Abstract
There has long been debate over the origins of dairy consumption within European populations. Whilst it was previously assumed that lactase persistence (LP) was under positive selection following the advent of agriculture, recent genetic studies of prehistoric human remains have revealed LP may have only emerged in Europe in the last 4000 years. These findings stand in contrast to organic residue analysis of Neolithic pottery indicating the utilisation of dairy products, and zooarchaeological mortality profiles consistent with dairying herds at Neolithic sites. The recent discovery of the milk protein β-lactoglobulin (BLG) within human dental calculus presents a new method via which to explore dairy product consumption in the archaeological past. Here, we apply shotgun proteomic analysis to dental calculus samples from three British Neolithic sites, revealing the earliest identification of BLG in human dental calculus to date. The presence of BLG peptides in individuals who are unlikely to possess LP provides new insight into dairying in the British Neolithic, suggesting the potential processing of milk by Neolithic populations to reduce the lactose content of dairy products.
Similar content being viewed by others
Introduction
The British Neolithic is a period which has long been characterised by the arrival and adoption of agriculture and perceived associated sedentary lifestyle, alongside the emergence of new forms of material culture and the construction of a range of monumental forms. Most recently, more support for the hypothesis that this ‘Neolithic package’ was brought to Britain by incoming European farmers has been recognised (Brace et al. 2019). Importantly, the Neolithic heralds a significant shift in subsistence, characterised by the introduction of domesticates into Britain, which included emmer wheat (Triticum dicoccum), einkorn wheat (Triticum momococcum) and barley (Hordeum vulgare), alongside emerging animal husbandry of domesticated species—predominately cattle, pig, sheep and goats (Thomas 1999; Rowley-Conwy 2004; Brown 2007). Our understanding of Neolithic subsistence has been obtained from a range of archaeological evidence, such as the faunal remains of domesticated animals (e.g., Albarella and Serjeantson 2002; Mulville and Grigson 2007), evidence of cereal cultivation—including carbonised cereal grains (e.g., Robinson 2000; Jones and Legge 2008), stable isotope analysis of skeletal material (e.g., Richards and Hedges 1999; Hedges et al. 2006; Richards 2008; Stevens et al. 2012; Schulting 2013), and organic residue analysis of pottery (e.g., Copley et al. 2008; Craig et al. 2015). Nevertheless, whilst we know that British Neolithic diets appear to be (isotopically) dominated by C3 terrestrial plant resources and terrestrial animals, with little to no marine input, there are still aspects of Neolithic subsistence which remain unclear. The contribution of wild plants, cereals, dairy and aquatic foodstuffs to the diet have all been debated (e.g., Robinson 2000; Thomas 2003; Richards et al. 2003; Milner et al. 2004; Rowley-Conwy 2004), and the contribution of crop-derived protein to Neolithic diets has also been suggested to typically be underestimated (Bogaard et al. 2013). The potential variability or homogeneity within British Neolithic diets is also still not fully understood.
Dental calculus, the mineralised bacterial biofilm of dental plaque, is consistently found within Neolithic skeletal assemblages and is now known to be a rich source of ancient biomolecules, notably ancient DNA and proteins (Warinner et al. 2014a, b), which can provide new insights not only into the oral microbiome but also past diet. Metaproteomic analysis of Neolithic dental calculus may therefore provide a new approach to early prehistoric dietary reconstruction, by providing direct and diagnostic evidence of consumed foodstuffs, including the consumption of milk or dairy products. Here we apply metaproteomic analysis to human dental calculus samples from three British Neolithic sites to assess the prevalence of dietary and milk proteins, and its efficacy as a method via which to increase our understanding of Neolithic diets. To date, the calculus from only a single Middle Neolithic individual has yet been analysed using metaproteomic methods (Mays et al. 2018). To our knowledge, this study represents the oldest human dental calculus samples analysed proteomically to date globally.
Dietary proteins within dental calculus
The periodic mineralisation of dental plaque to form calculus serendipitously entraps and preserves ancient biomolecules and microdebris, providing a direct record of consumed foodstuffs. A small number of studies have now explored dietary proteins preserved within human dental calculus. To date, evidence of putative plant proteins originating from oats (Avena sativa), peas (Pisum sativum) and Brassicaceae in archaeological dental calculus have been recovered, as well as a putative faunal blood haemoglobin (HBB) protein, and the milk proteins lactoperoxidase (LPO) and α-S1-casein (CSN1S1) (Hendy et al. 2018a; Jersie-Christensen et al. 2018; Mays et al. 2018). The detection of these proteins showcases how metaproteomic analysis can provide a new direct way of determining dietary information from the archaeological past. The most frequently detected dietary protein within metaproteomic studies of dental calculus has however thus far been the milk protein β-lactoglobulin (BLG).
Lactase persistence and the origins of milk consumption
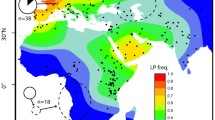
Within the archaeological literature, there has long been a debate over the origins of milk drinking and dairy product consumption (Leonardi et al. 2012). Milk is a significant nutritional resource, containing fats, sugars and proteins, as well as vitamins, minerals and essential amino acids (Bovenhuis et al. 2013). Lactose, a type of disaccharide sugar, is the main carbohydrate in milk, comprising 3.8–5.3% of total content. In order for humans to digest lactose however, it must be broken down by the β-galactosidase lactase-phloritzin hydrolase (more commonly known simply as ‘lactase’; EC 3.2.1.108), which allows it to be absorbed within the intestine (Luinge et al. 1993; Vesa et al. 2000; Ségurel and Bon 2017). As infants, humans have the ability to break down lactose. However, after weaning, the body naturally stops producing lactase, unless the individual has a genetic mutation which allows for its continued production into adulthood—known as lactase persistence (LP). Indeed, LP is thought to be the clearest example of gene-culture coevolution—the idea that cultural practices can alter the genome (Bersaglieri et al. 2004; Leonardi et al. 2012). Currently, around one third of the global population carry an LP mutation, with the highest frequencies (over 75% of the population) being found in Europe, East Africa, West Africa and the Middle East (Ségurel and Bon 2017), the same areas of the globe which have experienced a long history of dairying.
Previously, it has been suggested that the ability to digest raw milk must have provided a selective advantage and that LP would have been under positive selection following the advent of cattle, sheep and goat management and domestication (Cavalli-Sforza 1973; Beja-Pereira et al. 2003; Nielsen et al. 2007). It has generally been assumed that European Neolithic populations began dairying—and therefore also consuming milk products—soon after the introduction of agriculture into the region. These ideas have been traditionally supported by a number of lines of archaeological evidence, particularly organic residue analysis of Neolithic pottery showing the presence of milk lipids (e.g., Copley et al. 2003, 2005a, b; Craig et al. 2005; Salque et al. 2012, 2013; Cramp et al. 2014a, b; Smyth and Evershed 2015), and zooarchaeological analyses and mortality profiling of domesticated fauna (e.g., Legge 2005; Mulville et al. 2005; Vigne 2008; Greenfield and Arnold 2015). In almost all cases, these studies show that dairying was practised as soon as domesticated ruminants were available, i.e. at the start of the Early Neolithic period.
It was therefore commonly assumed that increased reliance on dairy products from the Early Neolithic would have driven the selection of LP during subsequent millennia. At least five different allelic variants have been identified as causative of LP globally, suggesting convergent evolution (Tishkoff et al. 2007; Jones et al. 2015; Liebert et al. 2017). Initial investigations of the European LCT gene responsible for LP (C-13910 > T; −13910*T allele (rs4988235)) suggested that selection favouring the LP allele may have begun in the Neolithic in conjunction with the spread of agriculture and increased use of domesticated animal species. These early studies using statistical approaches via simulation modelling (e.g., Gerbault et al. 2009; Itan et al. 2009), modern microsatellite diversity studies (e.g., Coelho et al. 2005) or modern allele frequencies and extended haplotypes (e.g., Bersaglieri et al. 2004; Myles et al. 2005) suggested that the LCT gene most likely emerged in Europe with the Neolithic transition (Bersaglieri et al. 2004; Coelho et al. 2005; Gerbault et al. 2011).
