Abstract
Despite unprecedented advances in understanding the intestinal microbiome, its potential to improve fields such as vaccinology has yet to be realized. This review briefly outlines the immunologic potential of the intestinal microbiome for vaccinology and highlights areas where the microbiome holds specific promise in vaccinology. Oral rotavirus vaccine effectiveness in low-income countries is used as a case study to describe how the intestinal microbiome may be employed to improve a vaccine’s immunogenicity. A top-down, evidence-based approach is proposed to identify effective microbiota-based applications for vaccine improvement. Applying evidence from field studies in pertinent populations that correlate microbiome composition with vaccine effectiveness to appropriate experimental platforms will lead to the identification of safe, vaccine-supporting microbiota targets that are relevant to populations in need of improvement in vaccine-induced immunity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The intestinal microbiome holds great promise for the field of vaccinology, but this potential has yet to be realized. |
Successful development of microbiota-targeted drugs to improve vaccine performance requires a top-down, reductive approach, translationally building on an evidence-base derived from relevant human populations. |
1 Introduction
Application of culture-independent sequencing techniques to the characterization of microbial communities on and in the human body has revolutionized our understanding of human health. The human microbiome, or the genomic catalogue of the bacteria, archaea, viruses (including bacteriophages), fungi, protozoa and helminths that inhabit our body space, is increasingly recognized to intersect with human health and disease [1, 2]. The human intestine is the most densely colonized surface in the human body, with a microbiome biomass of more than an order of magnitude greater than all other body sites [3]. Among the many functions ascribed to the intestinal microbiome are its roles in shaping mucosal and systemic immunity and protecting the human host from pathogen expansion and invasion [4, 5]. There is expanding interest in exploiting the intestinal microbiome for therapeutic goals, including improving vaccine-induced immunity. This review addresses the potential of the intestinal microbiome for vaccinology by providing a brief overview of evidence supporting a role for the intestinal microbiome in vaccine-induced immunity, and then uses rotavirus vaccines as a case study to outline how to move from a theoretical microbiome potential for vaccinology to an evidence-based application.
2 The Relevance of the Intestinal Microbiome to Vaccination
Principles of evolutionary biology support a long co-evolution between microbial consortia in the intestine and immune development in the human host. The intestinal tract in humans undergoes progressive, nonrandom, dynamic colonization after birth [6, 7], and normal immune development depends on this microbial colonization [8]. Germ-free animal studies are the most commonly cited example of this phenomenon as a lack of microbial exposures results in several immunologic abnormalities [9]. The presence of the intestinal microbiota in early life is particularly important for mucosal immunity and the normal maturation of gut-associated lymphoid tissues (GALT). Mice without microbial exposure have significantly fewer isolated lymphoid follicles (ILF) and smaller intestinal Peyer’s patches and mesenteric lymph nodes. These deficits result in diminished production of immunoglobulin (Ig) G1 and IgA by intestinal B cells. Furthermore, the absence of a microbiota in infancy results in deficits of intestinal T-cell subsets, as well as T-helper (Th) cell 2 skewing [8, 10, 11]. Robust adaptive immune responses to vaccination during infancy are possibly dependent on microbiota-mediated priming at the intestinal interface.
Intestinal homeostasis, reflecting a state where the human host tolerates the microbiota while still identifying potential pathogens, derives from a delicate interactive balance between intestinal microbiota, intestinal epithelium, and the host immune system [12]. Given this context, there are examples of how the intestinal microbiota can alter immune tone to potential pathogens and, by extension, may alter immune responses to vaccination. Depletion of the intestinal commensal microbiota can alter innate immune tone to, for example, enteric viruses [13,14,15]. Specific consortia of commensal microbiota can also directly and indirectly alter adaptive immunity to pathogens, inducing, for example, the differentiation of IgA-producing B cells and expansion of the functional diversity and repertoire of T effector Th17 and T-regulatory (Treg) cells [16]. Several reviews outline these interactions between the commensal microbiota and host immunity [4, 16,17,18,19].
The immunological basis for how the microbiome may hold specific promise for vaccination has been posited for many years [20, 21] and has recently been expanded upon [22,23,24,25]. The clearest demonstration of an effect of the intestinal microbiome on vaccination was in a study on inactivated influenza vaccine. Impaired antibody responses in germ-free and antibiotic-treated mice to the inactivated influenza vaccine could be restored by colonization with flagellated, but not aflagellated, Escherichia coli. TLR5 sensing of flagellin appeared to adjuvant the influenza vaccine response, promoting plasma cell differentiation in mice [26]. The aggregate of these findings support the immunological capacity for the microbiota, its cell wall components, or its metabolites to act as ‘endogenous vaccine adjuvants’ [22], potentially capable of being harnessed to amplify adaptive immune responses to specific pathogens.
However, missing in discussions on the potential of the microbiome for vaccinology is how to move translationally from this potential, often contained in animal studies, to understanding mechanisms and identifying safe and effective microbiome targets in relevant human populations. The focus of this review is to outline a potential multidisciplinary approach to addressing this gap, using rotavirus vaccines as a case example.
