Abstract
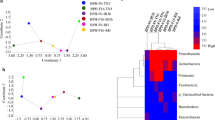
Nilaparvata lugens (Stål) is one of the major pests of rice throughout tropical and temperate Asia. Indiscriminate use of insecticides for suppressing N. lugens has resulted in the development of resistance to multiple insecticide classes, causing frequent control failures in the field. Analysis of gut bacterial diversity within an insect host is the initial step towards understanding the ecological roles of the symbionts. Present study aimed to survey the bacterial diversity associated with laboratory-reared (insecticide-susceptible) and field-collected (insecticide-resistant) populations of N. lugens by culture-dependent and PCR-Denaturing Gradient Gel Electrophoresis (DGGE) methods. Seventeen bacterial isolates were obtained by the culture-dependent method. Molecular characterization using the 16S rRNA gene and phylogenetic analysis revealed that the isolates belonged to Firmicutes and Proteobacteria. Taxonomic assignment placed these isolates into seven families representing 10 genera. Enterobacteriaceae was the most dominant family with its occurrence in four out of the five populations studied. The DGGE profiles indicated a low complex gut bacteria associated with N. lugens with limited number of bands. The Shannon-Wiener index ranged from 0.898 in insecticide-susceptible population to 0.946–1.035 in resistant populations. Sequencing and phylogenetic analysis revealed that the DGGE bands belonged to Firmicutes, Proteobacteria and Bacteriodetes. Results of this study illustrated that gut bacterial community associated with N. lugens is dominated by Proteobacteria and Firmicutes. Present findings could provide the basis for future work on the possible role of the bacterial symbionts in insecticide resistance and to formulate potential resistance management strategies.


Similar content being viewed by others
References
Anand, A. A. P., Vennison, S. J., Sankar, S. G., Prabhu, D. I. G., Vasan, P. T., Raghuraman, T., Geoffrey, C. J., & Vendan, S. E. (2010). Isolation and characterization of bacteria from the gut of Bombyx mori that degrade cellulose, xylan, pectin and starch and their impact on digestion. Journal of Insect Science, 10, 107.
Basanth, Y. S., Sannaveerappanavar, V. T., & Gowda, D. S. (2013). Susceptibility of different populations of Nilaparvata lugens from major rice growing areas of Karnataka, India to different groups of insecticides. Rice Science, 20, 371–378.
Broderick, N. A., Raffa, K. F., Goodman, R. M., & Handelsman, J. (2004). Census of the bacterial community of the gypsy moth larval midgut by using culturing and culture-independent methods. Applied and Environmental Microbiology, 70, 293–300.
Dillon, R. J., & Dillon, V. M. (2004). The gut bacteria of insects: nonpathogenic interactions. Annual Reviews of Entomology, 49, 71–92.
Dillon, R. J., Webster, G., Weightman, A. J., Dillon, V. M., Blanford, S., & Charnley, A. K. (2008). Composition of Acridid gut bacterial communities as revealed by 16S rRNA gene analysis. Journal of Invertebrate Pathology, 97, 265–272.
El-Sayed, W. S., & Ibrahim, R. A. (2015). Diversity and phylogenetic analysis of endosymbiotic bacteria of the date palm root borer Oryctes agamemnon (Coleoptera: Scarabaeidae). BMC Microbiology, 15, 88.
Engel, P., & Moran, N. A. (2013). The gut microbiota of insects–diversity in structure and function. FEMS Microbiology Reviews, 37, 699–735.
Gracy, R. G., Malathi, V. M., Jalali, S. K., Jose, V. L., & Thulasi, A. (2016). Variation in larval gut bacteria between insecticide-resistant and-susceptible populations of Helicoverpa armigera (Hübner) (Lepidoptera: Noctuidae). Phytoparasitica, 44, 477–490.
Hernández, N., Escudero, J. A., Millán, Á. S., González-Zorn, B., Lobo, J. M., Verdú, J. R., & Suárez, M. (2015). Culturable aerobic and facultative bacteria from the gut of the polyphagic dung beetle, Thorectes lusitanicus. Insect Science, 22, 178–190.
Hibino, H. (1996). Biology and epidemiology of rice viruses. Annual Reviews of Phytopathology, 34, 249–274.