Advances in next-generation sequencing technologies and the emergence of full genome characterisation of archaeological samples however now suggest that LP may not have been prevalent in the Neolithic, but instead present only in very low frequencies across the population, if present at all. The absence of LP in European Neolithic populations has now been noted in a number of studies (e.g., Burger et al. 2007; Gamba et al. 2014; Witas et al. 2015; Olalde et al. 2018; Brace et al. 2019), and Allentoft et al. (2015) suggest that LP had a very low frequency (5–10%) even in Bronze Age European populations, indicating that LP is the result of more recent positive selection. Similarly, a large-scale study by Mathieson et al. (2015) suggests that LP in Europe only became under strong selection within the last 4000 years. Most recently, none of the 67 British Neolithic individuals analysed in Brace et al. (2019) exhibited LP.
The current genetic evidence, therefore, suggests that European Neolithic populations did not possess LP, and as such, could not have easily digested large quantities of unprocessed dairy products or raw milk. Together, the combined genetic, organic residue analysis and zooarchaeological evidence suggests that whilst people were likely practising dairying in the Neolithic, that either (1) these products served a non-dietary function (i.e., were not being consumed); (2) that dairy products must have been consumed in very small quantities; or (3) were alternatively processed in such a way as to remove the lactose (e.g., through the production of cheeses or other processed dairy products).
The recent discovery of the preservation of the milk protein β-lactoglobulin (BLG) in human dental calculus (Warinner et al. 2014a) provides a method for detecting milk consumption in the archaeological past. Composed of 162 amino acid residues, BLG is the dominant protein within the whey fraction of milk, belonging to the lipocalin family of proteins (Kontopidis et al. 2004; Wal 2004; Monaci et al. 2006). The amino acid sequence of BLG differs between species, meaning it can be used as a species-specific indicator (Kontopidis et al. 2004), but is crucially not found within human milk (Crittenden and Bennett 2005; Restani et al. 2009), thereby excluding a host origin when detected in human dental calculus. BLG is only found in milk, making it a ‘tissue’-specific biomarker, and it is more resistant to microbial attack and enzymatic degradation than other milk proteins (Warinner et al. 2014a). Finally, it has been proposed that BLG could be used as a proxy for lactose, given that they partition together in the whey fraction of milk during processing (Warinner et al. 2014a). If correct, the presence of BLG in Neolithic calculus may provide an additional source of information on Neolithic dairy consumption and the origins of raw milk consumption. Although the presence of BLG in dental calculus has been explored in samples dating back to the Bronze Age (Warinner et al. 2014a) and a single individual from the Middle Neolithic (Mays et al. 2018), applications of metaproteomic analyses to early prehistoric calculus have not yet been widely explored.
Here we present the results of shotgun proteomic analysis by liquid chromatography-tandem mass spectrometry (LC/MS-MS) on human dental calculus samples from three British Neolithic sites (Hambledon Hill, Hazleton North and Banbury Lane; Fig. 1), to explore the presence and prevalence of dietary proteins more generally, and milk proteins specifically, within Neolithic populations. Sites chosen for the analysis date to the Early and Middle Neolithic, a period during which there is extensive evidence of domesticated fauna in the UK and of the utilisation of dairy. The three sites represent a range of site types, for which we have varying amounts of palaeodietary information, and which show differing degrees of evidence for dairying at the site level. Based on recent genetic analyses, individuals from all three sites are unlikely to have had LP (Brace et al. 2019). The goal of the analysis was to explore the presence of dietary proteins within calculus of an Early and Middle Neolithic date in the UK. Furthermore, if milk proteins were detected within the samples, we hoped to further explore the dichotomy in the British archaeological record between the apparent usage of dairy through material culture evidence and zooarchaeological remains, and the seeming absence or very low genetic prevalence of LP.
Site locations of dental calculus samples analysed (map created by the authors using the R package ggmap (Kahle and Wickham 2013))
Materials and methods
Ten dental calculus samples from three British Neolithic sites were analysed using a previously published shotgun metaproteomic approach utilising liquid chromatography-tandem mass spectrometry (LC/MS-MS) (Warinner et al. 2014a, b) (Table 1; Supplementary Information). The ten dental calculus samples analysed comprised of three individuals from the site of Hambledon Hill, five individuals from Hazleton North, and two individuals from Banbury Lane (Fig. 1). Hambledon Hill is an early Neolithic monument complex in Dorset, comprising of two causewayed enclosures, two long barrows and a range of other outer earthworks. The sampled individuals were recovered from both the main enclosure, dated to 3680–3310 cal. BC, and the second, smaller causewayed enclosure (known as the Stepleton enclosure), which is of a similar date, 3650–3370 cal. BC (Mercer and Healy 2008) (see Supplementary Information, Table S1). Hazleton North is a Cotswold-Severn chambered tomb near Cheltenham, Gloucestershire, dating to c.3800–3620 cal. BC (Saville et al. 1987; Meadows et al. 2007). The five individuals sampled for dental calculus here come from a number of different areas within the trapezoidal lateral long cairn (see Supplementary Information, Table S2). Finally, Banbury Lane is a Middle Neolithic triple-ditched monument, located in Northampton (Holmes 2012). The individuals sampled within this study all derive from a large pit of disarticulated human remains dug into the entrance to the innermost ditch of the monument, which has been dated to c.3360–3090 cal. BC (Holmes 2012) (see Supplementary Information, Table S3). All individuals analysed from the three sites were adults, apart from one individual from Hambledon Hill identified as infant/juvenile (HH74 HB3/ HH3) by McKinley (2008) (Table 1 and Tables S1–S3). Furthermore, the majority of the skeletal material sampled was disarticulated or fragmentary, although one articulated skeleton from both Hambledon Hill and Hazleton North respectively was analysed (Tables S1 and S2). Due to this, few of the individuals included within this study could be osteologically sexed. Although not all of the 10 individuals have been directly AMS dated, they all come from secure and well-dated archaeological contexts. Additional information on each of the individuals studied can be found in the Supplementary Information (Tables S1–S3).
Dental calculus was removed from archaeological specimens using a sterile dental scaler and stored in sterile 2.0-mL tubes. Peptides were extracted from decalcified dental calculus using a filter-aided sample preparation (FASP) protocol modified for degraded samples (Cappellini et al. 2014) according to previously published protocols (Warinner et al. 2014a, b). The extracted peptides were then analysed using shotgun protein tandem mass spectrometry (MS/MS) at the Central Proteomics Facility, Target Discovery Institute, Oxford on a Q-Exactive tandem mass spectrometer (see Supplementary Information).
