Abstract
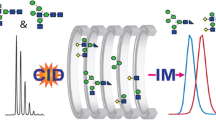
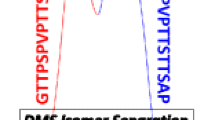
Presence of unresolved ion mobility (IM) profiles limits the efficient utilization of IM mass spectrometry (IM-MS) systems for isomer differentiation. Here, we introduce an automated ion mobility deconvolution (AIMD) computer software for streamlined deconvolution of overlapped IM-MS profiles. AIMD is based on a previously reported post-IM/collision-induced dissociation (CID) deconvolution approach [J. Am. Soc. Mass Spectrom. 23, 1873 (2012)] and, unlike the previously reported manual approach, it does not require resampling of post-IM/CID data. A novel data preprocessing approach is utilized to improve the accuracy and efficiency of the deconvolution process. Results from AIMD analysis of overlapped IM profiles of data from (1) Waters Synapt G1 for a binary mixture of isomeric peptides (amino acid sequences: GRGDS and SDGRG) and (2) Waters Synapt G2-S for a binary mixture of isomeric trisaccharides (raffinose and isomaltotriose) are presented.

ᅟ




Similar content being viewed by others
References
Han, X., Yang, K., Gross, R.W.: Multidimensional mass spectrometry-based shotgun lipidomics and novel strategies for lipidomic analyses. Mass Spectrom. Rev. 31, 134–178 (2012)
Mishur, R.J., Rea, S.L.: Applications of mass spectrometry to metabolomics and metabonomics: detection of biomarkers of aging and of age-related diseases. Mass Spectrom. Rev. 31, 70–95 (2012)
Marshall, A.G., Rodgers, R.P.: Petroleomics: chemistry of the underworld. Proc. Natl. Acad. Sci. U.S.A. 105, 18090–18095 (2008)
Bensimon, A., Heck, A.J., Aebersold, R.: Mass spectrometry-based proteomics and network biology. Annu. Rev. Biochem. 81, 379–405 (2012)
Domon, B., Aebersold, R.: Mass spectrometry and protein analysis. Science 312, 212–217 (2006)
Chait, B.T.: Mass spectrometry: bottom-up or top-down? Science 314, 65–66 (2006)
Konermann, L., Pan, J., Liu, Y.H.: Hydrogen exchange mass spectrometry for studying protein structure and dynamics. Chem. Soc. Rev. 40, 1224–1234 (2011)
Marshall, A.G., Hendrickson, C.L., Jackson, G.S.: Fourier transform ion cyclotron resonance mass spectrometry: a primer. Mass Spectrom. Rev. 17, 1–35 (1998)
Xian, F., Hendrickson, C.L., Marshall, A.G.: High resolution mass spectrometry. Anal. Chem. 84, 708–719 (2012)
Wenger, C.D., McAlister, G.C., Xia, Q., Coon, J.J.: Sub-part-per-million precursor and product mass accuracy for high-throughput proteomics on an electron transfer dissociation-enabled orbitrap mass spectrometer. Mol. Cell. Proteom. 9, 754–763 (2010)
Peterson, A.C., McAlister, G.C., Quarmby, S.T., Griep-Raming, J., Coon, J.J.: Development and characterization of a GC-enabled QLT-Orbitrap for high-resolution and high-mass accuracy GC/MS. Anal. Chem. 82, 8618–8628 (2010)
Fattahi, A., Solouki, T.: Conformational analysis of metal complexed model peptides and their fragment ions using FT-ICR MS and gas-phase H/D exchange reactions. Proceedings of the 49th ASMS Conference on Mass Spectrometry and Allied Topics. Chicago, IL, 27–31 May 2001
Fattahi, A., Zekavat, B., Solouki, T.: H/D exchange kinetics: experimental evidence for formation of different b fragment ion conformers/isomers during the gas-phase peptide sequencing. J. Am. Soc. Mass Spectrom. 21, 358–369 (2010)
Solouki, T., Szulejko, J.E., Bennett, J.B., Graham, L.B.: A preconcentrator coupled to a GC/FTMS: advantages of self-chemical ionization, mass measurement accuracy, and high mass resolving power for GC applications. J. Am. Soc. Mass Spectrom. 15, 1191–1200 (2004)
Solouki, T., Szulejko, J.E.: Bimolecular and unimolecular contributions to the disparate self-chemical ionizations of alpha-pinene and camphene isomers. J. Am. Soc. Mass Spectrom. 18, 2026–2039 (2007)
Mondello, L., Tranchida, P.Q., Dugo, P., Dugo, G.: Comprehensive two-dimensional gas chromatography–mass spectrometry: a review. Mass Spectrom. Rev. 27, 101–124 (2008)