A logical departure point for microbiome–vaccine interactions is in the identification of vaccines with a high likelihood of microbiome interaction. Identifying areas with the highest microbiome–vaccinology potential can be aided by pragmatically converging animal and human approaches. The first approach mines mechanistic, bottom-up evidence, focusing on pathogens that, in animal studies, have already been shown to interact with the intestinal microbiome. These are numerous and include enteric viruses [13, 27,28,29], enteric bacterial pathogens [30,31,32,33,34], systemic viruses, such as influenza viruses and HIV-1 [26, 35], and even protozoa such as malaria [36]. The second approach is a human, epidemiologic, top-down approach, concentrating on those instances in which both vaccine performance and intestinal microbiome composition are divergent in different populations. An example of an area in which both these approaches converge are the poor efficacy and effectiveness of oral vaccines for enteric pathogens, such as rotavirus, in infants in low- and middle-income countries, which will be explored further as a case study in this review. Of note, numerous other areas in vaccinology exist with similar potential, including age-dependent differences in the duration of antibody responses, such as the diminished influenza vaccine efficacy observed in elderly populations [37] or the rapid decline in vaccine protection characterizing several early-life neonatal vaccines [38].
3 Rotavirus Vaccines: A Case Study
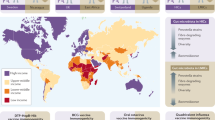
Oral vaccines have significantly diminished efficacy in low- and middle-income country settings [39, 40]. This phenomenon is true for both live-attenuated and heat-killed licensed vaccines against viral pathogens, such as rotavirus and poliovirus, and bacterial enteric pathogens, such as cholera and typhoid. The etiology of this diminished efficacy is certainly multifactorial and possibly vaccine-specific [41,42,43]. In parallel, bacterial intestinal microbiome composition and functional profile segregates by geography across all ages, and the intestinal microbiome composition differs significantly between high- and low-income country infants [44]. Other components of the intestinal microbiome share this divergence; the virome also differs significantly by geography, with a higher abundance of eukaryotic viruses in low-income country settings [45,46,47,48,49]. These parallel sets of differences in vaccine performance and microbiome composition along geographic lines tempt the assumption of microbiome-determined differences in oral vaccine performance between high- and low-income country infants. As the microbiome may be a modifiable etiology of oral vaccine under-performance, it is an attractive target in efforts to improve the public health impact of essential oral vaccines.
As noted above, interactions between viral and bacterial enteric pathogens, the intestinal microbiome, and host immunity have been extensively described in mouse models, and therefore provide in vivo support for the notion that the microbiome could influence enteric viral vaccine immunogenicity. The intestinal microbiome has been implicated in poliovirus, reovirus, mouse mammary tumor virus, rotavirus, and norovirus pathogenicity and immunity [13, 27,28,29, 50,51,52]. Importantly, the mechanisms underlying these interactions are multiple, ranging from direct effects, such as LPS stabilizing poliovirus and facilitating host cell entry [29], to indirect effects, such as antibiotic treatment permitting sterilizing innate immunity to persistent norovirus infection in mice. These viral-bacterial-immune pathways or ‘trans-kingdom interactions’ have been previously reviewed [53].
3.1 From Empiricism to Correlation
These mechanistic studies suggest that in addition to exploiting a more general ‘endogenous adjuvanting potential’ [22] of the microbiome, finding effective microbiome-based solutions for the improvement of oral vaccines may depend on pathogen-specific strategies. This approach is in opposition to empiric probiotic strategies, which have been the most commonly tested microbiota-targeted interventions to improve oral vaccine immunogenicity in humans [54]. Empiric probiotics such as Bifidobacterium and Lactobacillus are theorized to reduce intestinal mucosal inflammation, decrease pathogen mucosal binding, and increase adaptive antibody responses to pathogens [55, 56]. Three studies have tested the effect of these probiotics on oral vaccine performance in infants, with mixed results. Lactobacillus rhamnosus strain GG demonstrated modest and nonsignificant effects on rotavirus vaccine immunogenicity in Finnish [57] and Indian infants [58], respectively, and Bifidobacterium breve significantly lowered vaccine-specific antibodies to the oral cholera vaccine [54, 59]. An alternative empiric approach has been to use antibiotics to alter infants’ microbiome and improve vaccine immunogenicity. Theorizing that an increased enteropathogen burden in low-income countries may diminish oral vaccine immunogenicity, Indian infants received 3 days of azithromycin prior to oral polio vaccination. Despite reducing bacterial enteropathogens, there were no differences in polio vaccine immunogenicity between the two groups [60]. While these results are disappointing, they also underscore the limitations of an empiric approach, where known probiotics and antibiotics with general effects are applied to specific vaccines without effect. In order to fully assess whether the microbiome alterations can improve vaccine immunogenicity, a more targeted evidence-based approach is likely required where novel microbiota-based targets are identified for specific vaccines. The first step in this approach is to identify microbiome targets correlating with vaccine immunogenicity from the microbiota of low-income country infants themselves.
Rotavirus vaccines can be used as a case study to demonstrate such an approach. Currently licensed rotavirus vaccines are oral, live-attenuated vaccines, delivered in two or three doses 1 month apart, beginning at 6–10 weeks of age [61]. The vaccines have excellent effectiveness against severe rotavirus gastroenteritis in high-income countries (84%), however in low-income and some middle-income countries with high child mortality, their effectiveness is only between 45 and 57% [62]. Explanations for this difference in effectiveness often invoke microbiome differences related to malnutrition or a high pathogen burden [21, 39]; however, these associations have not been corroborated. Malnutrition has surprisingly little or no effect on most vaccine responses [63]. Studies correlating malnutrition indexes to rotavirus vaccine immunogenicity or efficacy have, with the exception of one study [64], predominantly not shown a correlation [65,66,67]. One study evaluating enteropathogen burden in relation to rotavirus immunogenicity in India showed that responders were actually more likely to harbor bacterial enteropathogens than nonresponders [68]. These data suggest that existing hypotheses triangulating microbiome composition with bacterial enteropathogen burden or malnutrition and rotavirus vaccine immunogenicity in low-income countries may not be valid.