Data analysis
Raw tandem mass spectra were converted to searchable Mascot generic format using Proteowizard version 3.0.7518 using the 200 most intense peaks in each mass spectrum. MS/MS ion database searching was performed on Mascot (Matrix Science™, version 2.5.1 (Perkins et al. 1999)), against all available sequences in UniProt and the Human Oral Microbiome Database (HOMD) (as in Warinner et al. 2014a). Searches were run twice, first as tryptic with one missed cleavage and a peptide tolerance of 10 ppm (Table S6), and then as semi-tryptic allowing up to two missed cleavages and a peptide tolerance of 10 ppm (Table 2). For both searches, MS/MS ion tolerance was set to 0.07 Da. Post-translational modifications were set as carbamidomethylation (fixed modification), and acetyl (protein N-term), deamidated (NQ), glutamine to pyroglutamate, methionine oxidation and hydroxylation of proline (variable modifications). Searches were performed against a decoy database to estimate protein false discovery rates. Mascot search results were filtered to a false discovery rate of less than 5% and an ion score of > 25. Searches for dietary proteins were conducted based on the strategy reported in Hendy et al. (2018b). First, the protein results were filtered to include only those proteins that were represented by at least two peptides, before assigning Mascot protein family identifications to broad taxonomic classifications of mammalian, plant or microbial sources. A conservative approach was taken, and any protein identified in our blank controls or injection blanks was assigned to the ‘contaminant’ category; these were principally derived from trypsin, human collagens and human keratins. For all putative dietary derived proteins, BLASTp was used to verify all peptide matches, and taxonomic assignment is reported based on the consensus peptide assignments for each individual (see Supplementary Information).
To validate whether the identified dietary proteins were derived from ancient endogenous proteins as opposed to modern contaminants, levels of deamidation were assessed following the method proposed in Mackie et al. (2018). The raw MS/MS files were run on MaxQuant (Cox and Mann 2008) version 1.6.2.6a against bovid β-lactoglobulin, as well as against a custom database including bovid and horse milk proteins, human salivary proteins, and common laboratory contaminant proteins (Supplementary Information, SI File 1). Bulk deamidation rates were assessed in the identified peptides using intensity-based calculations—for example, if a protein is 2% deamidated, this means that, for its summed intensities, only 2% of the total sum relates to peptides with deamidated residues. All parameters were default, apart from using a semi-tryptic search strategy with dependent peptides enabled, and the score cut-off for modified and unmodified peptides was set to 60. Carbamidomethyl (C) was added as a fixed modification, whilst variable modifications included oxidation (M), acetyl (protein N-term), deamidation (NQ), Gln->pyro-Glu, Glu->pyro-Glu and hydroxyproline.
Contamination controls
It is important to monitor and test for contamination because bovine proteins are used in some proteomics laboratories as instrument standards (e.g. bovine fetuin), among other purposes. Precautionary measures were taken to reduce the likelihood of contamination from modern protein sources and ensure that the reported proteins were endogenous to the dental calculus (Hendy et al. 2018b), including: (1) the use of a dedicated ancient protein laboratory where modern proteins are not extracted or analysed; (2) the inclusion of blank controls analysed in parallel with the experimental samples to monitor for contamination during protein extraction; (3) the inclusion of injection blanks between each dental calculus sample to monitor for protein carryover during LC-MS/MS analysis; and (4) the analysis of bulk deamidation rates in observed dietary peptides using intensity-based calculations. No milk or other dietary proteins were detected in extraction blanks or injection blanks in this study.
Results and discussion
Proteins were successfully recovered from all 10 of the dental calculus samples analysed, with total protein identifications ranging from 15 to 128 (following quality filtering) (Table S5). The identified proteins were assigned primarily to the human proteome and to microbial taxa commonly found within the oral microbiome, as well as to common contaminants identified in previous dental calculus proteomic analyses (principally trypsin, human keratins and human collagens) (Hendy et al. 2018a; Mackie et al. 2017; Warinner et al. 2014b) (Fig. S1). The only dietary protein identified within the sample set was BLG. The initial Mascot search strategy only considered peptides which conformed to tryptic cleavage patterns, identifying 34 BLG peptides in 6 individuals (Table S6). These identifications were augmented with spectral data searched using semi-tryptic modifications (Table 2), which identified 29 additional BLG peptides (i.e., not detected in the tryptic dataset) and an additional BLG positive individual (HH3). Using this combined dataset, over half of the dental calculus samples (n = 7) displayed evidence for BLG peptides (Table 2). BLG peptides were identified in all three assemblages (Hambledon Hill (n = 2), Hazleton North (n = 4), Banbury Lane (n = 1)), although with one individual from both Hambledon Hill and Hazelton North represented by only a single BLG peptide. No BLG peptides were identified within blank controls or injection blanks. Using the combined search strategy, 63 spectra (comprising 14 distinct peptides) were assigned to BLG. For each of the samples which tested positive for BLG, a consensus BLG sequence could be assigned to ruminants of the Pecora infraorder of Artiodactyla, with all seven samples containing Bovidae-specific peptides (see Supplementary Information).
Levels of deamidation were examined using MaxQuant to ensure that the recovered BLG peptides and human oral proteins conformed with damage expected from ancient proteins, as opposed to modern contaminants (Mackie et al. 2018). Whilst asparagine (N) tends to deamidate into aspartic acid (D) relatively rapidly (van Duin and Collins 1998; Robinson and Robinson 2001, 2004), deamidation of glutamine (Q) into glutamic acid (E) is markedly slower and therefore can be used to assess protein degradation in archaeological contexts (van Doorn et al. 2012; Wilson et al. 2012). As expected, contaminant proteins (e.g., human keratins, trypsin, etc.) displayed, on average, low-levels of deamidation, with 90.9% undamaged asparagines and 98.5% undamaged glutamines (Fig. S2). The subset of analysed human proteins displayed variable, but on average, more advanced deamidation, with only 17.7% undamaged asparagines and 33.4% undamaged glutamines across the samples (Fig. S2). The BLG peptides displayed the most advanced levels of deamidation. In total, five individuals displayed BLG peptides with at least one deaminating residue: two individuals (HH610 and HN7387) displayed BLG peptides with both Ns and Qs and three individuals (HN11456, BL309.12, HN4786) displayed peptides with either Ns or Qs. For further two individuals (HH3, HN7656), the detected BLG peptides contained neither Ns nor Qs. In the five individuals where at least one deamidating BLG peptide was observed, the deamidation levels were extremely advanced. On average, only 2% of the asparagines and 5% of the glutamines were undamaged across the samples (Fig. 2). These levels of degradation are more advanced than those reported in previous analyses of BLG from ancient dental calculus proteomes (Mays et al. 2018; Mackie et al. 2017), a result consistent with the greater antiquity of these Neolithic individuals compared to previous studies (e.g., Middle Bronze Age, Roman and Medieval individuals).
Of the two individuals from Hambledon Hill in which BLG peptides were detected here, both indicated BLG deriving from Bovidae (HH3; HH610), a family of ruminants which includes cattle, buffalo, bison, antelopes, sheep, goats and gazelles (Gentry 1992). Therefore, the BLG within the calculus of two individuals at Hambledon Hill may have derived from cattle, sheep or goat milk. However, one tryptic peptide in individual HH610 also suggested the specific presence of goat (Capra sp.) milk. These results are supported by the faunal assemblage from the site, with domesticated cattle (Bos taurus) being the most dominant species present, followed by smaller numbers of ovicaprids (Ovis aries and Capra hircus) (Legge 2008). The age and sex structure of the cattle remains at Hambledon Hill have previously been suggested to be indicative of a dairy herd (Whittle 1992, p. 221; Copley et al. 2003), although others have suggested that the restricted age distribution of the cattle means that the animals were unlikely to have derived from a resident herd (Legge 2008). The high number of cattle skulls has also been interpreted as being representative of an association between the deposition of human and faunal remains (Thomas 1999, p. 28), and parallels between the treatment of human and cattle remains has been suggested to represent ‘respectful consumption’ of beef (Jones 2007, p. 164). The large-scale deposition of cattle remains, however, alongside large amounts of burnt wheat, barley and hazelnuts, has prompted suggestions of feasting at the site (Jones and Legge 2008; Mercer 2008). Additionally, organic residue analysis of pottery from Hambledon Hill has indicated the presence of both porcine and ruminant fats, and ruminant adipose and dairy fats (Copley et al. 2003, 2005a, b, 2008). The presence of dairy fats in > 25% of potsherds analysed has been suggested to indicate that ‘dairying was a very important element of animal husbandry at Hambledon Hill’ (Copley et al. 2008, p. 535).