Indiragandhi, P., Anandham, R., Madhaiyan, M., Poonguzhali, S., Kim, G. H., Saravanan, V. S., & Sa, T. (2007). Cultivable bacteria associated with larval gut of prothiofos-resistant, prothiofos- susceptible and field-caught populations of diamond back moth, Plutella xylostella and their potential for antagonism towards entomopathogenic fungi and host insect nutrition. Journal of Applied Microbiology, 103, 2664–2675.
Kikuchi, Y., Hayatsu, M., Hosokawa, T., Nagayama, A., Tago, K., & Fukatsu, T. (2012). Symbiont-mediated insecticide resistance. Proceedings of the National Academy Sciences USA, 109, 8618–8622.
Kim, Y. J., Lee, Y. J., Kim, G. H., Lee, S. W., & Ahn, Y. J. (1999). Toxicity of tebufenpyrad to Tetranychus urticae (Acari: Tetranychidae) and Amblyseius womersleyi (Acari: Phytoseiidae) under laboratory and field conditions. Journal of Economic Entomology, 92, 187–192.
Kim, J. Y., Lee, J., Shin, N. R., Yun, J. H., Whon, T. W., Kim, M. S., Jung, M. L., Roh, S. W., Hyun, D. W., & Bae, J. W. (2013). Orbus sasakiae sp. nov., a bacterium isolated from the gut of the butterfly, Sasakia charonda, and emended description of the genus Orbus. International Journal Systematic and Evolutionary Microbiology, 63, 1766–1770.
Kumar, S., Stecher, G., & Tamura, K. (2016). MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33, 1870–1874.
Lane, D. J. (1991). 16S/23S rRNA sequencing. In E. Stackebrandt & M. Goodfellow (Eds.), Nucleic acid techniques in bacterial systematics (pp. 115–147). New York: Wiley.
Larkin, M. A., Blackshields, G., Brown, N. P., Chenna, R., McGettigan, P. A., McWilliam, H., Valentin, F., Wallace, I. M., Wilm, A., Lopez, R., Thompson, J. D., Gibson, T. J., & Higgins, D. G. (2007). ClustalW and ClustalX version 2. Bioinformatics, 23, 2947–2948.
Lin, X. L., Pan, Q. J., Tian, H. G., Douglas, A. E., & Liu, T. X. (2015). Bacteria abundance and diversity of different life stages of Plutella xylostella (Lepidoptera: Plutellidae), revealed by bacteria culture-dependent and PCR-DGGE methods. Insect Science, 22, 375–385.
Lindh, J. M., Terenius, O.’ Faye, I. (2005). 16S rRNA gene-based identification of midgut bacteria from field-caught Anopheles gambiae sensu lato and A. funestus mosquitoes reveals new species related to known insect symbionts. Applied and Environmental Microbiology, 71, 7217–7223.
Malathi, V. M., Jalali, S. K., Sidde Gowda, D. K., Mohan, M., & Venkatesan, T. (2015). Establishing the role of detoxifying enzymes in field-evolved resistance to various insecticides in the brown planthopper, (Nilaparvata lugens) in South India. Insect Science. https://doi.org/10.1111/1744-7917.12254.
Min, S., Lee, S. W., Choi, B. R., Lee, S. H., & Kwon, D. H. (2014). Insecticide resistance monitoring and correlation analysis to select appropriate insecticides against Nilaparvata lugens (Stål), a migratory pest in Korea. Journal of Asia-Pacific Entomology, 17, 711–716.
Mrazek, J., Stosova, L., Fliegerova, K., Kott, T., & Kopecny, J. (2008). Diversity of insect intestinal microflora. Folia Microbiologica, 53, 229–233.
Nakamura, N., Kawai, S., Yukuhiro, F., Ito, S., Gotoh, T., Kisimoto, R., Yanase, T., Matsumoto, Y., Kageyama, D., & Noda, H. (2009). Prevalence of Cardinium bacteria in planthoppers and spider mites and taxonomic revision of “Candidatus Cardinium hertigii” based on detection of a new Cardinium group from biting midges. Applied and Environmental Microbiology, 75, 6757–6763.
Ramya, S. L., Venkatesan, T., Murthy, K. S., Jalali, S. K., & Varghese, A. (2016). Degradation of acephate by Enterobacter asburiae, Bacillus cereus and Pantoea agglomerans isolated from diamondback moth, Plutella xylostella (L), a pest of cruciferous crops. Journal of Environmental Biology, 37, 611–618.