However, emerging evidence does support a bilateral correlation between microbiome composition and rotavirus vaccine immunogenicity (Table 1). Three studies have compared the pre-vaccination fecal bacterial microbiome composition between infants with and without rotavirus vaccine immune response (anti-RV IgA > 20 IU/mL). A study in India demonstrated no significant differences in microbiome composition between rotavirus vaccine responders and nonresponders, however responders had a higher abundance of bacterial enteropathogens, as described above [68]. Two studies, one in rural Ghana and one in Pakistan, did demonstrate geography-specific differences in pre-vaccination fecal microbiome composition between matched rotavirus vaccine responders and nonresponders [69, 70]. In Ghana, rotavirus vaccine immunogenicity correlated with a low abundance of bacteria from the Bacteroidetes phylum and a high abundance of bacteria related to Streptococcus bovis. In Pakistan, rotavirus vaccine immunogenicity correlated with an increased abundance of Proteobacteria, (such as bacteria related to Escherichia coli and Serratia). In aggregate, these results suggest that if the microbiome correlates with rotavirus vaccine immunogenicity, this correlation may depend on location. Yet the overarching data also suggest that immune-stimulatory bacteria, such as bacterial enteropathogens in India, Gamma Proteobacteria in Pakistan, and bacteria related to Streptococcus bovis (known to have pathogenic potential) correlate with vaccine immunogenicity across populations. These results are hypothesis-generating, signifying a possible ‘adjuvanting’ capacity of the cell membrane components of the endogenous microbiota in low- and middle-income country infants to effect vaccine immune response.
3.2 From Correlation to Causation
The most simple extrapolation from these field studies is to hypothesize that differentially-abundant taxa between rotavirus vaccine seroconverters and non-seroconverters could be employed to improve rotavirus vaccine performance in developing countries; however, there are two major challenges to such an extrapolation. First, all microbiome correlations to date have been made with immunogenicity and not efficacy outcomes. Anti-RV IgA seroconversion is an imperfect correlate of protection from severe RV gastroenteritis, particularly in low-income settings [71]. There is therefore a risk that altering the microbiome to effect an immune response to vaccination may have little effect on the clinical outcome of interest, i.e. severe gastroenteritis. Conversely, anti-RV IgA is the best correlate of clinical protection currently available [71].
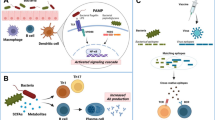
The second challenge is one that is shared by much of the microbiome research, i.e. how to distinguish correlation from causation and identify evidence-based mechanisms to rationally improve vaccine performance [72]. Identification of potential mechanisms mediating oral vaccine–microbiota interactions requires a top-down, stepwise, reductive approach. A useful example of how such a top-down strategy has been successful is in the Clostridium difficile field. Buffie et al. used longitudinal patient microbiota data to identify bacterial taxa correlating with resistance to C. difficile infection. Adoptive transfer of one of the protective taxa to mice correlated with C. difficile infection resistance. Subsequently, predictive gene functional profiling and metabolite analysis demonstrated that this taxa conferred infection resistance by synthesizing C. difficile-inhibiting metabolites from host-derived bile salts [30]. Similar approaches can be applied to the rotavirus and vaccinology fields (Fig. 1). The first step in this process is to test the capacity of microbial taxa derived from population-level and geography-specific correlations to actually alter vaccine immunogenicity in appropriate in vitro and in vivo experimental systems, as described below. Assuming that they do, the second step is to identify how. Population-level microbiome correlations could be a consequence of species membership, gene functional profiles, transcript or protein expression, metabolite profiles, or a combination of these effects [73, 74]. While risking a correlation with immunogenicity and not disease protection, employing a top-down approach ensures that microbiota-based therapeutic candidates remain relevant to the infant populations they are intended to serve. Simultaneously, a top-down approach capitalizes on human-derived high throughput genomic data, enabling unbiased identification of microbiome-vaccine interactions and thereby novel hypothesis generation.
Schematic overview of a top-down approach to identifying microbiota-derived targets for oral vaccines. a Well-conducted studies in appropriate low- and middle-income country settings can identify infants with and without adequate responses to vaccination. b Identification of differences in microbiota composition and community function between responder and nonresponder infants. c Differentially-abundant taxa, gene products, proteins, or metabolites can be tested for alteration of vaccine response and mechanism in appropriate experimental models: murine, gnotobiotic pig, human intestinal enteroid, and adult volunteers. d In parallel, microbiota differences distinguishing adult volunteer vaccine responders and nonresponders can be defined and tested in appropriate experimental models
3.3 Experimental Platforms
Numerous in vivo and in vitro experimental models can be employed to test potential microbiota–oral vaccine interactions, and their potential and limitations, using rotavirus vaccines as a case study, are explored below. A working model for rotavirus infection of adult mice was developed as early as the 1990s [75], and has broadly been employed to test rotavirus immunology. Mice with antibiotic-depleted microbiota and gnotobiotic mice have decreased rotavirus shedding and increased late anti-RV antibody responses following homologous rotavirus infection [76]. However, murine models have several disadvantages. Mice have significantly different intestinal physiology and microbiota from humans, and markedly decreased susceptibility to infection with human rotavirus strains. Neonatal mice, the best rotavirus infection model since the animals develop diarrhea [77], are challenging to manipulate in experimental settings, particularly in the context of microbiome manipulation or human fecal microbiome transfer. Murine immunity also differs considerably from humans, and divergence in toll-like receptor (TLR) recognition and innate immunity may have considerable relevance for microbiome–host immune interactions [78,79,80].
