Summary
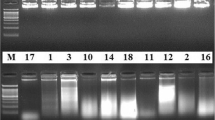
In this study, two PCR-based methodologies were evaluated for potential use in the determination of DNA diversity between 20 commercial sugarcane hybrids and 6 ‘outgroup’ varieties of S. spontaneum, S. officinarum and hybrids from early in the genealogy. The first method involved PCR amplification of sugarcane DNA in the presence of random, decamer primers (RAPDs), while the second protocol utilized specific microsatellite and telomere sequences as primers. A total of 41 RAPD primers (356 loci) were screened across the varieties of which 15 (160 loci) were used in the calculation of DNA diversity (expressed% similarity). This varied from 61 to 95%, with most of the commercial varieties showing more than 80% similarity in their DNA. The RAPD data indicated that there had been a gradual decline in DNA diversity (84% reduction) from the early inter-specific crosses to the commercial hybrids, probably as a result of backcrossing and in-breeding strategies used in the previous 5 to 6 generations of sugarcane breeding. The microsatellite and telomere data produced a much greater range in DNA similarity values (25–91%), probably due to the fact that these primers detect highly variable regions of the genome. It is suggested that these specific primers would not be suitable for determination of DNA diversity, but could be used more effectively in the development of a methodology for routine, rapid identification of sugarcane varieties.
Similar content being viewed by others
References
Aljanabi, S. M, R.J. Honeycutt, M. McClelland & B.W.S. Sobral, 1993. A genetic linkage map of Saccharum spontaneum (L.) SES 208. Genetics 134(4): 1249–1260.
Beckmann, J.S., & M. Soller, 1983. Restriction fragment length polymorphisms in genetic improvement: methodologies, mapping and cost. Theor. Appl. Genet. 67: 35–43.
Berding, N. & B.T. Roach, 1987. Germplasm maintenance and use. p. 143–210. In: D.J. Heinz (Ed). Developments in Crop Science II. Sugarcane improvement through breeding. (Elsevier).
Caetano-Anolles, G., B.J. Bassam & P.M. Gresshoff, 1991a. DNA amplification fingerprinting using very short arbitrary oligonucleotide primers. Biotechnology 9: 553–556.
Caetano-Anolles, G., B.J. Bassam & P.M. Gresshoff, 1991b. DNA amplification fingerprinting: a strategy for genome analysis. Plant Mol. Biol. Rep. 9: 294–307.
Dice, L.R., 1945. Measures of the amount of ecologic association between species. Ecology 26: 297–302.
Gale, M.D., S. Chao & P.J. Sharp, 1990. RFLP mapping in wheat-progress and problems. p. 353–364. In: J.P. Gustafson (Ed). Gene manipulation and plant improvement II. (Plenum Press, New York).
Hadrys, H., M. Balick and B. Schierwater, 1992. Amplification of random amplified polymorphic DNA (RAPDs) in molecular ecology. Mol. Ecol. 1: 55–63.
Haley, S.D., P.N. Miklas, L. Afanador & J.D. Kelly, 1994. Random amplified polymorphic DNA (RAPD) marker variability between and within pools of common bean. J. Amer. Soc. Hort. Sci. 119: 122–125.
Heinz, D.J., 1987. Sugarcane improvement: current productivity and future opportunities. In: Copersucar International Sugarcane Breeding Workshop Copersucar, Sao Paulo, pp 57–70
Honeycutt, H.J., B.W.S. Sobral, P. Kiem & J.E. Irvine, 1992. A rapid DNA extraction method for sugarcane and its relatives. Plant Mol. Biol. Rep. 10(1): 66–72.
Huff, D.R., R. Peakall & P.E. Smouse, 1993. RAPD variation within and among natural populations of outcrossing buffalograss (Buchloe dactyloides (Nutt.) Engelm.) Theor. Appl. Genet. 86: 927–934.
Jackson, P., 1994. Genetic relationships between attributes in sugarcane clones closely related to Saccharum spontaneum. Euphytica 79: 101–108.
Jeffreys, A.J., V. wilson & S.L. Thein, 1985. Hypervariable ‘minisatellite’ regions in human DNA. Nature 314: 67–73.
Ko, H.L., D.C. Cowan, R.J. Henry, G.C. Graham, A.B. Blakeney & L.G. Lewin, 1994. Random amplified polymorphic DNA analysis of Australian rice (Oryza sativa L.) varieties. Euphytica 80: 179–189.
Kolchinsky, A., & P.M. Gresshoff, 1994. Fingerprinting of plant varieties and species with telomeric primers. Plant Genome II Abstracts, pp 43. San Diego, California.