Reeson, A. F., Jankovic, T., Kasper, M. L., Rogers, S., & Austin, A. D. (2003). Application of 16S rDNA- DGGE to examine the microbial ecology associated with a social wasp, Vespula germanica. Insect Molecular Biology, 12, 85–91.
Sánchez, E., Donat, E., Ribes-Koninckx, C., Fernández-Murga, M. L., & Sanz, Y. (2013). Duodenal- mucosal bacteria associated with celiac disease in children. Applied and Environmental Microbiology, 79, 5472–5479.
Sanguinetti, C. J., Neto, E. D., & Simpson, A. J. G. (1994). Rapid silver staining and recovery of PCR products separated on polyacrylamide gels. BioTechniques, 17, 915–919.
Sasaki, M. (1996). Nitrogen recycling in the brown planthopper, Nilaparvata lugens: involvement of yeast-like endosymbionts in uric acid metabolism. Journal of Insect Physiology, 42, 125–129.
Shannon, C. E., & Weaver, W. (1963). The mathematical theory of communication. Urbana: University of Illinois Press.
Simpson, E. H. (1949). Measurement of diversity. Nature, 163, 688.
Singh, B. K., & Walker, A. (2006). Microbial degradation of organophosphorus compounds. FEMS Microbiology Reviews, 30, 428–471.
Tang, M., Lv, L., Jing, S. L., Zhu, L. L., & He, G. C. (2010). Bacterial symbionts of the Brown planthopper, Nilaparvata lugens (Homoptera: Delphacidae). Applied and Environmental Microbiology, 76, 1740–1745.
van den Bosch, T. J., & Welte, C. U. (2016). Detoxifying symbionts in agriculturally important pest insects. Microbial Biotechnology. https://doi.org/10.1111/1751-7915.12483.
Xia, X., Zheng, D., Zhong, H., Qin, B., Gurr, G. M., Vasseur, L., Lin, H., Bai, J., He, W., & You, M. (2013). DNA sequencing reveals the midgut microbiota of DBM, Plutella xylostella (L.) and a possible relationship with insecticide resistance. PLoS One, 8, e68852.
Xu, H. X., Zheng, X. S., Yang, Y. J., Wang, X., Ye, G. Y., & Lu, Z. X. (2014). Bacterial community in different populations of Rice Brown Planthopper, Nilaparvata lugens (Stål). Rice Science, 21, 59–64.
Xue, J., Zhou, X., Zhang, C. X., Yu, L., Fan, H., Wang, Z., Xu, H. J., Xi, Y., Zhu, Z. R., Zhou, W. W., Pan, P. L., Li, B. L., et al. (2014). Genomes of the rice pest brown planthopper and its endosymbionts reveal complex complementary contributions for host adaptation. Genome Biology. https://doi.org/10.1186/s13059-014-0521-0.
Acknowledgements
We acknowledge late Prof. D. K. Sidde Gowda, for his assistance and advice in the rearing of Nilaparvata lugens. Authors thank M/s Applied Maths, Germany, for providing evaluation license of Bionumerics software. The first author gratefully acknowledges the Department of Science and Technology (DST), Govt. of India, for the award of INSPIRE fellowship for Ph.D. work. We are thankful to staff of Regional Agricultural Research Station, Warangal and Nellore and Krishi Vigyan Kendra, Tiruchirappalli, for their kind help extended during field collection of Nilaparvata lugens. The Director, ICAR-NBAIR and the Director, ICAR-NIANP, Bengaluru, India, are thanked for providing necessary laboratory facilities for conducting this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Supplementary Fig. 1
Phylogenetic tree of partial 16S rRNA genes showing the cultured bacteria from the guts of different populations of N. lugens constructed by Neighbor-joining method. The numbers at the nodes represent bootstrap values out of 1000 replications. (GIF 56 kb)
ESM 1
(DOC 58 kb)
ESM 2
(DOC 41 kb)
Rights and permissions
About this article
Cite this article
Malathi, V.M., Jalali, S.K., Lyju, V.J. et al. Associated bacterial diversity of insecticide-susceptible and -resistant brown planthopper, Nilaparvata lugens (Homoptera: Delphacidae) analyzed by culture-dependent and -independent methods. Phytoparasitica 45, 683–693 (2017). https://doi.org/10.1007/s12600-017-0629-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12600-017-0629-3