Abstract
Introduction
The aim of this study was to investigate the distribution of antimicrobial susceptibility, biotypes and phylotypes of clinical Cutibacterium acnes (C. acnes, formerly Propionibacterium acnes) isolates as well as the relationship among demographic factors, C. acnes biotypes and phylotypes.
Methods
Cutibacterium acnes was collected from the skin lesions of acne patients who visited the dermatologic department of Huashan Hospital in Shanghai from October 2016 to March 2017. The agar dilution method was conducted to determine the minimum inhibitory concentrations (MICs) of C. acnes, the fermentation test to identify biotypes and then multiplex touchdown polymerase chain reaction (PCR) to identify phylotypes.
Results
Of the 63 C. acnes strains we isolated, 18 (28.6%), 31 (49.2%) and 4 (6.3%) strains were resistant to clindamycin, erythromycin and moxifloxacin, respectively; no strains were resistant to tetracycline, minocycline, fusidic acid or β-lactam, while metronidazole was completely resisted; 3 strains showed multidrug resistance (MDR). Biotype III (BIII) was the major biotype (50.8%) followed by BI and BV (both 15.9%), BII (12.7%) and lastly BIV (4.8%). IA1 was the predominant phylotype (71.4%) followed by IA2 (19.0%), II (4.8%), IB (3.2%) and IC (1.6%), while III was not detected. Significant differences were observed in the severity of disease: different degrees of acne severity reflected different biotype and phylotype distributions, and the biotype distribution of mild acne was different from that of moderate acne; the phylotype distribution of moderate acne varies from that of severe acne, too. Additionally, there was no significant difference in the distribution of biotypes or phylotypes between resistant and susceptible strains.
Conclusion
Erythromycin and clindamycin resistances are the most common in clinical C. acnes strains; BIII is the predominant biotype and IA1 is the major phylotype of C. acnes, which are mainly related to disease severity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
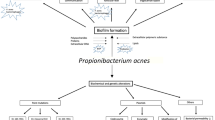
Acne is a common chronic inflammatory skin disease in both adolescents and adults that mainly involves epidermis and pilosebaceous units [1]. The pathogenesis of acne is complicated, and the colonization of Cutibacterium acnes (C. acnes, formerly Propionibacterium acnes, which was recently proposed for reclassification as Cutibacterium acnes [2]), is considered a crucial factor throughout the whole development of acne. C. acnes promotes the abnormal proliferation and differentiation of keratinocytes and increases sebum production. In addition, C. acnes can lead to the overexpression of Toll-like receptors (TLRs), protease-activated receptors (PARs) and matrix metalloproteinase (MMP) by keratinocytes, all of which can further activate the innate immune system [3, 4]. As the main therapy of acne, topical and oral antibiotics against C. acnes have been used for decades [5]. However, the increasing number of strains of drug-resistant C. acnes due to antibiotic abuse in acne treatment has aroused wide concerns [6]. Based on the fact that different strains determine different pathogenicity and resistance, classification of C. acnes is of great significance [7]. According to previous studies, we can make use of fermentation tests for biotyping [8] while using multiplex touchdown polymerase chain reaction (PCR) for genotyping [9]. So far, there have been few reports on the biotype and phylotype classification of clinical C. acnes isolates, and little is known about the biologic and phylogenetic typing of drug-resistant C. acnes. Therefore, in this study, we investigated the antimicrobial susceptibility and biotypes and phylotypes of clinical C. acnes isolates as well as the relationship between demographic factors and C. acnes biotypes and phylotypes.
Methods
Patients
Patients > 18 years of age with grade I–IV facial acne according to the Pillsbury scale were included in the present study. Patients who suffered from scar constitution; unable to cooperate or unwilling to sign informed consent; with severe medical complications and pregnant women were excluded from the study. This study was approved by the Ethics Committee of Huashan Hospital of Fudan University. Informed consent was obtained from all individual participants included in the study.
