Abstract
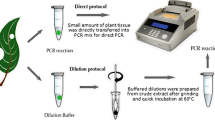
A polymerase chain reaction (PCR) approach using an air thermal cycler was developed to identify transgenic plants. The PCR amplification consisted of a pre-cycle denaturation (2 min at 94 °C), followed by 30 cycles of denaturation (10 sec at 94 °C), annealing (10 sec at 55 °C), and extension (1 min at 72 °C), and a post-cycle extension (7 min at 72 °C). The reaction could be completed in approximately 55 minutes. Denaturation and annealing times were found to be critical in obtaining repeatable amplification. This procedure was used to detect the neomycin phosphotransferase (NPT II) and β-glucuronidase (GUS) genes from genomic DNA of transformed petunia (Petunia hybrida cv. Mitchell) plants and shining willow (Salix lucida Muhl.) calli. Both AmpliTaq and Pfu DNA polymerases were capable of amplifying the target genes, but had significant differences in sensitivity.
Similar content being viewed by others
References
Asano Y, Ugaki M (1994). Transgenic plants of Agrostis alba obtained by electroporation-mediated direct gene transfer into protoplasts. Plant Cell Rep 13: 243–246.
Atkinson RG, Gardner RC (1991). Agrobacterium-mediated transformation of pepino and regeneration of transgenic plants. Plant Cell Rep 10: 208–212.
Berthomieu P, Meyer C (1991). Direct amplification of plant genomic DNA from leaf and root pieces using PCR. Plant Mol Bio 17: 555–557.
Blake NK, Ditterline RL, Stout RG (1991). Polymerase chain reaction used for monitoring multiple gene integration in Agrobacterium-mediated transformation. Crop Sci 31: 1686–1688.
Clarke HRG, Davis JM, Wilbert SM, et al. (1994). Wound-induced and developmental activation of a poplar tree chitinase gene promoter in transgenic tobacco. Plant Mol Bio 25: 799–815.
Does MP, Dekker BMM, Groot JAde, et al. (1991). A quick method to estimate the T-DNA copy number in transgenic plants at an early stage after transformation, using inverse PCR. Plant Mol Bio 17: 151–153.
Flaman J, Frebourg T, Moreau V, et al. (1994). A rapid PCR fidelity assay. Nucleic Acids Res 22: 3259–3260.
Grayburn WS, Vick BA (1995). Transformation of sunflower (Helianthus annuus L.) following wounding with glass beads. Plant Cell Rep 14: 285–289.
Gustafson CE, Alm RA, Trust TJ (1993). Effect of heat denaturation of target DNA on the PCR amplification. Gene 123: 241–244.
Ha SB, Wu FS, Thorne TK (1992). Transgenic turftype tall fescue (Festuca arundinacea Schreb.) plants regenerated from protoplasts. Plant Cell Rep 11: 601–604.
Hartman CL, Lee L, Day PR, et al. (1994). Herbicide resistant turfgrass (Agrostis palustris Huds.) by biolistic transformation. Bio/Technology 12: 919–923.
Hamill JD, Rounsley S, Spencer A, et al. (1991). The use of the polymerase chain reaction in plant transformation studies. Plant Cell Rep 10: 221–224.
Horsch RB, Fry JE, Hoffmann NL, et al. (1985). A simple and general method for transferring genes into plants. Science 227: 1229–1231.
Innis MA, Gelfand DH (1990). Optimization of PCRs. In: Innis MA, Gelfand DH, Sninsky JJ, et al. (eds), PCR protocols: a guide to methods and applications. San Diego: Academic Press, pp. 3–12.
Jong Jde, Mertens MMJ, Rademaker W (1994). Stable expression of the GUS reporter gene in chrysanthemum depends on binary plasmid T-DNA. Plant Cell Rep 14: 59–64.
Kaneyoshi J, Kobayashi S, Nakamura Y, et al. (1994). A simple and efficient gene transfer system of trifoliate orange (Poncirus trifoliata Raf.). Plant Cell Rep 13: 541–545.
Kenward KD, Altschuler M, Hildebrand D, et al. (1993). Accumulation of type I fish antifreeze protein in transgenic tobacco is cold-specific. Plant Mol Bio 23: 377–385.
Kuehnle AR, Sugii N (1992). Transformation of Dendrobium orchid using particle bombardment of protocorms. Plant Cell Rep 11: 484–488.
Lassner MW, Peterson P, Yoder JI (1989). Simultaneous amplification of multiple DNA fragments by polymerase chain reaction in the analysis of transgenic plants and their progeny. Plant Mol Bio Rep 7: 116–128.
Lundberg KS, Shoemaker DD, Adams MWW, et al. (1991). High fidelity amplification using a thermostable DNA polymerase isolated from Pyrococcus furiossus. Gene 108: 1–6.
Mayerhofer R, Koncz-Kalman Z, Nawrath C, at al. (1991). T-DNA integration: a mode of illegitimate recombination in plants. EMBO J 10: 697–704.
McGarvey P, Kaper JM (1991). A simple and rapid method for screening transgenic plants using the PCR. Biotechniques 11: 428–432.
Moore GA, Jacono CC, Neidigh JL, et al. (1992). Agrobacterium-mediated transformation of Citrus stem segments and regeneration of transgenic plants. Plant Cell Rep 11: 238–242.
Rocha SP, Maynard CA (1990). In vitro multiplication and Agrobacterium-transformation of willow (Salix lucida). In Vitro Cell Dev Bio 26: 43A (Abstract 111).
Rogers SO, Bendich AJ (1985). Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Bio 5: 69–76.
Sagi L, Panis B, Remy S, et al. (1995). Genetic transformation of banana and plantain (Musa spp.) via particle bombardment. Bio/Technology 13: 481–485.
Saiki RK, Scharf S, Faloona F, et al. (1985). Enzymatic amplification of β-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230: 1350–1354.
Swerdlow H, Dew-Jager K, Gesteland RF (1993). Rapid cycle sequencing in an air thermal cycler. BioTechniques 15: 512–519.
Wittwer CT, Garling DJ (1991). Rapid cycle DNA amplification: time and temperature optimization. BioTechniques 10: 76–83.
Xing Z (1995). Genetic transformation and regeneration of willows (Salix spp.). UMI Number 9608282. Ann Arbor, MI: Bell & Howell Co.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xing, Z., Satchwell, M. & Maynard, C.A. A rapid polymerase chain reaction method for early screening of transgenic plants. Methods Cell Sci 18, 7–13 (1996). https://doi.org/10.1007/BF00123517
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00123517