Gnotobiotic and neonatal pigs with humanized microbiota are alternative robust models by which to study human rotavirus vaccine strain interaction with microbiota [81,82,83]. Pigs are immunologically closer to humans, with long gestations and development of immunocompetence in the neonate [84]. They can be infected with human rotavirus strains, and also develop diarrhea upon infection [83]. Gnotobiotic pigs can be transplanted with human fecal microbiota [85, 86] and can thereby be used to interrogate how specific alterations in microbiota composition may affect immune responses to rotavirus vaccines [82, 87]. Gnotobiotic pigs with defined or humanized microbiota have been employed to study the immune effect of empiric probiotics on human rotavirus infection [87, 88]. Alternatively, gnotobiotic pigs transplanted with feces from infants with and without biomarkers of environmental enteropathy have been used to interrogate a possible effect of enteropathy on immunity to rotavirus infection [82]. The primary limitations of this model are cost and time, with relatively few facilities able to perform gnotobiotic studies. While mice are likely more evolutionarily divergent, pigs also diverge anatomically (spiral colon) and immunologically from humans, especially in their lymphoid tissue structure and trafficking [89]. Additionally, there is considerable difference between the composition of the initial human fecal inoculum and colonization in the gnotobiotic pig with alteration of composition over time, reflecting limitations in the efficacy and stability of microbiota transfer [82, 86].
An emerging and as yet underutilized platform for studying enteric pathogen-microbiome interactions is the human intestinal enteroid model. Derived from donated human intestinal cells, human intestinal enteroids are three-dimensional cultures that contain all existing differentiated epithelial cell types [90]. Enteroids can support replication of human rotavirus strains [91] and have also been successfully used to culture norovirus [92]. They are also slowly becoming robust enough to survive the addition of simple bacterial cultures or their supernatants [93]. While missing adaptive immunity, lymphatics, and complex microbiota, the system permits elegant and simplified examination of rotavirus–bacterial–innate immune interactions [94].
A final platform, more frequently used in vaccine development than in proof-of-principle or mechanistic studies, is the human clinical trial. Clinical studies in infants have already evaluated the effect of empiric probiotics (with known safety profiles [54, 58]) and short-course antibiotic regimens (such as azithromycin) on oral vaccine immunogenicity [60]. However, testing of novel probiotics with unknown safety profiles or longer courses of antibiotics with more dramatic, potentially durable alterations in microbiome composition in vulnerable infant populations is ethically untenable. Adult volunteer studies may make these trials possible and could support detection of a causal relationship between microbiome alteration and oral vaccine immunogenicity and the bandwidth of a potential effect.
As a proof-of-principle, targeted antibiotics can be used in adult volunteers to confirm that specific microbiota phenotypes from field studies correlate with vaccine immunogenicity. Rotavirus vaccine seroconversion, for example, correlates with an increased abundance of specific Proteobacteria in Pakistan and a decreased abundance of Bacteroidetes in Ghana. Vancomycin can diminish Bacteroidetes abundance but is also known to result in a simultaneous bloom of Proteobacteria [95, 96]. A proof-of-principle clinical trial is therefore currently underway evaluating the effect of vancomycin on rotavirus vaccine immunogenicity in an adult volunteer model (ClinicalTrials.gov identifier: NCT02154061). The use of such a model is also not without limitations. The adult microbiota is significantly different from the infant microbiota [44]. Additionally, adults undergo multiple asymptomatic rotavirus infections over their lifetime, therefore adult rotavirus vaccination likely results in an anamnestic boosting response, and not a priming response that is needed in infants in low-income settings. Nevertheless, successful alteration of rotavirus vaccine immunogenicity through targeted antibiotics would support a more causal role for the intestinal microbiome in determining vaccine immunogenicity, help corroborate microbiota components with the highest potential for vaccine improvement in infants, and build hypotheses for subsequent mechanistic evaluations.
Thus, upon identifying specific microbiota components or their metabolites in field studies or adult volunteer studies that differentiate vaccine responders from nonresponders, these components can be tested for mechanism, efficacy, and safety in in vitro and in vivo models, as any potential drug target would (Fig. 1). Rather than depending on known probiotics, deriving microbiota targets from an evidence base of relevant field studies and then working towards application is classically translational, but has not yet been fully applied to vaccinology. Yet such applications are not without risk. The endogenous intestinal microbiota can support host immune responses against enteric pathogens but they are also active participants in the induction of immune tolerance and maintenance of mutualism [24]. Alteration of the microbiota at an early age risks disruption of these roles with the potential induction of (auto)inflammation. The microbiota and their components straddle a complicated regulatory zone between classical drugs, requiring Investigational New Drug (IND) applications, and food or dietary supplements [97]. However, specific application of microbiome components to vaccinology, which assumes widespread dissemination of an effective product among infants, demands an especially evidence-based approach, with mechanistic underpinning, clear efficacy and safety outcomes, and strong regulatory rigor.