Isolation and Identification of C. acnes
Samples were isolated from the lesions of acne patients who visited the Dermatologic Department of Huashan Hospital in Shanghai, China, from October 2016 to March 2017. After being sterilized with iodine tincture, the acne lesions (comedones, papules and pustules) were compressed with a comedone extractor. Then the contents were transported immediately with a sample smear and inoculated on Brucella agar (Becton-Dickinson, USA) containing 5% (v/v) lysed defibrinated sheep’s blood (Zhuzhai Blood Reagent Supply Station, Shanghai), supplemented with vitamin K1 (Sigma-Aldrich, USA) and incubated in an anaerobic chamber (Ruskinn Invivo2 400 Hypoxia workstation, 10% CO2, 10% H2, 80% N2) at 37 °C for 48–72 h. The cultured microorganisms were identified as C. acnes with an API-20A system (bioMérieux, France). The bacteria were then placed in 40% glycerin broth and stored at − 70 °C. The standard C. acnes strain (ATCC 6919) was purchased from the Guangdong Institute of Microbiology, China.
Antimicrobial Agents
Metronidazole, clindamycin, erythromycin, fusidic acid, moxifloxacin, ceftriaxone, cefoxitin, minocycline, tetracycline, ampicillin, sulbactam and imipenem were purchased from the China National Institutes for Food and Drug Control. Ampicillin and sulbactam were mixed at a 2:1 ratio according to recommendations of the Clinical and Laboratory Standards Institute (CLSI).
Determination of Susceptibility to Antibiotics of C. acnes
Agar dilution assays were employed to measure the minimum inhibitory concentrations (MICs) of clinical C. acnes isolates. All the antimicrobials were prepared in concentrations ranging from 0.06 to 128 μg/ml with a two-fold serial dilution. A standard inoculum of 105 colony-forming units per 1 μl was also prepared and delivered by a multipoint inoculator (Denley A40, Denley Ltd., Billingshurst, Sussex, UK). The MIC, defined as the lowest concentration with no visible bacterial growth, was determined after incubation under anaerobic conditions at 35 °C for 48 h. A standard strain (ATCC 6919) was used as a quality control strain. The breakpoints for the antimicrobial agents were defined based on CLSl 2016 M100-S23 criteria except for fusidic acid, which was based on the European Committee on Antimicrobial Susceptibility Testing (EUCAST) 2016 criteria.
Statistical Analysis
Data were analyzed using SPSS version 24.0. The distribution of biotypes and phylotypes of Cutibacterium acnes, according to different clinical characteristics, and the distribution differences of biotypes and phylotypes between resistant and susceptible strains were analyzed with Fisher’s exact test. P < 0.05 was considered statistically significant. For the relationship between disease severity and C. acnes biotypes and phylotypes, the Bonferroni test for post hoc comparisons was performed. P < 0.0167 was considered statistically significant.
Grading of Acne Severity
Acne severity was classified into four grades according to the Pillsbury scale: grade I (≤ 30 eruptions, mainly comedones with a small number of papules and pustules), grade II (31–50 eruptions, comedones with an intermediate amount of papules and pustules), grade III (51–100 extensively distributed eruptions, many papules and pustules with occasional large inflammatory lesions, less than 3 nodules) and grade IV (> 100 eruptions, > 3 nodules or cysts). Grades I and IV were defined as mild and severe acne, respectively, while grades II and III were defined as moderate acne.