Of the four individuals from Hazleton North in which BLG peptides were detected, all were also specific to Bovidae, but with peptides distinct from Capra, suggesting that individuals from Hazleton North were likely exploiting milk from cattle and/or sheep, but not goat. Stable isotopic analysis of δ13C and δ15N on human remains from the site has revealed a diet high in animal protein, supplemented by C3 plants (Hedges et al. 2008). Although Hazleton North does not possess a faunal assemblage akin to that at Hambledon Hill, the remains of domesticated cattle were nonetheless recovered within the chambered areas of the tomb at the site (Levitan 1990), and therefore, is consistent with the utilisation of cattle milk by the individuals at Hazleton North.
One individual from Banbury Lane (BL309.12) was also found to have BLG specific to Bovidae, but not Capra, using the tryptic search strategy. Unlike the other two sites, Banbury Lane has very little faunal material or pottery associated with either the monument itself or the human remains (Holmes 2012).
The combined results obtained here from British sites indicate that Neolithic populations were utilising milk and/or dairy products from either cattle, goat or sheep (or potentially combinations of these). This evidence is consistent with the recent discovery of bovine milk consumption in the Middle Neolithic individual recovered from Stonehenge (Mays et al. 2018). The results obtained here also show varying numbers of BLG spectra detected across all individuals, as has been observed in previous studies (Warinner et al. 2014a; Mays et al. 2018). For example, two individuals analysed here (HH3 and HN7656) exhibited only one BLG-specific spectrum each, whereas in contrast, individual HH610 from Hambledon Hill had a total of 36 spectra matching BLG peptides (13 unique peptides). The reasons as to why some individuals have a significantly higher number of BLG spectra are still unclear but is something which certainly warrants further future study. Additional study of the abundance of BLG spectra within different dental calculus samples may reveal if this is linked to the amount of dairy consumed by an individual, or instead if it is the result of preservational biases or environments, or the timings and nature of calculus formation. In order to accurately assess this, however, a much broader scale study, with the likely inclusion of modern dental calculus samples from individuals with known diets, would be needed. Further study should also reveal if individuals with no evidence of BLG peptides within their calculus (as observed in this study, and within other proteomic studies (Warinner et al. 2014a; Hendy et al. 2018a)) were truly not consuming dairy products, or if the absence of milk proteins is instead the result of differential calculus formation processes or taphonomic effects.
It is important to note, however, that due to recent discoveries of LP prevalence in the European Neolithic (Burger et al. 2007; Gamba et al. 2014; Witas et al. 2015; Olalde et al. 2018; Brace et al. 2019), it is unlikely that the British Neolithic individuals studied here would have carried the genetic mutation associated with lactase persistence. Human remains from neither Hambledon Hill nor Hazleton North have been subjected to ancient DNA analysis to date, but none of the individuals (n = 3) genetically analysed thus far from Banbury Lane have a derived lactase persistence allele (Brace et al. 2019). The presence of BLG within the dental calculus however indicates the consumption of dairy products, as supported by other archaeological evidence for dairying from the period, as discussed above. It is important to note however that BLG may reflect either the regular consumption of small quantities of raw milk—based on the observation that individuals without LP can tolerate up to 240 mL of milk per day with negligible symptoms (Suarez et al. 1995; Swallow 2003)—or alternatively, the consumption of processed milk products with reduced lactose content. Distinguishing between these two consumption scenarios is not possible based on the current data and analytical technique. Nevertheless, as organic residue analysis has revealed that the processing of dairy in pottery vessels was widespread from the Early Neolithic onwards in Britain (Copley et al. 2005b), the likely absence of LP, but presence of BLG, is therefore more consistent with the hypothesis that Neolithic populations were processing milk to remove the lactose, but in such a way that at least some BLG was retained.
Lactose can be removed from or decreased in milk products through a range of processing methods. For example, cheese contains little or no lactose, as it is removed during processing with the whey fraction of the milk. Indeed, 98% of lactose is removed in the whey during most cheese production (Izco et al. 2002). The production of cheese within prehistory has however been suggested to have been beneficial for past populations not only due to the reduced lactose content, thereby making it more readily digestible and suitable for non-LP individuals, but also because it allows for the preservation of milk products in a transportable form (Salque et al. 2013). Lactose content is also known to be much decreased in fermented milk products, such as yoghurt, kefir and buttermilk (Alm 1982; O’Brien 1999). As such, fermented milk products have previously been suggested to be suitable for consumption by lactose intolerant individuals (Alm 1982). Although yoghurt does contain small amounts of lactose, it is believed that this lactose is more easily digested than that found within whole milk, due to hydrolysis and autodigestion of lactose by the yoghurt bacteria—thus improving its absorption and creating a ‘lactase activity’ in the gastrointestinal tract (Kolars et al. 1984; Savaiano 2014).
BLG is known to be present in processed milk products, but generally in significantly lower quantities than whole milk, and decreases the more the whole milk is processed. BLG is known to be absent or at very low levels in hard cheeses, for example, due to the removal of the whey fraction of the milk during processing. BLG is however present in yoghurts and other fermented milk products, and it has been noted that it is not markedly subject to hydrolysis or proteolysis during the fermentation process (Bertrand-Harb et al. 2003; Tzvetkova et al. 2007). The heating or fermentation of milk will decrease the levels of BLG within it, but to varying degrees dependent upon the type and intensity of processing utilised (Czerwenka et al. 2007; Bu et al. 2013).
The idea of Neolithic milk processing to reduce lactose content has been proposed previously (e.g., in the form of cheese making (Salque et al. 2013)) and is plausible given the recent genetic evidence for the absence of LP in Neolithic populations. Of the three sites analysed here, only the pottery from Hambledon Hill has previously had organic residue analysis undertaken upon it, indicating the exploitation of dairy fats (Copley et al. 2003). Overall, the evidence for dairy consumption (as evidenced through BLG peptides within calculus detected here and lipid residue analyses on pottery), combined with genetic support for the absence of LP, suggests that either British Neolithic populations were processing raw milk in an attempt to remove or reduce the lactose content, or alternatively consuming relatively small quantities of raw milk. As such, these observations open a range of exciting new research avenues exploring how Neolithic populations may have been consuming and processing raw milk, the required technologies needed to do this, the kinds of products which may have been created, the potential variability which may have existed within these processes in the past, and notions of cuisine. It is conceivable that past populations chose to utilise the milk of different animals purposively, and processed this in different ways, due to cultural reasons or even taste. We can consider the modern regional variability which exists in cheese production in the UK—for example, varying in terms of the types of milk used, the processing methods, if the finished product is a soft or hard cheese, how long the cheese is left to mature for, and often having Protected Designation of Origin or Protected Geographical Indication status (British Cheese Board 2018)—as an indication that similar regional differences may have also existed in the prehistoric past in dairy processing and production. The fact that milk lipids are frequently found in British Neolithic pottery, often with thermally modified compounds, such as long-chain ketones, also suggests that raw milk was processed by heating.
Conclusions
Proteomic analysis of dental calculus obtained from 10 British Neolithic skeletons has revealed the presence of the milk protein β-lactoglobulin (BLG) in seven of the analysed skeletons, providing the earliest direct evidence for the consumption of dairy products in Britain and the earliest identification of BLG in human dental calculus to date. Analysis of deamidation patterns suggests that the BLG is indeed a degraded endogenous protein and not the result of laboratory contamination. Although only a small sample size, the presence of Bovidae-specific peptides within the calculus samples provides new insight into prehistoric patterns of milk consumption and can also be tied into larger discussions regarding dietary variables driving natural selection in humans and gene-culture co-evolution. Whilst previous studies suggest that early Neolithic individuals lacked LP (Brace et al. 2019), future genomic analyses targeting individuals with direct evidence for BLG may provide greater insight into selection for raw milk consumption. Additionally, the indication that the individuals studied here are unlikely to have had LP, but were consuming and utilising dairy products, indicates a range of exciting new research avenues exploring how Neolithic populations may have been consuming and processing raw milk and the potential variability which may have existed within these processes in the past.
