Abstract
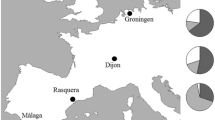
Mitochondrial DNA, sex linked allozymes, and chromosome A gene arrangement data, from eight European natural populations of Drosophila subobscura, were analyzed to determine the existence of latitudinal clines. Strong north-south correlations with latitude were found for gene arrangements and for the Hbdh and 6Pgdh allozymes. Gametic associations between the A2 gene arrangement, the Hbdh 96 and the 6Pgdh 96 alleles, point out some kind of epistatic interaction. At mtDNA level, the Hae III, A variant did not show a previously found north-south clinal variation.
Similar content being viewed by others
References
AfonsoJ. M., A.Volz, M.Hernàndez, H.Ruttkay, A. M.Gonzàlez, J. M.Larruga, V. M.Cabrera & D.Sperlich, 1990. Mitochondrial DNA variation and genetic structure in Old-World populations of Drosophila subobscura. Mol. Biol. Evol. 7: 123–142.
AviseJ. C., 1986. Mitochondrial DNA and the evolutionary genetics of higher animals. Phil. Trans. R. Soc. Lond.. B 312, 325–342.
BeckenbachA. & A.Prevosti, 1986. Colonization of North America by the European species Drosophila subobscura and D. ambigua. Am. Midland Nat. 115: 10–18.
BrncicD. & M.Budnik, 1980. Colonization of Drosophila subobscura Coll. in Chile. Drosophila Inf. Serv. 55: 20.
CabreraV. M., A. M.Gonzàlez, J. M.Larruga & C.Vega, 1983. Linkage disequilibrium in chromosome A of Drosophila subobscura. Genetica 61: 3–8.
ClarkA. G. & E. M. S.Lyckegaard, 1988. Natural selection with nuclear and cytoplasmic transmission. III. Joint analysis of segregation and mtDNA in Drosophila melanogaster. Genetics 118: 471–481.
FosM., M. A.Dominguez, A.Latorre & A.Moya, 1990. Mitochondrial DNA evolution in experimental populations of Drosophila subobscura. Proc. Natl. Acad. Sci. USA 87: 4198–4201.
HaleL. R. & R.Singh, 1987. Mitochondrial DNA variation and genetic structure in populations of Drosophila melanogaster. Mol. Biol. Evol. 4: 622–637.
KrimbasC. B., 1967. The genetics of Drosophila subobscura populations. III. Inversion polymorphism and climatic factors. Mol. Gen. Genet. 99: 133–150.
KrimbasC. B. & M.Loukas, 1980. The inversion polymorphism of Drosophila subobscura, pp. 163–234 in Evolutionary Biology, Vol 12, edited by M.Hecht, W. C.Steere and B.Wallace. Plenum, New York.
Kunze-MühlE. & E.Müller, 1958. Weitere Untersuchungen über die Chromosomale Struktur und natürlichen Strukturtypen von Drosophila subobscura. Chromosoma 9: 559–570.
MacRaeA. F. & W. W.Anderson, 1988. Evidence for non-neutrality of mitochondrial DNA haplotypes in Drosophila pseudoobscura. Genetics 120: 485–494.
MantelN., 1967. The detection of disease clustering and a generalized regression approach. Cancer Research 27: 209–220.
ManiatisT., E. F.Fritsch & J.Sambrook, 1982. Molecular Cloning. A Laboratory Manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
MisraR. K. & E. C. R.Reeve, 1964. Clines in body dimensions in populations of Drosophila subobscura. Genet. Res. 5: 240–256.
PfriemP., 1983. Latitudinal variation in wing size in Drosophila subobscura and its dependence on polygenes of chromosome O. Genetica 61: 221–232.
PinskerW. & D.Sperlich, 1979. Allozyme variation in natural populations of Drosophila subobscura along a north-south gradient. Genetica 50: 207–219.
PinskerW. & D.Sperlich, 1981. Geographical pattern of allozyme and inversion polymorphism on chromosome O of Drosophila subobscura and its evolutionary origin. Genetica 57: 51–64.
PinskerW. & I.Böhm, 1989. Allozyme divergence between different gene arrangements in Drosophila subobscura. J. Evol. Biol. 2: 353–366.