Fermentation Tests
Determination of the biotype of C. acnes was performed based on previously reported procedures [10]. Both basal medium (pH 7.0) and carbohydrate medium (basal medium plus 1 g of carbohydrate per dl) were inoculated with Pasteur pipettes. Of the carbohydrates used, deoxyribose, ribose and xylose were sterilized by filtration and added to autoclaved basal media. Incubation was performed at 37 °C for 7 days. The pH values of the cultures and uninoculated media were measured. Then, the “sugar-specific pH” (SpH) for each carbohydrate was calculated by the equation SpH = (A − B) − (C − D), where A, B, C and D are the pH of the carbohydrate medium, inoculated carbohydrate medium, basal medium and inoculated basal medium, respectively. SpH ≥ 0.35 was interpreted as fermentation positive and SpH < 0.35 as fermentation negative. Ribose (+), erythritol (+) and sorbitol (+) strains were defined as biotype I (BI); ribose (+), erythritol (+) and sorbitol (–) strains as biotype II (BII); ribose (+), erythritol (–) and sorbitol (+) strains as biotype III (BIII); ribose (+), erythritol (–) and sorbitol (–) strains as biotype IV (BIV); ribose (–), erythritol (–) and sorbitol (–) strains as biotype V (BV).
Multiplex Touchdown PCR
Multiplex touchdown PCR was employed according to the report of Barnard et al. [11], which can identify phylotypes of C. acnes isolates in a single reaction based on its pattern of reaction with six primer sets that target the 16S rRNA gene (all isolates), ATPase (types IA1, IA2 and IC), sodA (types IA2 and IB), atpD (type II), recA (type III) housekeeping genes and a Fic family toxin gene (type IC). Bacterial genomic DNA was prepared with TaKaRa Taq. PCR amplification was carried out using longGene L96G. The samples contained 10 × PCR buffer 2 µl, primers set mix 2 µl, dNTP mix 2 µl, rTaq 0.3 µl, DNA templates 2 µl and ddH2O 11.7 µl in a total volume of 20 µl. The samples were initially heated at 94 °C for 1 min followed by 14 cycles consisting of 94 °C for 30 s, 66 °C (decreasing incrementally by 0.3 °C per cycle) for 30 s and 72 °C for 1 min, followed by 11 cycles at 94 °C for 30 s, 62 °C for 30 s and 72 °C for 1 min, culminating with a final cycle at 72 °C for 10 min. A negative control (PCR water) and six positive control samples representing all phylotypes were included in all experiments. The PCR products were analyzed by electrophoresis on 1.5% (wt/vol) agarose gels.
Results
Culture and Identification of C. acnes
Samples were collected from 100 patients with acne, and 69 Cutibacterium strains were identified. Of these strains, C. acnes was found in 63 patients and Cutibacterium granulosum in 6 patients.
MICs of C. acnes
The MIC range of clindamycin and erythromycin for C. acnes was ≤ 0.06 to > 128 μg/ml. Among the isolated C. acnes strains, 58.7% (37/63) were susceptible to clindamycin and 28.6% (18/63) were resistant to clindamycin; 50.8% (32/63) were susceptible to erythromycin and 49.2% (31/63) were resistant to erythromycin. The MIC range of moxifloxacin was ≤ 0.06 to > 2 μg/ml, and 6.3% (4/63) were resistant to moxifloxacin. In addition, three C. acnes strains were simultaneously resistant to clindamycin, erythromycin and moxifloxacin, hence exhibiting multidrug resistance (MDR). The MIC range was 0.125–1 μg/ml for tetracycline and 0.06–1 μg/ml for minocycline. No strains were resistant to tetracycline or minocycline. The MIC range was 0.25–2 μg/ml for fusidic acid. Fusidic acid showed good antimicrobial susceptibility against most C. acnes strains (93.7% were susceptible and 6.3% showed moderate sensitivity), and resistance was not observed. In vitro, β-lactam antibiotics demonstrated favorable antibiotic activities, while metronidazole was completely resisted (Table 1).