Donato, P., Cacciola, F., Tranchida, P.Q., Dugo, P., Mondello, L.: Mass spectrometry detection in comprehensive liquid chromatography: basic concepts, instrumental aspects, applications, and trends. Mass Spectrom. Rev. 31, 523–559 (2012)
Kanu, A.B.., Dwivedi, P., Tam, M., Matz, L., Hill Jr., H.H.: Ion mobility-mass spectrometry. J. Mass Spectrom. 43, 1–22 (2008)
Wyttenbach, T., Pierson, N.A., Clemmer, D.E., Bowers, M.T.: Ion mobility analysis of molecular dynamics. Annu. Rev. Phys. Chem. 65, 175–196 (2014)
Rokushika, S., Hatano, H., Bairn, M.A., Hill, H.H.: Resolution measurement for ion mobility spectrometry. Anal. Chem. 57, 1902–1907 (1985)
Siems, W.F., Wu, C., Tarver, E.E., Hill, H.H.: Measuring the resolving power of ion mobility spectrometers. Anal. Chem. 66, 4195–4201 (1995)
Shvartsburg, A.A., Smith, R.D.: Fundamentals of traveling wave ion mobility spectrometry. Anal. Chem. 80, 9689–9699 (2008)
Clowers, B.H., Dwivedi, P., Steiner, W.E., Hill Jr., H.H., Bendiak, B.: Separation of sodiated isobaric disaccharides and trisaccharides using electrospray ionization-atmospheric pressure ion mobility-time of flight mass spectrometry. J. Am. Soc. Mass Spectrom. 16, 660–669 (2005)
Dwivedi, P., Wu, C., Matz, L.M., Clowers, B.H., Siems, W.F., Hill Jr., H.H.: Gas-phase chiral separations by ion mobility spectrometry. Anal. Chem. 78, 8200–8206 (2006)
Dwivedi, P., Bendiak, B., Clowers, B.H., Hill Jr., H.H.: Rapid resolution of carbohydrate isomers by electrospray ionization ambient pressure ion mobility spectrometry-time-of-flight mass spectrometry (ESI-APIMS-TOFMS). J. Am. Soc. Mass Spectrom. 18, 1163–1175 (2007)
Shvartsburg, A.A., Singer, D., Smith, R.D., Hoffmann, R.: Ion mobility separation of isomeric phosphopeptides from a protein with variant modification of adjacent residues. Anal. Chem. 83, 5078–5085 (2011)
Ibrahim, Y.M., Shvartsburg, A.A., Smith, R.D., Belov, M.E.: Ultrasensitive identification of localization variants of modified peptides using ion mobility spectrometry. Anal. Chem. 83, 5617–5623 (2011)
Shvartsburg, A.A., Smith, R.D.: Accelerated high-resolution differential ion mobility separations using hydrogen. Anal. Chem. 83, 9159–9166 (2011)
Li, H., Giles, K., Bendiak, B., Kaplan, K., Siems, W.F., Hill Jr., H.H.: Resolving structural isomers of monosaccharide methyl glycosides using drift tube and traveling wave ion mobility mass spectrometry. Anal. Chem. 84, 3231–3239 (2012)
Fasciotti, M., Sanvido, G.B., Santos, V.G., Lalli, P.M., McCullagh, M., de Sa, G.F., Daroda, R.J., Peter, M.G., Eberlin, M.N.: Separation of isomeric disaccharides by traveling wave ion mobility mass spectrometry using CO2 as drift gas. J. Mass Spectrom. 47, 1643–1647 (2012)
Liu, Y., Clemmer, D.E.: Characterizing oligosaccharides using injected-ion mobility/mass spectrometry. Anal. Chem. 69, 2504–2509 (1997)
Badman, E.R., Hoaglund-Hyzer, C.S., Clemmer, D.E.: Dissociation of different conformations of ubiquitin ions. J. Am. Soc. Mass Spectrom. 13, 719–723 (2002)
Zucker, S.M., Lee, S., Webber, N., Valentine, S.J., Reilly, J.P., Clemmer, D.E.: An ion mobility/ion trap/photodissociation instrument for characterization of ion structure. J. Am. Soc. Mass Spectrom. 22, 1477–1485 (2011)
Lee, S., Li, Z., Valentine, S.J., Zucker, S.M., Webber, N., Reilly, J.P., Clemmer, D.E.: Extracted fragment ion mobility distributions: A new method for complex mixture analysis. Int. J. Mass Spectrom. 309, 154–160 (2012)
Silveira, J.A., Fort, K.L., Kim, D., Servage, K.A., Pierson, N.A., Clemmer, D.E., Russell, D.H.: From solution to the gas phase: stepwise dehydration and kinetic trapping of Substance P reveals the origin of peptide conformations. J. Am. Chem. Soc. 135, 19147–19153 (2013)
Zekavat, B., Solouki, T.: Chemometric data analysis for deconvolution of overlapped ion mobility profiles. J. Am. Soc. Mass Spectrom. 23, 1873–1884 (2012)
