Abstract
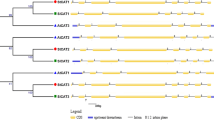
Two catalase genes,cat1 andcat2, have been isolated from the castor bean genome. They were located in the same direction on a chromosome at a distance of 2.4 kb,cat1 being on the downstream side ofcat2. The two genes contained introns at the same positions except that one of the 7 introns incat1 is missing incat2 and the corresponding introns differed in size and sequence between the two genes. The translated regions of the two genes had the same number of nucleotides and exhibited 81.3% nucleotide sequence identity. In addition to introns, the nucleotide sequences of the 5′-and 3′-flanking regions are highly divergent between the two genes. In etiolated seedlings,cat1 mRNA was present abundantly in endosperms and cotyledons and only in a small amount in roots. Thecat1 mRNA could not be detected in hypocotyls. By contrast,cat2 mRNA is most abundant in hypocotyls and roots, while endosperms and cotyledons contained only low levels ofcat2 mRNA. Although neithercat1 norcat2 mRNA could be detected in dry seeds, both mRNAs showed temporal accumulation in the endosperm in response to germination. These results suggest that expression of two tightly linked catalase genes of castor bean,cat1 andcat2, are differentially regulated during development.
Similar content being viewed by others
References
Beevers H: Microbodies in higher plants. Annu Rev Plant Physiol 30: 159–193 (1979).
Bethards LA, Skadsen SW, Scandalios JG: Isolation and characterization of a cDNA clone for theCat2 gene in maize and its homology with other catalases. (Correction.) Proc Natl Acad Sci USA 87: 6927 (1990).
Chevalier C, Yamaguchi J, McCourt P: Nucleotide sequence of a cDNA for catalase from Arabidopsis thaliana. Plant Physiol 99: 1726–1728 (1992).
Drumm H, Shopfer P. Effect of phytochrome on development of catalase activity and isozyme pattern in mustard (Simapis alba L.) seedlings. Planta 120: 13–20 (1974).
Eising R, Trelease RN, Ni W: Biogenesis of catalase in glyoxysomes and leaf-type peroxisomes of sunflower cotyledons. Arch Biochem Biophys 278: 258–264 (1989).
Frischauf A-M, Lehrach H, Poustka A, Murray N: Lambda replacement vectors carrying polylinker sequences. J Mol Biol 170: 827–842 (1983).
González E: The C-terminal domain of plant catalase: Implication for a glyoxysomal targeting sequence. Eur J Biochem 199: 211–215 (1991).
Hattori T, Fukumoto H, Nakagawa S, Nakamura K: Sucrose-induced expression of gene coding for the tuberous root storage protein, sporamin, of sweet potato in leaves and petioles. Plant Cell Physiol 32: 79–86 (1991).
Hattori T, Nakamura K. Genes coding for the tuberous root protein of sweet potato: identification of putative regulatory sequence in the 5′ upstream region. Plant Mol Biol 11: 417–426 (1988).
Havir EA, McHale NA: Biochemical and developmental characterization of multiple forms of catalase in tobacco leaves. Plant Physiol 84: 450–455 (1987).
Kunce CM, Trelease RN. Heterogeneity of catalase in maturing and germinated cottonseeds. Plant Physiol 81: 1134–1139 (1986).
Maniatis T, Fritsch EF, Sambrook J: Molecular Cloning: A Molecular Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY (1982).
Messing J. New M13 vecters for cloning. Meth Enzymol 101: 20–78 (1983).
Mori H, Higo K, Higo H, Minobe Y, Matsui H, Chiba S: Nucleotide and derived amino acid sequence of a catalase cDNA isolated from rice immature seeds. Plant Mol Biol 18: 973–976 (1992).
Ni W, Trelease RN: Two genes encode the two subunits of cottonseed catalase. Arch Biochem Biophys 289: 237–243 (1991).
Ni W, Trelease RN: Post-transcriptional regulation of catalase isozyme expression in cotton seeds. Plant Cell 3: 737–744 (1991).
Olsen LJ, Harada JJ: Biogenesis of peroxisomes in higher plants. In: Huang AHC, Taiz L (eds) Molecular Approaches to Compartmentation and Metabolic Regulation, pp. 129–137. American Society of Plant Physiologists (1991).
Ota Y, Ario T, Hayashi K, Nakagawa T, Hattori T, Maeshima M, Asahi T: Tissue-specific isoforms of catalase subunits in castor bean seedlings. Plant Cell Physiol 33: 225–232 (1992).
Redinbaugh MG, Sabre M, Scandalios JG: The distribution of catalase activity, isozyme protein, and transcript in the tissues of the developing maize seedling. Plant Physiol 92: 375–380 (1990).
Redingbaugh MG, Wadsworth GJ, Scandalios JG: Characterization of catalase transcripts and their differential expression in maize. Biochim Biophys Acta 951: 104–116 (1988).
Sakajo S, Nakamura K, Asahi T: Molecular cloning and nucleotide sequence of full-length cDNA for sweet potato catalase mRNA. Eur J Biochem 165: 437–442 (1987).
Singh R, Singh D: Peroxidase, polyphenol oxidase and catalase isozymes during germination and early plant development of tall and dwarf wheats (Triticum aestivum L.). Biol Plant 17: 235–240 (1975).
Vieira J, Messing J: Production of single stranded plasmid DNA. Meth Enzymol 153: 3–11 (1987).
Yanish-Perron C, Vieira J, Messing J: Improved M13 phage cloning vectors and host strans: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33: 103–119 (1985).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Suzuki, M., Ario, T., Hattori, T. et al. Isolation and characterization of two tightly linked catalase genes from castor bean that are differentially regulated. Plant Mol Biol 25, 507–516 (1994). https://doi.org/10.1007/BF00043878
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00043878