Clinical Characteristics and Biotypes of C. acnes
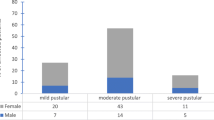
All patients were grouped according to family history, age, gender, disease duration, disease severity and previous treatment. Of these patients, 57.1% were male and 42.9% female. The mean age was 22.4 ± 4.5 years, and the mean course of disease was 13.1 ± 1.1 months. According to the Pillsbury grading system, grade I was found in 5 patients, grade II in 16, grade III in 23 and grade IV in 19. The numbers of patients treated with antibiotics or retinoids were: oral antibiotics (n = 31), topical antibiotics (n = 28), oral isotretinoin (n = 14) and topical retinoids (n = 29). BIII was the major biotype (50.8%; 32/63), BI and BV each accounted for 15.9% (10/63), BII accounted for 12.7% (8/63), and BIV accounted for only 4.8% (3/63). The distribution of BIII had little correlation with all but only one disease demographic factor: disease severity. The biotype distribution was significantly different among four degrees of acne severity (P = 0.048 < 0.05), while Bonferroni correction showed that there were no significant statistical differences in six pairwise comparisons (P > 0.0083). When we classified acne severity as three degrees including mild, moderate and severe, Fisher’s exact test showed that the biotype distribution of C. acnes was significantly different among the three disease degrees (P = 0.031 < 0.05), and Bonferroni correction suggested that statistical difference was only found between mild and moderate acne (P = 0.016 < 0.0167). The standard C. acnes strain (ATCC 6919) was biotyped as BIII (Table 2).
Clinical Characteristics and Phylotypes of C. acnes
IA1 was the predominant phylotype (71.4%, 45/63), followed by IA2 (19.0%, 12/63), II (4.8%, 3/63), IB (3.2%, 2/63) and IC (1.6%, 1/63), while III was not detected. In accordance with the biotype distribution, disease severity was the only disease demographic factor that had correlation with the distribution of IA1. The phylotype distribution was significantly different among the four degrees of acne severity (P = 0.033 < 0.05), while Bonferroni correction showed that there were no significant statistical differences in six pairwise comparisons (P > 0.0083). When acne severity was classified as three degrees including mild, moderate and severe, Fisher’s exact test showed that the phylotype distribution of C. acnes was significantly different among the three disease degrees (P = 0.015 < 0.05), and Bonferroni correction suggested that statistical difference was only found between moderate and severe acne (P = 0.011 < 0.0167). The standard C. acnes strain (ATCC 6919) was determined as IA1 (Table 3).
Distribution of Biotypes of Antibiotic-Resistant and -Susceptible C. acnes
The overwhelming majority of C. acnes biotypes were found to have antibiotic-resistant and -susceptible subtypes. The only exception was biotype I, which lacked a moxifloxacin-resistant subtype. In addition, among the three MDR strains, BIII, BII and BV were found, respectively. Consequently, BIII was the dominant biotype in both antibiotic-resistant and -susceptible C. acnes strains, and there was no significant difference in the distribution of biotypes between resistant and susceptible strains (P > 0.05) (Table 4).
Distribution of Phylotypes of Antibiotic-Resistant and -Susceptible C. acnes
All phylotypes were found to have antibiotic-susceptible subtypes except the undetected phylotype III. Clindamycin- and moxifloxacin-resistant subtypes were only observed in phylotype IA1 and IA2, while erythromycin-resistant subtypes also in the phylotype IB and II division. Moreover, the three MDR C. acnes isolates were identified to be two IA1 and one IA2 strains. There was no significant difference in the distribution of phylotypes between resistant and susceptible strains (P > 0.05) (Table 5).
Discussion
Cutibacterium acnes is an anaerobic gram-positive bacillus and exists as a skin commensal in the human skin microbiota [12]. As the predominant bacterium in sebaceous sites, C. acnes is confirmed as the most important acne-inducing pathogen. Antibiotic therapy against C. acnes has remained the major acne therapy for years. However, antibiotic abuse has led to increased levels of resistance, decreased C. acnes sensitivity to treatment and ultimately treatment failure [13].