Lu, Y.H., A. D'Hont, F. Paulet, L. Grivet, M. Arnaud & J.C. Glaszmann, 1994a. Molecular diversity and genome structure in modern sugarcane varieties. Euphytica 78: 217–226.
Lu, Y.H., A. D'Hont, D.I.T. Walker, P.S. Rao, P. Feldmann & J.C. Glaszmann, 1994b. Relationship among ancestral species of sugarcane revealed with RFLP using single copy maize nuclear probes. Euphytica 78: 7–18.
Nei, M. & W.H. Li, 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. 76: 5269–5273.
Orozco-Castillo, C., K.J. Chalmers, R. Waugh and W. Powell, 1994. Detection of genetic diversity and selective gene introgression in coffee using RAPD markers. Theor. Appl. Genet. 87: 934–940
Price, S., 1957. Cytological studies in Saccharum and allied genera. III. Chromosome numbers in interspecific hybrids. Bot. Gaz. 144–159.
Price, S., 1965. Interspecific hybridization in sugarcane breeding. Proc. Int. Soc. Sugar Cane Technol. 14: 217–223.
Ranjekar, P.K., W. Ramakrishna, K.V. Choudhari, M.D. Lagu & V.S. Gupta, 1994. DNA fingerprinting in rice using simple repetitive DNA sequences and their characterization in rice. Plant Genome II Abstracts, pp 56. San Diego, California.
Roach, B.T., 1986. Evaluation and use of sugarcane germplasm. Proceedings of the 19th Congress of the International Society of Sugar Cane Technologists 1: 492–503.
Roach, B.T., 1989. Origin and improvement of the genetic base of sugarcane. Proceedings of Australian Society of Sugar Cane Technologists 11: 34–47.
Roach, B.T. & J. Daniels, 1987. A review of the origin and improvement of sugarcane. Copersucar International Sugarcane Breeding workshop 1: 1–33.
Rohlf, F.J., 1993. Numerical Taxonomy and Multivariate Analysis System (NTSYS-pc). Exeter Software, U.K.
Kus-Kortekaas, W., M.J.M. Smulders, P. Arens & B. Vosman, 1994. Direct comparison of genetic variation in tomato detected by a GACA-containing microsatellite probe and by random amplified polymorphic DNA Genome 37: 375–381.
Saiki, R.K., S. Scharf, F. Faloona, K.B. Mullis, G.T. Horn, H.A. Erlich & N. Arnheim, 1985. Enzymic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230: 1350–1354.
Sobral, B.W.S., D.P.V. Braga, E.S. LaHood & P. Kiem, 1994. Phylogenetic analysis of chloroplast restriction site mutations in the Saccharinae (Griseb.) subtribe of the Andropogoneae (Dumort.) tribe. Theor. Appl. Genet. (in press).
Sreenivasan, T.V., B.S. Ahloowalia & D.J. Heinz, 1987. Cytogenetics. p. 211–253. In: D.J. Heinz (Ed): Developments in Crop Science II. Sugarcane improvement through breeding. (Elsevier).
Tautz, D., 1989 Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nuc. Acids Res. 17: 4127–4138.
Tautz, D., M. Trick & G.A. Dover, 1986. Cryptic simplicity in DNA is a major source of genetic variation. Nature 322: 652–656.
Vosman, B., P. Arens, W. Rus-Kortekaas & M.J.M. Smulders, 1992. Identification of highly polymorphic DNA regions in tomato. Theor. Appl. Genet. 85: 239–244.
Weising, K., F. Weigand, A.J. Driesel, G. Kahl, H. Zischler & J.T. Epplen, 1989. Polymorphic simple GATA/GACA repeats in plant genomes. Nuc. Acids Res. 17: 10128.
Welsh, J. & M. McClelland, 1990. Fingerprinting genomes using PCR with arbitrary primers. Nuc. Acids Res. 18: 7213–7218.
Williams, C.E., & D.A. St. Clair, 1993. Phenetic relationships and levles of variability detected by restriction fragment length polymorphisms and random amplified polymorphic DNA analysis of cultivated and wild accessions of Lycopersicon esculentum. Genome 36: 619–629.
Williams, J.G.K., A.R. Kubelik, J.K. Livak, J.A. Rafalski & S.V. Tingey, 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nuc. Acids Res. 18: 6531–6535.
Yang, X., & C. Quiros, 1993. Identification of celery cultivars with RAPD markers. Theor. Appl. Genet. 86: 205–212.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Harvey, M., Botha, F.C. Use of PCR-based methodologies for the determination of DNA diversity between Saccharum varieties. Euphytica 89, 257–265 (1996). https://doi.org/10.1007/BF00034614
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00034614