4 Conclusion
The rapid expansion of knowledge on the intestinal microbiome holds great promise for human health, and certainly for vaccinology. This review outlines the immunologic potential for the application of the intestinal microbiota to vaccines, focusing on improving the immunogenicity of specific enteric vaccines in the very young in low- and middle-income countries. Oral enteric vaccines, specifically rotavirus vaccines, provide a case study for how the immunogenic potential of the microbiome could be exploited to improve vaccine performance. Employing a top-down approach—studying differences in microbiome composition between vaccine responders and nonresponders in relevant populations and then moving to appropriate in vitro and in vivo experimental platforms—ensures that microbiota targets will remain relevant to both specific vaccines and patient populations. An adult volunteer model is perhaps an underutilized study model to bridge questions of correlation and causation in vaccines. Ultimately, a strong epidemiologic evidence-base with mechanistic underpinning and rigorous safety testing provides hope that effective microbiota-based drug targets can improve the protection elicited by vaccines and the protection of human health.
References
Ursell LK, Metcalf JL, Parfrey LW, Knight R. Defining the human microbiome. Nutr Rev. 2012;70(1):S38–44.
Lynch SV, Pedersen O. The Human intestinal microbiome in health and disease. New Engl J Med. 2016;375(24):2369–79.
Sender R, Fuchs S, Milo R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 2016;14(8):e1002533.
Kamada N, Núñez G. Regulation of the immune system by the resident intestinal bacteria. Gastroenterology. 2014;146(6):1477–88.
Buffie CG, Pamer EG. Microbiota-mediated colonizationresistance against intestinal pathogens. Nat Rev Immunol. 2013;13:791–801.
Milani C, Duranti S, Bottacini F, Casey E, Turroni F, Mahony J, et al. The first microbial colonizers of the human gut: composition, activities, and health implications of the infant gut microbiota. Microbiol Mol Biol Rev. 2017;81(4):e00036-17.
Nguyen QN, Himes JE, Martinez DR, Permar SR. The impact of the gut microbiota on humoral immunity to pathogens and vaccination in early infancy. PLoS Pathog. 2016;12(12):e1005997-6.
Gensollen T, Iyer SS, Kasper DL, Blumberg RS. How colonization by microbiota in early life shapes the immune system. Science. 2016;352(6285):539–44.
Bauer H, Horowitz RE, Levenson SM, Popper H. The response of the lymphatic tissue to the microbial flora. Studies on germfree mice. Am J Pathol. 1963;42:471–83.
Gaboriau-Routhiau V, Rakotobe S, Lecuyer E, Mulder I, Lan A, Bridonneau C, et al. The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity. 2009;31(4):677–89.
Mazmanian SK, Liu CH, Tzianabos AO, Kasper DL. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell. 2005;122(1):107–18.
Mowat AM. To respond or not to respond — a personal perspective of intestinal tolerance. Nature Rev Immunol. 2018;18(6):405–15.
Baldridge MT, Nice TJ, McCune BT, Yokoyama CC, Kambal A, Wheadon M, et al. Commensal microbes and interferon—determine persistence of enteric murine norovirus infection. Science. 2014;347(6219):266–9.
Zhang B, Chassaing B, Shi Z, Uchiyama R, Zhang Z, Denning TL, et al. Viral infection. Prevention and cure of rotavirus infection via TLR5/NLRC4-mediated production of IL-22 and IL-18. Science. 2014;346(6211):861–5.
Abt MC, Osborne LC, Monticelli LA, Doering TA, Alenghat T, Sonnenberg GF, et al. Commensal bacteria calibrate the activation threshold of innate antiviral immunity. Immunity. 2012;37(1):158–70.
Honda K, Littman DR. The microbiota in adaptive immune homeostasis and disease. Nature. 2016;535(7610):75–84.
Hooper LV, Gordon JI. Commensal host-bacterial relationships in the gut. Science. 2001;292(5519):1115–8.
Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336(6086):1268–73.
Kamada N, Chen GY, Inohara N, Núñez G. Control of pathogens and pathobionts by the gut microbiota. Nat Immunol. 2013;14(7):685–90.
Ferreira RBR, Antunes LCM, Finlay BB. Should the human microbiome be considered when developing vaccines? PLoS Pathog. 2010;6(11):e1001190.
Valdez Y, Brown EM, Finlay BB. Influence of the microbiota on vaccine effectiveness. Trends Immunol. 2014;35(11):526–37.
Collins N, Belkaid Y. Do the microbiota influence vaccines and protective immunity to pathogens? Engaging our endogenous adjuvants. Cold Spring Harb Perspect Biol. 2018;10(2):a028860.
Littman DR. Do the microbiota influence vaccines and protective immunity to pathogens? If so, is there potential for efficacious microbiota-based vaccines? Cold Spring Harb Perspect Biol. 2018;10(2):a029355.
Macpherson AJ. Do the microbiota influence vaccines and protective immunity to pathogens? Cold Spring Harb Perspect Biol. 2018;10(2):a029363-11.
Lynn DJ, Pulendran B. The potential of the microbiota to influence vaccine responses. J Leukoc Biol. 2018;103(2):225–31.
Oh JZ, Ravindran R, Chassaing B, Carvalho FA, Maddur MS, Bower M, et al. TLR5-mediated sensing of gut microbiota is necessary for antibody responses to seasonal influenza vaccination. Immunity. 2014;41(3):478–92.
Kuss SK, Best GT, Etheredge CA, Pruijssers AJ, Frierson JM, Hooper LV, et al. Intestinal microbiota promote enteric virus replication and systemic pathogenesis. Science. 2011;334(6053):249–52.