PrevostiA., 1955. Geographical variability in quantitative traits in populations of Drosophila subobscura. Cold Spring Harbor Symp. Quant. Biol. 20: 294–298.
PrevostiA., 1964. Chromosomal polymorphism in Drosophila subobscura from Barcelona (Spain). Genet. Res. 5: 27–38.
Prevosti, A., 1966. Inversion heterozygosity and size in a natural population of Drosophila subobscura. Symposium of the Mutational Process: Mutation in Populations, Prague, 9–11.
PrevostiA., 1967. Inversion heterozygosity and selection for wing length in Drosophila subobscura. Genet. Res. 10: 81–93.
PrevostiA., J.Ocaña & G.Alonso, 1975: Distances between populations of Drosophila subobscura, based on chromosome arrangement frequencies. Theoret. Appl. Genet. 45: 231–241.
PrevostiA., M. P.Garcia, L.Serra, M.Aguadé, G.Ribò & E.Sagarra, 1983. Association between allelic isozyme alleles and chromosomal arrangements in european populations and chilean colonizers of Drosophila subobscura, pp. 171–191 in Isozymes, vol. 10, Genetics and Evolution, edited by M. C.Ratazzi, J. G.Seandalios and G. S.Whitt, Liss, NY.
PrevostiA., R.Frutos, G.Alonso, A.Latorre, M.Monclùs & M. J.Martinez, 1984. Genetic differentiation between natural populations of Drosophila subobscura in the Western Mediterranean Area with respect to chromosomal variation. Génét. Sél. Evol. 16: 143–156.
PrevostiA., L.Serra, G.Ribò, M.Aguadé, E.Sagarra, M.Monclùs & M. P.Garcia, 1985. The colonization of Drosophila subobscura in Chile. II. Clines in the chromosomal arrangements. Evolution 39: 838–844.
PrevostiA., G.Ribò, L.Serra, M.Aguadé, J.Balaña, M.Monclùs & F.Mestres, 1988. Colonization of America by Drosophila subobscura: Experiment in natural populations that supports the adaptive role of chromosomal-inversion polymorphism. Proc. Natl. Acad. Sci. USA 85: 5597–5600.
RozasJ. & M.Aguadé, 1990. Evidence of extensive genetic exchange in the rp49 region among polymorphic chromosome inversions in Drosophila subobscura. Genetics 126: 417–426.
RozasJ., M.Hernàndez, V. M.Cabrera & A.Prevosti, 1990. Colonization of America by Drosophila subobscura: Effect of the founder event on the mitochondrial DNA polymorphism. Mol. Biol. Evol. 7: 103–109.
SauraA., S.Lakovaara, J.Lokki & P.Lankinen, 1973. Genic variation in central and marginal populations of Drosophila subobscura. Hereditas 75: 33–46.
SimmonsG. M., M. E.Kreitman, W. F.Quattlebaum & N.Miyashita, 1989. Molecular analysis of the alleles of alcohol dehydrogenase along a cline in Drosophila melanogaster. I. Maine, North Carolina, and Florida. Evolution 43: 393–409.
SperlichD., W.Pinsker & P.Pfriem, 1980. Inversion, allozyme and lethal frequencies in natural populations of Drosophila subobscura. Genetika 12: 91–101.
TrippaG., R.Scozzari & P.Pfriem, 1983. Clinal variation at the Peptidase-1 (Pept-1) locus in natural populations of Drosophila subobscura. Experientia 39: 787–788.
VoelkerR. A., C. C.Cockerham, F. M.Johnson, H. E.Schaffer, T.Mukai & L. E.Mettler, 1978. Inversions fail to account for allozyme clines. Genetics 88: 515–527.
WilsonA. C., R. L.Cann, S. M.Carr, M.George, U. B.Gyllensten, K. M.Helmbychowski, R. G.Higuchi, S. R.Palumbi, E. M.Prager, R. D.Sage & M.Stonenking, 1985. Mitochondrial DNA and two perspectives on evolutionary genetics. Biol. J. Linnean Soc. 26: 375–400.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Larruga, J.M., Rozas, J., Hernández, M. et al. Latitudinal differences in sex chromosome inversions, sex linked allozymes, and mitochondrial DNA variation in Drosophila subobscura . Genetica 92, 67–74 (1993). https://doi.org/10.1007/BF00057509
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00057509