The first resistant C. acnes strain was reported by Leyden in 1976 [14], and the resistance rate of C. acnes has increased from 20% in 1978 to 62% in 1996 [15]. Many countries have reported that > 50% of C. acnes strains are resistant to antibiotics, especially to macrolides [6]. In general, resistance to erythromycin and clindamycin are the most common, while tetracycline remains relatively susceptible. Studies show high resistance rates of erythromycin and clindamycin in Europe, such as > 70% in France and even 91% in Spain, while tetracycline resistance ranges from 0% to 26.4% [16, 17]. Likewise, there is great variability among resistance rates in different Asian countries. A Japanese study by Nakase et al. [18] reported 23.2% resistance to erythromycin, 18.8% to clindamycin, 4.3% to doxycycline and 0% to minocycline. However, in Hong Kong, a recent study found clindamycin resistance the most common (53.5%), followed by erythromycin (20.9%) and tetracyclines (16.3%) [19]. The different resistance rates between countries may be associated with different prescribing patterns and usage limitations. Current research also found that the presence of C. acnes resistance was related to age, duration and severity of disease [18, 19]. The relationship between resistance and history of antibiotic use remains controversial. Previous antibiotic treatment does not seem to be a premise for the presence of resistance because resistant C. acnes have also been found in patients without antibiotic therapy, which might be due to contact-mediated spread of resistant C. acnes [17, 20].
Our results suggest that erythromycin resistance (49.2%) is the most prevalent in clinical C. acnes isolates, followed by clindamycin (28.6%), which is consistent with the overall trend throughout the world. As the two most frequently prescribed topical antibiotics, erythromycin and clindamycin have been used in acne treatment for decades. However, it has been reported that the efficacy of topical erythromycin reduced from 70% in 1975 to 25% in 2000 with widespread usage [21], which led to a decreased therapeutic effect and its infrequent use in acne treatment now [22]. Cyclines are mostly used for moderate to severe acne and remain effective against C. acnes, which is demonstrated in this study since no strains of minocycline- or tetracycline-resistant C. acnes were observed. Quinolones are used for acne in some regions [23], but it is not recommended to use these potent antibiotics in acne because they should be reserved for serious conditions such as infections by methicillin-resistant Staphylococcus aureus [24]. Fusidic acid is an antibiotic targeting gram-positive cocci and has been proven to have antibacterial and immunomodulatory effects [25]. In spite of the rare emergence of resistance and good antimicrobial susceptibility of fusidic acid, the application of fusidic acid in acne treatment is considered overused and is still disputed [26]. Moreover, it should be noted that three moxifloxacin-resistant C. acnes strains also showed resistance to both erythromycin and clindamycin, indicating multidrug resistance (MDR), which is defined as non-susceptibility to at least one agent in three or more antimicrobial categories [27]. Current research indicates that overexpression of efflux pumps may be an important mechanism of MDR [28]. However, the role of the efflux pump in multidrug-resistant C. acnes remains unknown. On the whole, dermatologists should strictly follow the application principle and specification of antibiotics in treating acne to prevent the rise of drug-resistant strains.
As early as 1972, Johnson and Cummins [29] classified C. acnes into serotypes I and II according to the findings from the agglutination test. In addition, they found serotype I had galactose as a cell wall sugar, while serotype II lacked galactose. Then, in 1979, Kishishita et al. [10] compared the biochemical and serologic characteristics of 128 C. acnes strains from healthy volunteers with 3 standard strains (ATCC 6919, 11827 and 11828). They classified C. acnes into five biotypes (BI to BV) according to the fermentation tests of ribose, erythritol and sorbitol. Several studies [8, 30] have indicated that BIII is the predominant biotype of C. acnes, probably due to its higher lipase activity than other biotypes. Higaki et al. [8] investigated the relationship between the severity of acne and C. acnes biotype, finding that BIII was the major biotype of C. acnes in all levels of acne severity but most commonly isolated from severe acne lesions. However, the sample sizes in these studies are generally small, which is inadequate for accurate conclusions. In our study, among the 63 isolated C. acnes strains, BIII was the most common biotype (50.8%), consistent with previous studies. BIII was also the dominant biotype in severe (57.9%) and moderate acne (53.8%). It is speculated that C. acnes of BIII has some specific characteristics, such as production of free fatty acids, which may contribute to its growth and proliferation in severe acne lesions [8]. Additionally, our results suggested that different degrees of acne severity reflected different biotype distributions, and the biotype distribution of mild acne was different from that of moderate acne.
