 441–448 (1994). https://doi.org/10.1007/BF00125378
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00125378