The lack of other dietary proteins detected within the Neolithic samples analysed here echoes the results of other metaproteomic studies of dental calculus, suggesting dietary proteins are of low abundance in dental calculus and that BLG appears to have enhanced resistance to degradation within calculus, leading to its long-term survival (Mackie et al. 2017; Mays et al. 2018; Hendy et al. 2018a; Jersie-Christensen et al. 2018). This research however clearly highlights the utility of metaproteomic approaches in exploring prehistoric dairying. To our knowledge, this study represents the earliest direct evidence of milk consumption globally and also the earliest identification of the milk whey protein β-lactoglobulin (BLG) to date.
The presence of Bovidae-specific peptides within the calculus samples analysed here opens up the possibility that British Neolithic populations may have been exploiting multiple species for dairy products including cows, sheep and goat, although zooarchaeological evidence from the sites is most consistent with cattle milk. Future studies targeting a greater diversity of British Neolithic sites may indicate distinctions between the utilisation of different ruminant species for milk exploitation, and whether this varied on a population level (i.e., between sites), or was more closely aligned to social constructs. Despite the small sample size, the detection of BLG in seven of the ten tested individuals may provide a tentative glimpse into the prevalence of dairy consumption. The detection of BLG in the majority of individuals across three sites may suggest that milk consumption was not confined to a small number of individuals within Neolithic society but was more likely a widespread dietary practice—a hypothesis that certainly warrants further investigation with a larger number of individuals, archaeological sites and burial contexts. With further future work, the idea that we may be able to distinguish if certain members of a society consumed differential amounts of dairy products or dairy from different animals—for example, along the lines of sex, gender, age or social standing—is truly fascinating and would provide a unique new insight into British Neolithic culture and social structure. Furthermore, if milk is such an accessible foodstuff in the Neolithic, what was the underlying driver of the observed selection pressure for lactase persistence? Increased genetic testing of archaeological remains in the future, including the analysis of individuals with evidence of BLG, may provide more insight into milk consumption and processing, and the gene-culture co-evolution of LP.
Data availability
Mass spectrometry data are available via the ProteomeXchange Consortium under the accession PXD012893.
References
Albarella U, Serjeantson D (2002) A passion for pork: butchery and cooking at the British Neolithic site of Durrington Walls. In: Miracle PT, Milner N (eds) Consuming passions and patterns of consumption. MacDonald Institute, Cambridge, pp 33–49
Allentoft ME, Sikora M, Sjögren K-G, Rasmussen S, Rasmussen M, Stenderup J, Damgaard PB, Schroeder H, Ahlström T, Vinner L, Malaspinas AS, Margaryan A, Higham T, Chivall D, Lynnerup N, Harvig L, Baron J, Casa PD, Dąbrowski P, Duffy PR, Ebel AV, Epimakhov A, Frei K, Furmanek M, Gralak T, Gromov A, Gronkiewicz S, Grupe G, Hajdu T, Jarysz R, Khartanovich V, Khokhlov A, Kiss V, Kolář J, Kriiska A, Lasak I, Longhi C, McGlynn G, Merkevicius A, Merkyte I, Metspalu M, Mkrtchyan R, Moiseyev V, Paja L, Pálfi G, Pokutta D, Pospieszny Ł, Price TD, Saag L, Sablin M, Shishlina N, Smrčka V, Soenov VI, Szeverényi V, Tóth G, Trifanova SV, Varul L, Vicze M, Yepiskoposyan L, Zhitenev V, Orlando L, Sicheritz-Pontén T, Brunak S, Nielsen R, Kristiansen K, Willerslev E (2015) Population genomics of Bronze Age Eurasia. Nature 522:167–172
Alm L (1982) Effect of fermentation on lactose, glucose, and galactose content in milk and suitability of fermented milk products for lactose intolerant individuals. J Dairy Sci 65:346–352
Beja-Pereira A, Luikart G, England PR, Bradley DG, Jann OC, Bertorelle G, Chamberlain AT, Nunes TP, Metodiev S, Ferrand N, Erhardt G (2003) Gene-culture coevolution between cattle milk protein genes and human lactase genes. Nat Genet 35:311–313
Bersaglieri T, Sabeti PC, Patterson N, Vanderploeg T, Schaffner SF, Drake JA, Rhodes M, Reich DE, Hirschhorn JN (2004) Genetic signatures of strong recent positive selection at the lactase gene. Am J Hum Genet 74:1111–1120
Bertrand-Harb C, Ivanova IV, Dalgalarrondo M, Haertllé T (2003) Evolution of β-lactoglobulin and α-lactalbumin content during yoghurt fermentation. Int Dairy J 13:39–45
Bogaard A, Fraser R, Heaton THE, Wallace M, Vaiglova P, Charles M, Jones G, Evershed RP, Styring AK, Andersen NH, Arbogast RM, Bartosiewicz L, Gardeisen A, Kanstrup M, Maier U, Marinova E, Ninov L, Schafer M, Stephan E (2013) Crop manuring and intensive land management by Europe’s first farmers. Proc Natl Acad Sci U S A 110:12589–12594
Bovenhuis H, Visker MHPW, Lundén A (2013) Selection for milk fat and milk protein composition. Adv Anim Biosci 4:612–617
Brace S, Diekmann Y, Booth TJ, van Dorp L, Faltyskova Z, Rohland N, Mallick S, Olalde I, Ferry M, Michel M, Oppenheimer J, Broomandkhoshbacht N, Stewardson K, Martiniano R, Walsh S, Kayser M, Charlton S, Hellenthal G, Armit I, Schulting R, Craig OE, Sheridan A, Parker Pearson M, Stringer C, Reich D, Thomas MG, Barnes I (2019) Ancient genomes indicate population replacement in Early Neolithic Britain. Nat Ecol Evol 3:765–771. https://doi.org/10.1038/s41559-019-0871-9
British Cheese Board (2018) British protected name cheeses. In: British Cheese Board. http://www.britishcheese.com/othercheese/british_protected_name_cheeses-89. Accessed 17 Jul 2018
Brown A (2007) Dating the onset of cereal cultivation in Britain and Ireland: the evidence from charred cereal grains. Antiquity 81:1042–1052
Bu G, Luo Y, Chen F, Liu K, Zhu T (2013) Milk processing as a tool to reduce cow’s milk allergenicity: a mini-review. Dairy Sci Technol 93:211–223
Burger J, Kirchner M, Bramanti B, Haak W, Thomas MG (2007) Absence of the lactase-persistence-associated allele in early Neolithic Europeans. Proc Natl Acad Sci U S A 104:3736–3741
Caffell A, Holst M (2012) The Human Bone. In: Holmes M (ed) Archaeological excavation on land off Banbury Lane. Pineham, Northampton. Assessment Report and Updated Project Design. Northamptonshire Archaeology Report 12/133
Cappellini E, Gentry A, Palkopoulou E et al (2014) Resolution of the type material of the Asian elephant, Elephas maximus Linnaeus, 1758 (Proboscidea, Elephantidae). Zool J Linnean Soc 170:222–232