Uchiyama R, Chassaing B, Zhang B, Gewirtz AT. Antibiotic treatment suppresses rotavirus infection and enhances specific humoral immunity. J Infect Dis. 2014;210(2):171–82.
Robinson CM, Jesudhasan PR, Pfeiffer JK. Bacterial lipopolysaccharide binding enhances virion stability and promotes environmental fitness of an enteric virus. Cell Host Microb. 2014;15(1):36–46.
Buffie CG, Bucci V, Stein RR, McKenney PT, Ling L, Gobourne A, et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature. 2015;517(7533):205–8.
Rivera-Chávez F, Zhang LF, Faber F, Lopez CA, Byndloss MX, Olsan EE, et al. Depletion of butyrate-producing Clostridia from the gut microbiota drives an aerobic luminal expansion of Salmonella. Cell Host Microb. 2016;19(4):443–54.
Brandl K, Plitas G, Mihu CN, Ubeda C, Jia T, Fleisher M, et al. Vancomycin-resistant enterococci exploit antibiotic-induced innate immune deficits. Nature. 2008;455(7214):804–7.
Butterton JR, Ryan ET, Shahin RA, Calderwood SB. Development of a germfree mouse model of Vibrio cholerae infection. Infect Immun. 1996;64(10):4373–7.
Fanning S, Hall LJ, Cronin M, Zomer A, MacSharry J, Goulding D, et al. Bifidobacterial surface-exopolysaccharide facilitates commensal-host interaction through immune modulation and pathogen protection. Proc Natl Acad Sci USA. 2012;109(6):2108–13.
Williams WB, Liao HX, Moody MA, Kepler TB, Alam SM, Gao F, et al. Diversion of HIV-1 vaccine-induced immunity by gp41-microbiota cross-reactive antibodies. Science. 2015;349(6249):aab1253-3.
Villarino NF, LeCleir GR, Denny JE, Dearth SP, Harding CL, Sloan SS, et al. Composition of the gut microbiota modulates the severity of malaria. Proc Natl Acad Sci USA. 2016;113(8):2235–40.
Darvishian M. J Bijlsma, M, Hak E, van den Heuvel ER. Effectiveness of seasonal influenza vaccine in community- dwelling elderly people: a meta-analysis of test-negative design case-control studies. Lancet Infect Dis. 2014;14(12):1228–39.
Trotter CL, Andrews NJ, Kaczmarski EB, Miller E, Ramsay ME. Effectiveness of meningococcal serogroup C conjugate vaccine 4 years after introduction. Lancet. 2004;364(9431):365–7.
Levine MM. Immunogenicity and efficacy of oral vaccines in developing countries: lessons from a live cholera vaccine. BMC Biol. 2010;8:129.
John J. Role of injectable and oral polio vaccines in polio eradication. Expert Rev Vac. 2009;8(1):5–8.
Parker EP, Ramani S, Lopman BA, Church JA, Iturriza-Gómara M, Prendergast AJ, et al. Causes of impaired oral vaccine efficacy in developing countries. Future Microbiol. 2017;61(9):3994.
Velasquez DE, Parashar U, Jiang B. Decreased performance of live attenuated, oral rotavirus vaccines in low-income settings: causes and contributing factors. Expert Rev Vac. 2018;17(2):145–61.
Desselberger U. Differences of rotavirus vaccine effectiveness by country: likely causes and contributing factors. Pathogens. 2017;6(4):65–78.
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, et al. Human gut microbiome viewed across age and geography. Nature. 2012;486(7402):222–7.
Lim ES, Zhou Y, Zhao G, Bauer IK, Droit L, Ndao IM, et al. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat Med. 2015;21(10):1228–34.
Lim ES, Wang D, Holtz LR. The bacterial microbiome and virome milestones of Infant development. Trends Microbiol. 2016;24(10):801–10.
Reyes A, Haynes M, Hanson N, Angly FE, Heath AC, Rohwer F, et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature. 2011;466(7304):334–8.
Reyes A, Blanton LV, Cao S, Zhao G, Manary M, Trehan I, et al. Gut DNA viromes of Malawian twins discordant for severe acute malnutrition. Proc Natl Acad Sci USA. 2015;112(38):11941–6.
Holtz LR, Cao S, Zhao G, Bauer IK, Denno DM, Klein EJ, et al. Geographic variation in the eukaryotic virome of human diarrhea. Virology. 2014;468–470:556–64.
Kane M, Case LK, Kopaskie K, Kozlova A, MacDearmid C, Chervonsky AV, et al. Successful transmission of a retrovirus depends on the commensal microbiota. Science. 2011;334(6053):245–9.
Nice TJ, Baldridge MT, McCune BT, Norman JM, Lazear HM, Artyomov M, et al. Interferon-λ cures persistent murine norovirus infection in the absence of adaptive immunity. Science. 2015;347(6219):269–73.
Ma C, Wu X, Nawaz M, Li J, Yu P, Moore JE, et al. Molecular characterization of fecal microbiota in patients with viral diarrhea. Curr Microbiol. 2011;63(3):259–66.
Pfeiffer JK, Virgin HW. Transkingdom control of viral infection and immunity in the mammalian intestine. Science. 2016;351(6270):aad5872-2.
Maidens C, Childs C, Przemska A, Bin Dayel I, Yaqoob P. Modulation of vaccine response by concomitant probiotic administration. Br J Clin Pharmacol. 2012;75(3):663–70.