Multilocus sequence typing (MLST) of C. acnes currently represents the clear gold standard for accurate phylotype identification, but its expensive, time-consuming and technically undemanding characteristics restrict the application. Consequently, Barnard et al. [11] developed multiplex touchdown PCR as a rapid, precise, high-throughput typing method for phylogenetic classification. It can identify the six highly distinct evolutionary phylotypes of C. acnes in a single reaction, known as IA1, IA2, IB, IC, II and III, which display differences in inflammatory properties, production of virulence determinants and association with various conditions [31,32,33]. We utilized this technique, and our study revealed that IA1 was the predominant phylotype of skin C. acnes isolates, consistent with former researches [34,35,36]. Neuraminidase and dermatan-sulfate binding adhesins DsA1 and DsA2 were assumed to be virulence factors of type IA division on account of its exclusive association with acne and the recurrent nature of the disease [9, 37].
Furthermore, we studied the biotype and phyotype distribution of resistant and susceptible C. acnes. Biotype BIII and phylotype IA1 took the lead in both groups. BIII occupied the main biotype status of antibiotic-susceptible and -resistant C. acnes, including erythromycin (53.1% vs. 48.4%), clindamycin (48.6% vs. 55.6%) and moxifloxacin (50.9% vs. 50.0%). IA1 is also the major phylotype of isolated C. acnes strains susceptible and resistant to erythromycin (71.9% vs. 71.0%), clindamycin (70.3% vs. 77.8%) and moxifloxacin (71.2% vs. 75.0%). The three MDR strains were classified into the biotype BII, BIII and BV group each and two strains into IA1 as well as one strain into the IA2 group of phylotypes. Among the different approaches utilized by C. acnes to confer resistance to antibiotics, specific point mutations in the rRNA operon represent a major mechanism. McDowell et al. [35] conducted MLST analysis of 40 antibiotic-resistant C. acnes strains and found 95% strains had at least one specific point mutation in the rRNA operon. Besides, 85% of the resistant isolates belonged to the type IA1 group. Nevertheless, it is difficult to draw any definitive conclusions on the significance of these data until a wider prospective study is conducted and a greater number of isolates are analyzed. In our study, there was no significant difference in the distribution of biotypes or phylotypes between susceptible and resistant C. acnes.
Conclusions
Increases in C. acnes resistance have become a crucial issue in the treatment of acne. Resistance to erythromycin and clindamycin is common, while cyclines still have good antimicrobial susceptibility. BIII group and IA1 group are the major biotype and phylotype of C. acnes, respectively, and the predominant types in moderate and severe acne as well. Different degrees of acne severity also reflect different biotype and phylotype distributions. Our findings may provide evidence for clinical studies about drug resistance and help elucidate the biologic and phylogenetic characteristics of C. acnes.
References
Williams HC, Dellavalle RP, Garner S. Acne vulgaris. Lancet. 2012;379:361–72.
Scholz CF, Kilian M. The natural history of cutaneous propionibacteria, and reclassification of selected species within the genus Propionibacterium to the proposed novel genera Acidipropionibacterium gen. nov., Cutibacterium gen. nov. and Pseudopropionibacterium gen. nov. Int J Syst Evol Microbiol. 2016;66:4422–32.
Beylot C, Auffret N, Poli F, et al. Propionibacterium acnes: an update on its role in the pathogenesis of acne. J Eur Acad Dermatol Venereol. 2014;28:271–8.
Del Rosso JQ, Kircik LH. The sequence of inflammation, relevant biomarkers, and the pathogenesis of acne vulgaris: what does recent research show and what does it mean to the clinician? J Drugs Dermatol. 2013;12:s109–15.
Humphrey S. Antibiotic resistance in acne treatment. Skin Ther Lett. 2012;17:1–3.