Cavalli-Sforza LL (1973) Analytic review: some current problems of human population genetics. Am J Hum Genet 25:82–104
Coelho M, Luiselli D, Bertorelle G, Lopes AI, Seixas S, Destro-Bisol G, Rocha J (2005) Microsatellite variation and evolution of human lactase persistence. Hum Genet 117:329–339
Copley MS, Berstan R, Dudd SN, Docherty G, Mukherjee AJ, Straker V, Payne S, Evershed RP (2003) Direct chemical evidence for widespread dairying in prehistoric Britain. Proc Natl Acad Sci U S A 100:1524–1529
Copley MS, Berstan R, Mukherjee AJ et al (2005b) Dairying in antiquity. III Evidence from absorbed lipid residues dating to the British Neolithic. J Archaeol Sci 32:523–546
Copley MS, Berstan R, Dudd SN, Aillaud S, Mukherjee AJ, Straker V, Payne S, Evershed RP (2005a) Processing of milk products in pottery vessels through British prehistory. Antiquity 79:895–908
Copley M, Berstan R, Stott A, Evershed R (2008) Organic residue analysis of pottery vessels: determination of vessel use and radiocarbon dates. In: Mercer R, Healy F (eds) Hambledon Hill, Dorset. England. Excavation and survey of a Neolithic monument complex and its surrounding landscape. English Heritage, Swindon, pp 527–535
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26:1367–1372
Craig OE, Chapman J, Heron C, Willis LH, Bartosiewicz L, Taylor G, Whittle A, Collins M (2005) Did the first farmers of central and eastern Europe produce dairy foods? Antiquity 79:882–894
Craig OE, Shillito L-M, Albarella U, Viner-Daniels S, Chan B, Cleal R, Ixer R, Jay M, Marshall P, Simmons E, Wright E, Pearson MP (2015) Feeding Stonehenge: cuisine and consumption at the Late Neolithic site of Durrington Walls. Antiquity 89:1096–1109
Cramp LJE, Evershed RP, Lavento M, Halinen P, Mannermaa K, Oinonen M, Kettunen J, Perola M, Onkamo P, Heyd V (2014a) Neolithic dairy farming at the extreme of agriculture in northern Europe. Proc Biol Sci 281:20140819
Cramp LJE, Jones J, Sheridan A, Smyth J, Whelton H, Mulville J, Sharples N, Evershed RP (2014b) Immediate replacement of fishing with dairying by the earliest farmers of the Northeast Atlantic archipelagos. Proc Biol Sci 281:20132372
Crittenden RG, Bennett LE (2005) Cow’s milk allergy: a complex disorder. J Am Coll Nutr 24:582S–591S
Czerwenka C, Maier I, Potocnik N, Pittner F, Lindner W (2007) Absolute quantitation of β-lactoglobulin by protein liquid chromatography−mass spectrometry and its application to different milk products. Anal Chem 79:5165–5172
Gamba C, Jones ER, Teasdale MD, McLaughlin RL, Gonzalez-Fortes G, Mattiangeli V, Domboróczki L, Kővári I, Pap I, Anders A, Whittle A, Dani J, Raczky P, Higham TFG, Hofreiter M, Bradley DG, Pinhasi R (2014) Genome flux and stasis in a five millennium transect of European prehistory. Nat Commun 5:5257
Gentry AW (1992) The subfamilies and tribes of the family Bovidae. Mammal Rev 22:1–32
Gerbault P, Moret C, Currat M, Sanchez-Mazas A (2009) Impact of selection and demography on the diffusion of lactase persistence. PLoS One 4:e6369
Gerbault P, Liebert A, Itan Y, Powell A, Currat M, Burger J, Swallow DM, Thomas MG (2011) Evolution of lactase persistence: an example of human niche construction. Philos Trans R Soc Lond Ser B Biol Sci 366:863–877
Greenfield HJ, Arnold ER (2015) “Go(a)t milk?” New perspectives on the zooarchaeological evidence for the earliest intensification of dairying in south eastern Europe. World Archaeol 0:1–27
Hedges REM, Stevens RE, Pearson JA (2006) Carbon and nitrogen stable isotope compositions of animal and human bone. In: Benson D, Whittle A (eds) Building memories: the Neolithic Cotswold long barrow at Ascott-under-Wychwood. Oxfordshire. Oxbow Books, Oxford, pp 255–262
Hedges R, Saville A, O’Connell T (2008) Characterizing the diet of individuals at the Neolithic chambered tomb of Hazleton North, Gloucestershire, England, using stable isotopic analysis. Archaeometry 50:114–128
Hendy J, Warinner C, Bouwman A, Collins MJ, Fiddyment S, Fischer R, Hagan R, Hofman CA, Holst M, Chaves E, Klaus L, Larson G, Mackie M, McGrath K, Mundorff AZ, Radini A, Rao H, Trachsel C, Velsko IM, Speller CF (2018a) Proteomic evidence of dietary sources in ancient dental calculus. Proc R Soc B 285:20180977
Hendy J, Welker F, Demarchi B, Speller C, Warinner C, Collins MJ (2018b) A guide to ancient protein studies. Nat Ecol Evol 2:791–799
Holmes M (ed) (2012) Archaeological excavation on land off Banbury Lane, Pineham, Northampton Assessment Report and Updated Project Design. Northamptonshire Archaeology Report 12/133
Itan Y, Powell A, Beaumont MA, Burger J, Thomas MG (2009) The origins of lactase persistence in Europe. PLoS Comput Biol 5:e1000491
Izco JM, Tormo M, Jiménez-Flores R (2002) Rapid simultaneous determination of organic acids, free amino acids, and lactose in cheese by capillary electrophoresis. J Dairy Sci 85:2122–2129
Jersie-Christensen RR, Lanigan LT, Lyon D, Mackie M, Belstrøm D, Kelstrup CD, Fotakis AK, Willerslev E, Lynnerup N, Jensen LJ, Cappellini E, Olsen JV (2018) Quantitative metaproteomics of medieval dental calculus reveals individual oral health status. Nat Com 9:4744
Jones M (2007) Feast: Why Humans Share Food. Oxford University Press, Oxford
Jones G, Legge AJ (2008) Evaluating the role of cereal cultivation in the Neolithic: charred plant remains from Hambledon Hill. In: Mercer R, Healy F (eds) Hambledon Hill, Dorset, England. Excavation and survey of a Neolithic monument complex and its surrounding landscape. English Heritage, Swindon, pp 469–476
Jones BL, Oljira T, Liebert A, Zmarz P, Montalva N, Tarekeyn A, Ekong R, Thomas MG, Bekele E, Bradman N, Swallow DM (2015) Diversity of lactase persistence in African milk drinkers. Hum Genet 134:917–925
Kahle D, Wickham H (2013) ggmap: spatial visualization with ggplot2. The R Journal 5:144–161
Kolars JC, Levitt MD, Aouji M, Savaiano DA (1984) Yogurt--an autodigesting source of lactose. N Engl J Med 310:1–3
Kontopidis G, Holt C, Sawyer L (2004) Invited review: beta-lactoglobulin: binding properties, structure, and function. J Dairy Sci 87:785–796
Legge T (2005) Milk use in prehistory: the osteological evidence. In: Mulville J, Outram AK (eds) The Zooarchaeology of Fats, Oils, Milk and Dairying. Oxbow, Oxford, pp 8–13
Legge AJ (2008) Livestock and Neolithic society at Hambledon Hill. In: Mercer R, Healy F (eds) Hambledon Hill, Dorset. England. Excavation and survey of a Neolithic monument complex and its surrounding landscape. English Heritage, Swindon, pp 536–586
Leonardi M, Gerbault P, Thomas MG, Burger J (2012) The evolution of lactase persistence in Europe. A synthesis of archaeological and genetic evidence. Int Dairy J 22:88–97