Panigrahi P, Parida S, Nanda NC, Satpathy R, Pradhan L, Chandel DS, et al. A randomized synbiotic trial to prevent sepsis among infants in rural India. Nature. 2017;548(7668):407–12.
Patel R, DuPont HL. New approaches for bacteriotherapy: prebiotics, new-generation probiotics, and synbiotics. Clin Infect Dis. 2015;60(Suppl 2):S108–21.
Isolauri E, Joensuu J, Suomalainen H, Luomala M, Vesikari T. Improved immunogenicity of oral D x RRV reassortant rotavirus vaccine by Lactobacillus casei GG. Vaccine. 2003;13(3):310–2.
Lazarus RP, John J, Shanmugasundaram E, Rajan AK, Thiagarajan S, Giri S, et al. The effect of probiotics and zinc supplementation on the immune response to oral rotavirus vaccine: a randomized, factorial design, placebo-controlled study among Indian infants. Vaccine. 2018;36(2):273–9.
Matsuda F, Chowdhury MI, Saha A, Asahara T, Nomoto K, Tarique AA, et al. Evaluation of a probiotics, Bifidobacterium breve BBG-01, for enhancement of immunogenicity of an oral inactivated cholera vaccine and safety: a randomized, double-blind, placebo-controlled trial in Bangladeshi children under 5 years of age. Vaccine. 2011;29(10):1855–8.
Grassly NC, Praharaj I, Babji S, Kaliappan SP, Giri S, Venugopal S, et al. The effect of azithromycin on the immunogenicity of oral poliovirus vaccine: a double-blind randomised placebo-controlled trial in seronegative Indian infants. Lancet Infect Dis. 2016;16(8):905–14.
Kirkwood CD, Ma LF, Carey ME, Steele AD. The rotavirus vaccine development pipeline. Vaccine. 2017. https://doi.org/10.1016/j.vaccine.2017.03.076
Jonesteller CL, Burnett E, Yen C, Tate JE, Parashar UD. Effectiveness of rotavirus vaccination: a systematic review of the first decade of global post-licensure data, 2006–2016. Clin Infect Dis. 2017;65(5):840–50.
Savy M, Edmond K, Fine PEM, Hall A, Hennig BJ, Moore SE, et al. Landscape analysis of interactions between nutrition and vaccine responses in children. J Nutr. 2009;139(11):2154S–218S.
Gastañaduy PA, Steenhoff AP, Mokomane M, Esona MD, Bowen MD, Jibril H, et al. Effectiveness of monovalent rotavirus vaccine after programmatic implementation in botswana: a multisite prospective case–control study. Clin Infect Dis. 2016;62(Suppl 2):S161–7.
Gruber JF, Hille DA, Liu GF, Kaplan SS, Nelson M, Goveia MG, et al. Heterogeneity of rotavirus vaccine efficacy among infants in developing countries. Pediatr Infect Dis J. 2017;36(1):72–8.
Emperador DM, Velasquez DE, Estivariz CF, Lopman Ben, Jiang B, Parashar U, et al. Interference of monovalent, bivalent, and trivalent oral poliovirus vaccines on monovalent rotavirus vaccine immunogenicity in rural Bangladesh. Clin Infect Dis. 2016;62(2):150–6.
Perez-Schael I, Salinas B, Tomat M, Linhares AC, Guerrero ML, Ruiz-Palacios GM, et al. Efficacy of the human rotavirus vaccine RIX4414 in malnourished children. J Infect Dis. 2007;196(4):537–40.
Parker EPK, Praharaj I, Zekavati A, Lazarus RP, Giri S, Operario DJ, et al. Influence of the intestinal microbiota on the immunogenicity of oral rotavirus vaccine given to infants in south India. Vaccine. 2018;36(2):264–72.
Harris V, Ali A, Fuentes S, Korpela K, Kazi M, Tate J, et al. Rotavirus vaccine response correlates with the infant gut microbiota composition in Pakistan. Gut Microb. 2017;3(1):1–9.
Harris VC, Armah G, Fuentes S, Korpela KE, Parashar U, Victor JC, et al. Significant correlation between the infant gut microbiome and rotavirus vaccine response in rural Ghana. J Infect Dis. 2017;215(1):34–41.
Patel M, Glass RI, Jiang B, Santosham M, Lopman B, Parashar U. A systematic review of anti-Rotavirus serum IgA antibody titer as a potential correlate of rotavirus vaccine efficacy. J Infect Dis. 2013;208(2):284–94.
Hanage WP. Microbiome science needs a healthy dose of scepticism. Nature. 2014;512(7514):247–8.
Huttenhower C, Knight R, Brown CT, Caporaso JG, Clemente JC, Gevers D, et al. Advancing the microbiome research community. Cell. 2014;159(2):227–30.
Britton RA, Cani PD, editors. Bugs as drugs. Therapeutic microbes for the prevention and treatment of disease. Washington DC: American Society of Microbiology Press; 2018.
Ward RL, McNeal MM, Sheridan JF. Development of an adult mouse model for studies on protection against rotavirus. J Virol. 1990;64(10):5070–5.
Uchiyama R, Chassaing B, Zhang B, Gewirtz AT. Antibiotic treatment suppresses rotavirus infection and enhances specific humoral immunity. J Infect Dis. 2014;210(2):171–82.