Windig, W., Guilment, J.: Interactive self-modeling mixture analysis. Anal. Chem. 63, 1425–1432 (1991)
Zekavat, B., Miladi, M., Becker, C., Munisamy, S.M., Solouki, T.: Combined use of post-ion mobility/collision-induced dissociation and chemometrics for b fragment ion analysis. J. Am. Soc. Mass Spectrom. 24, 1355–1365 (2013)
Zekavat, B., Miladi, M., Al-Fdeilat, A.H., Somogyi, A., Solouki, T.: Evidence for sequence scrambling and divergent H/D exchange reactions of doubly-charged isobaric b-type fragment ions. J. Am. Soc. Mass Spectrom. 25, 226–236 (2014)
Hoffmann, W., Hofmann, J., Pagel, K.: Energy-resolved ion mobility-mass spectrometry-A concept to improve the separation of isomeric carbohydrates. J. Am. Soc. Mass Spectrom. 25, 471–479 (2014)
Pedrioli, P.G., Eng, J.K., Hubley, R., Vogelzang, M., Deutsch, E.W., Raught, B., Pratt, B., Nilsson, E., Angeletti, R.H., Apweiler, R., Cheung, K., Costello, C.E., Hermjakob, H., Huang, S., Julian, R.K., Kapp, E., McComb, M.E., Oliver, S.G., Omenn, G., Paton, N.W., Simpson, R., Smith, R., Taylor, C.F., Zhu, W., Aebersold, R.: A common open representation of mass spectrometry data and its application to proteomics research. Nat. Biotechnol. 22, 1459–1466 (2004)
Hoyes, J.B., Bateman, R.H., Wildgoose, J.L.: A high resolution Orthogonal TOF with selectable drift length. Proceedings of the 48th ASMS Conference on Mass Spectrometry and Allied Topics. Long Beach, CA, 11–15 June 2000
Bartmess, J.E., Georgiadis, R.M.: Empirical methods for determination of ionization gauge relative ensitivities for different gases. Vacuum 33, 149–153 (1983)
Roepstorff, P., Fohlman, J.: Proposal for a common nomenclature for sequence ions in mass spectra of peptides. Biomed. Mass Spectrom. 11, 601 (1984)
Domon, B., Costello, C.E.: A systematic nomenclature for carbohydrate fragmentations in FAB-MS/MS spectra of glycoconjugates. Glycoconj. J. 5, 397–409 (1988)
Kessner, D., Chambers, M., Burke, R., Agus, D., Mallick, P.: ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008)
Chambers, M.C., Maclean, B., Burke, R., Amodei, D., Ruderman, D.L., Neumann, S., Gatto, L., Fischer, B., Pratt, B., Egertson, J., Hoff, K., Kessner, D., Tasman, N., Shulman, N., Frewen, B., Baker, T.A., Brusniak, M.Y., Paulse, C., Creasy, D., Flashner, L., Kani, K., Moulding, C., Seymour, S.L., Nuwaysir, L.M., Lefebvre, B., Kuhlmann, F., Roark, J., Rainer, P., Detlev, S., Hemenway, T., Huhmer, A., Langridge, J., Connolly, B., Chadick, T., Holly, K., Eckels, J., Deutsch, E.W., Moritz, R.L., Katz, J.E., Agus, D.B., MacCoss, M., Tabb, D.L., Mallick, P.: A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012)
Malinowski, E.R.: Determination of the number of factors and the experimental error in a data matrix. Anal. Chem. 49, 612–617 (1977)
Meloun, M., Čapek, J., Mikšík, P., Brereton, R.G.: Critical comparison of methods predicting the number of components in spectroscopic data. Anal. Chim. Acta. 423, 51–68 (2000)
Tauler, R., Smilde, A., Kowalski, B.: Selectivity, local rank, three-way data analysis and ambiguity in multivariate curve resolution. J. Chemom. 9, 31–58 (1995)
Giles, K., Williams, J.P., Campuzano, I.: Enhancements in travelling wave ion mobility resolution. Rapid Commun. Mass Spectrom. 25, 1559–1566 (2011)
Acknowledgments
The authors greatly appreciate financial support provided by Baylor University. They also thank Dr. Christopher Becker and Dr. C. Kevin Chambliss of Baylor University for providing the sugar samples used in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 51 kb)
Rights and permissions
About this article
Cite this article
Brantley, M., Zekavat, B., Harper, B. et al. Automated Deconvolution of Overlapped Ion Mobility Profiles. J. Am. Soc. Mass Spectrom. 25, 1810–1819 (2014). https://doi.org/10.1007/s13361-014-0963-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-014-0963-3