Levitan B (1990) The non-human vertebrate remains. In: Saville A (ed) Hazleton North, Gloucestershire, 1979–1982: The excavation of a Neolithic long cairn of the Cotswold-Severn group. Historic Buildings & Monuments Commission for England, London, pp 199–213
Liebert A, López S, Jones BL, Montalva N, Gerbault P, Lau W, Thomas MG, Bradman N, Maniatis N, Swallow DM (2017) World-wide distributions of lactase persistence alleles and the complex effects of recombination and selection. Hum Genet 136:1445–1453. https://doi.org/10.1007/s00439-017-1847-y
Luinge HJ, Hop E, Lutz ETG, van Hemert JA, de Jong EAM (1993) Determination of the fat, protein and lactose content of milk using Fourier transform infrared spectrometry. Anal Chim Acta 284:419–433
Mackie M, Hendy J, Lowe AD et al (2017) Preservation of the metaproteome: variability of protein preservation in ancient dental calculus. STAR: Sci Technol Archaeol Res 3:74–86
Mackie M, Rüther P, Samodova D, di Gianvincenzo F, Granzotto C, Lyon D, Peggie DA, Howard H, Harrison L, Jensen LJ, Olsen JV, Cappellini E (2018) Palaeoproteomic profiling of conservation layers on a 14th century Italian wall painting. Angew Chem 57:7369–7374
Mathieson I, Lazaridis I, Rohland N, Mallick S, Patterson N, Roodenberg SA, Harney E, Stewardson K, Fernandes D, Novak M, Sirak K, Gamba C, Jones ER, Llamas B, Dryomov S, Pickrell J, Arsuaga JL, de Castro JMB, Carbonell E, Gerritsen F, Khokhlov A, Kuznetsov P, Lozano M, Meller H, Mochalov O, Moiseyev V, Guerra MAR, Roodenberg J, Vergès JM, Krause J, Cooper A, Alt KW, Brown D, Anthony D, Lalueza-Fox C, Haak W, Pinhasi R, Reich D (2015) Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528:499–503
Mays S, Roberts D, Marshall P, Pike AWG, van Heekeren V, Bronk Ramsey C, Dunbar E, Reimer P, Linscott B, Radini A, Lowe A, Dowle A, Speller C, Vallender J, Bedford J (2018) Lives before and after Stonehenge: an osteobiographical study of four prehistoric burials recently excavated from the Stonehenge World Heritage Site. J Archaeol Sci Rep 20:692–710
McKinley JI (2008) The human remains. In: Mercer R, Healey F (eds) Hambledon Hill, Dorset. England. Excavation and survey of a Neolithic monument complex and its surrounding landscape. English Heritage, Swindon, pp 477–521
Meadows J, Barclay A, Bayliss A (2007) A short passage of time: the dating of the Hazleton long cairn revisited. Camb Archaeol J 17:45–64
Mercer R (2008) The Neolithic uses of the complex. In: Mercer R, Healy F (eds) Hambledon Hill, Dorset, England. Excavation and survey of a Neolithic monument complex and its surrounding landscape. English Heritage, Swindon, pp 753–760
Mercer RJ, Healy F (2008) Hambledon Hill, Dorset. England, English Heritage
Milner N, Craig OE, Bailey GN, Pedersen K, Andersen SH (2004) Something fishy in the Neolithic? A re-evaluation of stable isotope analysis of Mesolithic and Neolithic coastal populations. Antiquity 78:9–22
Monaci L, Tregoat V, van Hengel AJ, Anklam E (2006) Milk allergens, their characteristics and their detection in food: a review. Eur Food Res Technol 223:149–179
Mulville J, Grigson C (2007) The Animal Nones. In: Benson D, Whittle A (eds) Building Memories: The Neolithic Cotswold Long Barrow at Ascott-Under-Wychwood. Oxfordshire. Oxbow Books, Oxford, pp 237–254
Mulville J, Bond J, Craig O (2005) The white stuff, milking in the Outer Scottish Isles. In: Mulville J, Outram AK (eds) The Zooarchaeology of Fats, Oils, Milk and Dairying. Oxbow, Oxford, pp 167–182
Myles S, Bouzekri N, Haverfield E, Cherkaoui M, Dugoujon JM, Ward R (2005) Genetic evidence in support of a shared Eurasian-North African dairying origin. Hum Genet 117:34–42
Nielsen R, Hellmann I, Hubisz M, Bustamante C, Clark AG (2007) Recent and ongoing selection in the human genome. Nat Rev Genet 8:857–868
O’Brien J (1999) Sugar profiles of cultured dairy products in the UK. J Hum Nutr Diet 12:245–250
Olalde I, Brace S, Allentoft ME, Armit I, Kristiansen K, Booth T, Rohland N, Mallick S, Szécsényi-Nagy A, Mittnik A, Altena E, Lipson M, Lazaridis I, Harper TK, Patterson N, Broomandkhoshbacht N, Diekmann Y, Faltyskova Z, Fernandes D, Ferry M, Harney E, de Knijff P, Michel M, Oppenheimer J, Stewardson K, Barclay A, Alt KW, Liesau C, Ríos P, Blasco C, Miguel JV, García RM, Fernández AA, Bánffy E, Bernabò-Brea M, Billoin D, Bonsall C, Bonsall L, Allen T, Büster L, Carver S, Navarro LC, Craig OE, Cook GT, Cunliffe B, Denaire A, Dinwiddy KE, Dodwell N, Ernée M, Evans C, Kuchařík M, Farré JF, Fowler C, Gazenbeek M, Pena RG, Haber-Uriarte M, Haduch E, Hey G, Jowett N, Knowles T, Massy K, Pfrengle S, Lefranc P, Lemercier O, Lefebvre A, Martínez CH, Olmo VG, Ramírez AB, Maurandi JL, Majó T, McKinley JI, McSweeney K, Mende BG, Modi A, Kulcsár G, Kiss V, Czene A, Patay R, Endrődi A, Köhler K, Hajdu T, Szeniczey T, Dani J, Bernert Z, Hoole M, Cheronet O, Keating D, Velemínský P, Dobeš M, Candilio F, Brown F, Fernández RF, Herrero-Corral AM, Tusa S, Carnieri E, Lentini L, Valenti A, Zanini A, Waddington C, Delibes G, Guerra-Doce E, Neil B, Brittain M, Luke M, Mortimer R, Desideri J, Besse M, Brücken G, Furmanek M, Hałuszko A, Mackiewicz M, Rapiński A, Leach S, Soriano I, Lillios KT, Cardoso JL, Pearson MP, Włodarczak P, Price TD, Prieto P, Rey PJ, Risch R, Rojo Guerra MA, Schmitt A, Serralongue J, Silva AM, Smrčka V, Vergnaud L, Zilhão J, Caramelli D, Higham T, Thomas MG, Kennett DJ, Fokkens H, Heyd V, Sheridan A, Sjögren KG, Stockhammer PW, Krause J, Pinhasi R, Haak W, Barnes I, Lalueza-Fox C, Reich D (2018) The Beaker phenomenon and the genomic transformation of Northwest Europe. Nature. 555:190–196. https://doi.org/10.1038/nature25738
Perkins DN, Pappin DJ, Creasy DM, Cottrell JS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20:3551–3567
Restani P, Ballabio C, Di Lorenzo C et al (2009) Molecular aspects of milk allergens and their role in clinical events. Anal Bioanal Chem 395:47–56
Richards MP (2008) Hambledon Hill stable isotope values. In: Mercer R, Healy F (eds) Hambledon Hill, Dorset, England. Excavation and survey of a Neolithic monument complex and its surrounding landscape. English Heritage, Swindon, pp 522–527
Richards MP, Hedges R (1999) A Neolithic revolution? New evidence of diet in the British Neolithic. Antiquity 73:891–897
Richards MP, Schulting RJ, Hedges REM (2003) Archaeology: sharp shift in diet at onset of Neolithic. Nature 425:366
Robinson MA (2000) Further consideration of Neolithic charred cereals, fruit and nuts. In: Fairbairn AS (ed) Plants in Neolithic Britain and Beyond. Oxbow, Oxford, pp 85–90