Walsh TR, Efthimiou J, Dreno B. Systematic review of antibiotic resistance in acne: an increasing topical and oral threat. Lancet Infect Dis. 2016;16:e23–33.
McDowell A, Nagy I, Magyari M, Barnard E, Patrick S. The opportunistic pathogen Propionibacterium acnes: insights into typing, human disease, clonal diversification and CAMP factor evolution. PLoS One. 2013;8:e70897.
Higaki S, Nakamura M, Morohashi M, Yamagishi T. Propionibacterium acnes biotypes and susceptibility to minocycline and Keigai-rengyo-to. Int J Dermatol. 2004;43:103–7.
Lomholt HB, Kilian M. Population genetic analysis of Propionibacterium acnes identifies a subpopulation and epidemic clones associated with acne. PLoS One. 2010;5:e12277.
Kishishita M, Ushijima T, Ozaki Y, Ito Y. Biotyping of Propionibacterium acnes isolated from normal human facial skin. Appl Environ Microbiol. 1979;38:585–9.
Barnard E, Nagy I, Hunyadkurti J, Patrick S, McDowell A. Multiplex touchdown PCR for rapid typing of the opportunistic pathogen Propionibacterium acnes. J Clin Microbiol. 2015;53:1149–55.
Fitz-Gibbon S, Tomida S, Chiu BH, et al. Propionibacterium acnes strain populations in the human skin microbiome associated with acne. J Invest Dermatol. 2013;133:2152–60.
Thiboutot D, Gollnick H, Bettoli V, et al. New insights into the management of acne: an update from the Global Alliance to Improve Outcomes in Acne group. J Am Acad Dermatol. 2009;60:S1–50.
Leyden JJ. Antibiotic resistant acne. Cutis. 1976;17:593–6.
Crawford WW, Crawford IP, Stoughton RB, Cornell RC. Laboratory induction and clinical occurrence of combined clindamycin and erythromycin resistance in Corynebacterium acnes. J Invest Dermatol. 1979;72:187–90.
Dumont-Wallon G, Moyse D, Blouin E, Dreno B. Bacterial resistance in French acne patients. Int J Dermatol. 2010;49:283–8.
Ross JI, Snelling AM, Carnegie E, et al. Antibiotic-resistant acne: lessons from Europe. Br J Dermatol. 2003;148:467–78.
Nakase K, Nakaminami H, Takenaka Y, Hayashi N, Kawashima M, Noguchi N. Relationship between the severity of acne vulgaris and antimicrobial resistance of bacteria isolated from acne lesions in a hospital in Japan. J Med Microbiol. 2014;63:721–8.
Luk NM, Hui M, Lee HC, et al. Antibiotic-resistant Propionibacterium acnes among acne patients in a regional skin centre in Hong Kong. J Eur Acad Dermatol Venereol. 2013;27:31–6.
Schafer F, Fich F, Lam M, Garate C, Wozniak A, Garcia P. Antimicrobial susceptibility and genetic characteristics of Propionibacterium acnes isolated from patients with acne. Int J Dermatol. 2013;52:418–25.
Tzellos T, Zampeli V, Makrantonaki E, Zouboulis CC. Treating acne with antibiotic-resistant bacterial colonization. Expert Opin Pharmacother. 2011;12:1233–47.
Austin BA, Fleischer AB Jr. The extinction of topical erythromycin therapy for acne vulgaris and concern for the future of topical clindamycin. J Dermatolog Treat. 2017;28:145–8.
Kawada A, Wada T, Oiso N. Clinical effectiveness of once-daily levofloxacin for inflammatory acne with high concentrations in the lesions. J Dermatol. 2012;39:94–6.
Jacobs MR, Appelbaum PC. Nadifloxacin: a quinolone for topical treatment of skin infections and potential for systemic use of its active isomer, WCK 771. Expert Opin Pharmacother. 2006;7:1957–66.
Christiansen K. Fusidic acid non-antibacterial activity. Int J Antimicrob Agents. 1999;12(Suppl 2):S73–8.