Gouvea VS, Alencar AA, Barth OM, de Castro L, Fialho AM, Araújo HP, et al. Diarrhoea in mice infected with a human rotavirus. J Gen Virol. 1986;67(Pt 3):577–81.
Nguyen TLA, Vieira-Silva S, Liston A, Raes J. How informative is the mouse for human gut microbiota research? Dis Model Mech. 2015;8(1):1–16.
Forstnerič V, Ivičak-Kocjan K, Ljubetič A, Jerala R, Benčina M. Distinctive recognition of flagellin by human and mouse toll-like receptor 5. PLoS One. 2016;11(7):e0158894-18.
Shay T, Jojic V, Zuk O, Rothamel K, Puyraimond-Zemmour D, Feng T, et al. Conservation and divergence in the transcriptional programs of the human and mouse immune systems. Proc Natl Acad Sci USA. 2013;110(8):2946–51.
Kandasamy S, Vlasova AN, Fischer D, Kumar A, Chattha KS, Rauf A, et al. Differential effects of Escherichia coli Nissle and Lactobacillus rhamnosus strain GG on human rotavirus binding, infection, and B cell immunity. J Immunol. 2016;196(4):1780–9.
Twitchell EL, Tin C, Wen K, Zhang H, Becker-Dreps S, Azcarate-Peril MA, et al. Modeling human enteric dysbiosis and rotavirus immunity in gnotobiotic pigs. Gut Pathog. 2016;8:51.
Yuan L, Saif LJ. Induction of mucosal immune responses and protection against enteric viruses: rotavirus infection of gnotobiotic pigs as a model. Vet Immunol Immunopathol. 2002;87(3–4):147–60.
Hein WR, Griebel PJ. A road less travelled: large animal models in immunological research. Nat Rev Immunol. 2003;3(1):79–84.
Pang X, Hua X, Yang Q, Ding D, Che C, Cui L, et al. Inter-species transplantation of gut microbiota from human to pigs. ISME J. 2007;1(2):156–62.
Zhang Q, Widmer G, Tzipori S. A pig model of the human gastrointestinal tract. Gut Microb. 2014;4(3):193–200.
Wen K, Tin C, Wang H, Yang X, Li G, Giri-Rachman E, et al. Probiotic Lactobacillus rhamnosus GG enhanced Th1 cellular immunity but did not affect antibody responses in a human gut microbiota transplanted neonatal gnotobiotic pig model. PLoS One. 2014;9(4):e94504-10.
Huang H-C, Vlasova AN, Kumar A, Kandasamy S, Fischer DD, Deblais L, et al. Effect of antibiotic, probiotic, and human rotavirus infection on colonisation dynamics of defined commensal microbiota in a gnotobiotic pig model. Benef Microb. 2018;9(1):71–86.
Haley PJ. The lymphoid system: a review of species differences. J Toxicol Pathol. 2017;30(2):111–23.
Sato T, Stange DE, Ferrante M, Vries RGJ, Van Es JH, Van den Brink S, et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology. 2011;141(5):1762–72.
Saxena K, Blutt SE, Ettayebi K, Zeng X-L, Broughman JR, Crawford SE, et al. Human intestinal enteroids: a new model to study human rotavirus infection, host restriction, and pathophysiology. J Virol. 2015;90(1):43–56.
Ettayebi K, Crawford SE, Murakami K, Broughman JR, Karandikar U, Tenge VR, et al. Replication of human noroviruses in stem cell-derived human enteroids. Science. 2016;353(6306):1387–93.
Blutt SE, Crawford SE, Ramani S, Zou WY, Estes MK. Engineered human gastrointestinal cultures to study the microbiome and infectious diseases. Cell Mol Gastroenterol Hepatol. 2018;5(3):241–51.
Saxena K, Simon LM, Zeng X-L, Blutt SE, Crawford SE, Sastri NP, et al. A paradox of transcriptional and functional innate interferon responses of human intestinal enteroids to enteric virus infection. Proc Natl Acad Sci USA. 2017;114(4):E570–9.
Isaac S, Scher JU, Djukovic A, Jiménez N, Littman DR, Abramson SB, et al. Short- and long-term effects of oral vancomycin on the human intestinal microbiota. J Antimicrob Chemother. 2017;72(1):128–36.
Vrieze A, Out C, Fuentes S, Jonker L, Reuling I, Kootte RS, et al. Impact of oral vancomycin on gut microbiota, bile acid metabolism, and insulin sensitivity. J Hepatol. 2014;60(4):824–31.
Sanders ME. Advancing probiotic research in humans in the United States: challenges and strategies. Gut Microb. 2016;7(2):1–4.
Parker EPK. Influence of the intestinal microbiota on the immune response to oral poliovirus and rotavirus vaccines in a low-income community in South India, doctoral thesis. London: Imperial College; 2016. Available at: https://spiral.imperial.ac.uk/handle/10044/1/42504. Accessed 11 Jun 2018.
Huda MN, Lewis Z, Kalanetra KM, Rashid M, Ahmad SM, Raqib R, et al. Stool microbiota and vaccine responses of infants. Pediatrics. 2014;134(2):e362–72.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
No funding was received for the preparation of this review.
Conflict of Interest
Dr Harris has no conflicts of interest to declare.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/), which permits any noncommercial use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Harris, V.C. The Significance of the Intestinal Microbiome for Vaccinology: From Correlations to Therapeutic Applications. Drugs 78, 1063–1072 (2018). https://doi.org/10.1007/s40265-018-0941-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40265-018-0941-3