Robinson NE, Robinson AB (2001) Deamidation of human proteins. Proc Natl Acad Sci U S A 98:12409–12413
Robinson NE, Robinson A (2004) Molecular clocks: deamidation of asparaginyl and glutaminyl residues in peptides and proteins. Althouse press, Cave Junction
Rogers J (1990) The human skeletal material. In: Saville A (ed) Hazleton North, Gloucestershire, 1979–1982: The excavation of a Neolithic long cairn of the Cotswold-Severn group. Historic Buildings & Monuments Commission for England, London, pp 182–198
Rowley-Conwy P (2004) How the West was lost: A reconsideration of agricultural origins in Britain, Ireland, and Southern Scandinavia. Curr Anthropol 45:S83–S113
Salque M, Radi G, Tagliacozzo A, Pino Uria B, Wolfram S, Hohle I, Stauble H, Whittle A, Hofmann D, Pechtl J, Schade-Lindig S, Eisenhauer U, Evershed RP (2012) New insights into the Early Neolithic economy and management of animals in Southern and Central Europe revealed using lipid residue analyses of pottery vessels. Anthropozoologica 47:45–61
Salque M, Bogucki PI, Pyzel J, Sobkowiak-Tabaka I, Grygiel R, Szmyt M, Evershed RP (2013) Earliest evidence for cheese making in the sixth millennium BC in northern Europe. Nature 493:522–525
Savaiano DA (2014) Lactose digestion from yogurt: mechanism and relevance. Am J Clin Nutr 99:1251S–1255S
Saville A, Gowlett JAJ, Hedges REM (1987) Radiocarbon dates from the chambered tomb at Hazleton (Glos.): a chronology for neolithic collective burial. Antiquity 61:108–119
Schulting R (2013) On the Northwestern Fringes: Earlier Neolithic subsistence in Britain and Ireland as seen through faunal remains and stable isotopes. In: Colledge S, Conolly J, Dobney K, Manning K, Shennan S (eds) The Origins and Spread of Domestic Animals in Southwest Asia and Europe. Left Coast Press Inc., Walnut Creek, pp 313–338
Ségurel L, Bon C (2017) On the evolution of lactase persistence in humans. Annu Rev Genomics Hum Genet 18:297–319
Smyth J, Evershed RP (2015) Milking the megafauna: Using organic residue analysis to understand early farming practice. Environ Archaeol 0:1–16
Stevens RE, Lightfoot E, Allen T, Hedges REM (2012) Palaeodiet at Eton College Rowing Course, Buckinghamshire: isotopic changes in human diet in the Neolithic, Bronze Age, Iron Age and Roman periods throughout the British Isles. Archaeol Anthropol Sci 4:167–184
Suarez FL, Savaiano DA, Levitt MD (1995) A comparison of symptoms after the consumption of milk or lactose-hydrolyzed milk by people with self-reported severe lactose intolerance. N Engl J Med 333:1–4
Swallow DM (2003) Genetics of lactase persistence and lactose intolerance. Annu Rev Genet 37:197–219
Thomas J (1999) Understanding the Neolithic. Routledge, Oxon
Thomas J (2003) Thoughts on the “Repacked” Neolithic Revolution. Antiquity 77:67–74
Tishkoff SA, Reed FA, Ranciaro A, Voight BF, Babbitt CC, Silverman JS, Powell K, Mortensen HM, Hirbo JB, Osman M, Ibrahim M, Omar SA, Lema G, Nyambo TB, Ghori J, Bumpstead S, Pritchard JK, Wray GA, Deloukas P (2007) Convergent adaptation of human lactase persistence in Africa and Europe. Nat Genet 39:31–40
Tzvetkova I, Dalgalarrondo M, Danova S et al (2007) Hydrolysis of major dairy proteins by lactic acid bacteria from Bulgarian yogurts. J Food Biochem 31:680–702
van Doorn NL, Wilson J, Hollund H, Soressi M, Collins MJ (2012) Site-specific deamidation of glutamine: a new marker of bone collagen deterioration. Rapid Commun Mass Spectrom 26:2319–2327
van Duin ACT, Collins MJ (1998) The effects of conformational constraints on aspartic acid racemization. Org Geochem 29:1227–1232
Vesa TH, Marteau P, Korpela R (2000) Lactose intolerance. J Am Coll Nutr 19:165S–175S
Vigne J-D (2008) Zooarchaeological aspects of the Neolithic diet transition in the Near East and Europe, and their putative relationships with the Neolithic demographic transition. In: Bocquet-Appel J-P, Bar-Yosef O (eds) The Neolithic Demographic Transition and its Consequences. Springer Verlag, New York, pp 179–205
Wal J-M (2004) Bovine milk allergenicity. Ann Allergy Asthma Immunol 93:S2–S11
Warinner C, Rodrigues JFM, Vyas R, Trachsel C, Shved N, Grossmann J, Radini A, Hancock Y, Tito RY, Fiddyment S, Speller C, Hendy J, Charlton S, Luder HU, Salazar-García DC, Eppler E, Seiler R, Hansen LH, Castruita JAS, Barkow-Oesterreicher S, Teoh KY, Kelstrup CD, Olsen JV, Nanni P, Kawai T, Willerslev E, von Mering C, Lewis CM, Collins MJ, Gilbert MTP, Rühli F, Cappellini E (2014b) Pathogens and host immunity in the ancient human oral cavity. Nat Genet 46:336–344
Warinner C, Hendy J, Speller C et al (2014a) Direct evidence of milk consumption from ancient human dental calculus. Sci Rep 4:7104
Whittle A (1992) Neolithic Europe: A Survey. Cambridge University Press, Cambridge
Wilson J, van Doorn NL, Collins MJ (2012) Assessing the extent of bone degradation using glutamine deamidation in collagen. Anal Chem 84:9041–9048
Witas HW, Płoszaj T, Jędrychowska-Dańska K, Witas PJ, Masłowska A, Jerszyńska B, Kozłowski T, Osipowicz G (2015) Hunting for the LCT-13910*T allele between the Middle Neolithic and the Middle Ages suggests its absence in dairying LBK people entering the Kuyavia region in the 8th millennium BP. PLoS One 10:e0122384
Acknowledgements
We would like to give many thanks to Northamptonshire Archaeology (MOLA)—particularly Andy Chapman, Adam Yates and Mark Holmes—and also to Malin Holst, Anwen Caffell and York Osteoarchaeology Ltd. for permissions, access and the sharing of information on the Banbury Lane assemblage. For permissions for the sampling of the Hambledon Hill material and access to collections, we would like to give many thanks to Dorset County Museum, Richard Breward and Jonathan Murden. Finally, for access to collections and permissions for the sampling of the Hazleton North material, many thanks go to Corinium Museum, Alison Brookes and Heather Dawson. We would also like to thank Simon Hickinbotham, University of York, for his assistance with generating computational R scripts for protein family classification. We are also grateful to the two anonymous reviewers for their insightful comments on the manuscript.
Funding
This research was funded by the UK Natural Environment Research Council (NERC) [grant number NE/K500987/1] and the Arts and Humanities Research Council (AHRC) [grant number AH/N005015/1].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Charlton, S., Ramsøe, A., Collins, M. et al. New insights into Neolithic milk consumption through proteomic analysis of dental calculus. Archaeol Anthropol Sci 11, 6183–6196 (2019). https://doi.org/10.1007/s12520-019-00911-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12520-019-00911-7