Farrell DJ, Castanheira M, Chopra I. Characterization of global patterns and the genetics of fusidic acid resistance. Clin Infect Dis. 2011;52(Suppl 7):S487–92.
Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18:268–81.
Hernando-Amado S, Blanco P, Alcalde-Rico M, et al. Multidrug efflux pumps as main players in intrinsic and acquired resistance to antimicrobials. Drug Resist Updates. 2016;28:13–27.
Johnson JL, Cummins CS. Cell wall composition and deoxyribonucleic acid similarities among the anaerobic coryneforms, classical propionibacteria, and strains of Arachnia propionica. J Bacteriol. 1972;109:1047–66.
Higaki S, Kitagawa T, Kagoura M, Morohashi M, Yamagishi T. Correlation between Propionibacterium acnes biotypes, lipase activity and rash degree in acne patients. J Dermatol. 2000;27:519–22.
McDowell A, Perry AL, Lambert PA, Patrick S. A new phylogenetic group of Propionibacterium acnes. J Med Microbiol. 2008;57:218–24.
Nagy I, Pivarcsi A, Kis K, et al. Propionibacterium acnes and lipopolysaccharide induce the expression of antimicrobial peptides and proinflammatory cytokines/chemokines in human sebocytes. Microbes Infect. 2006;8:2195–205.
Valanne S, McDowell A, Ramage G, et al. CAMP factor homologues in Propionibacterium acnes: a new protein family differentially expressed by types I and II. Microbiology. 2005;151:1369–79.
Cebrian R, Arevalo S, Arias-Santiago S, et al. Optimization of genotypic and biochemical methods to profile P. acnes isolates from a patient population. J Microbiol Methods. 2017;141:17–24.
McDowell A, Barnard E, Nagy I, et al. An expanded multilocus sequence typing scheme for propionibacterium acnes: investigation of ‘pathogenic’, ‘commensal’ and antibiotic resistant strains. PLoS One. 2012;7:e41480.
Yu Y, Champer J, Garban H, Kim J. Typing of Propionibacterium acnes: a review of methods and comparative analysis. Br J Dermatol. 2015;172:1204–9.
McDowell A, Gao A, Barnard E, et al. A novel multilocus sequence typing scheme for the opportunistic pathogen Propionibacterium acnes and characterization of type I cell surface-associated antigens. Microbiology. 2011;157:1990–2003.
Acknowledgements
We thank the participants of the study.
Funding
This study was funded by the National Natural Science Foundation of China (no. 81402615), a scientific research project supported by Huashan Hospital, Fudan University (2013QD07), the major research and development project of innovative drugs, Ministry of Science and Technology (2017ZX09304005), and the Health Skin and Hair Research of Chinese Medical Association-L’Oreal. The National Natural Science Foundation of China also funded the Rapid Service Fees.
Authorship
All named authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this article, take responsibility for the integrity of the work as a whole and have given their approval for this version to be published.
Disclosures
Nanxue Zhang, Ruoyue Yuan, Kevin Z Xin, Zhong Lu and Ying Ma have nothing to disclose.
Compliance with Ethics Guidelines
This study was approved by the Ethics Committee of Huashan Hospital of Fudan University. This study was performed in accordance with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Informed consent to participate was obtained from all individual participants included in the study.
Data Availability
The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Enhanced Digital Features
To view enhanced digital features for this article go to https://doi.org/10.6084/m9.figshare.9676064.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License ( http://creativecommons.org/licenses/by-nc/4.0/ ), which permits any noncommercial use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zhang, N., Yuan, R., Xin, K.Z. et al. Antimicrobial Susceptibility, Biotypes and Phylotypes of Clinical Cutibacterium (Formerly Propionibacterium) acnes Strains Isolated from Acne Patients: An Observational Study. Dermatol Ther (Heidelb) 9, 735–746 (2019). https://doi.org/10.1007/s13555-019-00320-7
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13555-019-00320-7