Abstract
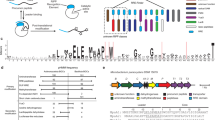
The ribosomally synthesized and post-translationally modified peptide (RiPP), pyrroloquinoline quinone (PQQ), is a dehydrogenase cofactor synthesized by, but not exclusively used by, certain prokaryotes. RiPPs represent a rapidly expanding and diverse class of natural products—many of which have therapeutic potential—and the biosynthetic pathways for these are gaining attention. Five gene products from the pqq operon (PqqA, PqqB, PqqC, PqqD, and PqqE) are essential for PQQ biosynthesis. The substrate is the peptide PqqA, which is presented to the radical SAM enzyme PqqE by the small protein PqqD. PqqA is unstructured in solution, and only binds to PqqE when in complex with PqqD. PqqD is a member of a growing family of RiPP chaperone proteins (or domains in most cases) that present their associated peptide substrates to the initial RiPP biosynthesis enzymes. An X-ray crystal structure exists for dimeric Xanthomonas campestris PqqD (PDB ID: 3G2B), but PqqD is now known to act as a monomer under physiological conditions. In this study, the PqqD truncation from naturally fused Methylobacterium extorquens (Mex) PqqCD was overexpressed in Escherichia coli and MexPqqA was chemically synthesized. Solution NMR 1H-,15N-HSQC chemical shift studies have identified the PqqD residues involved in binding PqqA, and 1H, 13C, and 15N peak assignments for PqqD alone and for PqqD bound to PqqA are reported herein.


Similar content being viewed by others
References
Anthony C (2001) Pyrroloquinoline quinone (PQQ) and quinoprotein enzymes. Antioxidant Redox Signal 3:757–774
Anthony C (2004) The quinoprotein dehydrogenases for methanol and glucose. Arch Biochem Biophys 428(1):2–9
Bauerly KA, Storms DH, Harris CB, Hajizadeh S, Sun MY, Cheung CP, Satre MA, Fascetti AJ, Tchaparian E, Rucker RB (2006) Pyrroloquinoline quinone nutritional status alters lysine metabolism and modulates mitochondrial DNA content in the mouse and rat. Biochim Biophys Acta 1760(11):1741–1748
Bauerly K, Harris C, Chowanadisai W, Graham J, Havel PJ, Tchaparian E, Satre M, Karliner JS, Rucker RB (2011) Altering pyrroloquinoline quinone nutritional status modulates mitochondrial, lipid, and energy metabolism in rats. PLoS ONE 6(7):e21779
Choi O, Kim J, Kim J-G, Jeong Y, Moon JS, Park CS, Hwang I (2008) Pyrroloquinoline quinone is a plant growth promotion factor produced by Pseudomonas fluorescens B16. Plant Physiol 146:657–668
Chowanadisai W, Bauerly KA, Tchaparian E, Wong A, Cortopassi GA, Rucker RB (2010) Pyrroloquinoline quinone stimulates mitochondrial biogenesis through cAMP response element-binding protein phosphorylation and increased PGC-1alpha expression. J Biol Chem 285(1):142–152
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Duine JA (1999) The PQQ story. J Biosci Bioeng 88(3):231–236
Goddard TD, Kneller DG (2014) SPARKY 3. University of California, San Francisco
Goodwin PM, Anthony C (1998) The biochemistry, physiology and genetics of PQQ and PQQ-containing enzymes. Adv Microb Physiol 40:1–80
Goosen N, Horsman HP, Huinen RG, van de Putte P (1989) Acinetobacter calcoaceticus genes involved in biosynthesis of the coenzyme pyrrolo-quinoline-quinone: nucleotide sequence and expression in Escherichia coli K-12. J Bacteriol 171(1):447–455
Harris CB, Chowanadisai W, Mishchuk DO, Satre MA, Slupsky CM, Rucker RB (2013) Dietary pyrroloquinoline quinone (PQQ) alters indicators of inflammation and mitochondrial-related metabolism in human subjects. J Nutr Biochem 24(12):2076–2084
Houck DR, Hanners JL, Unkefer CJ, van Kleef MA, Duine JA (1989) PQQ: biosynthetic studies in Methylobacterium AM1 and Hyphomicrobium X using specific 13C labeling and NMR. Antonie Van Leeuwenhoek 56(1):93–101
Kasahara T, Kato T (2003) Nutritional biochemistry: a new redox-cofactor vitamin for mammals. Nature 422(6934):832
Kay LE, Xu GY, Singer AU, Muhandiram DR, Forman-Kay JD (1993) A gradient-enhanced HCCH-TOCSY experiment for recording side-chain 1H and 13C correlations in H2O samples of proteins. J Magn Reson B 101:333–337
Kay LE, Xu GY, Yamazaki T (1994) Enhanced-sensitivity triple-resonance spectroscopy with minimal H2O saturation. J Magn Reson A109:129–133
Killgore J, Smidt C, Duich L, Romero-Chapman N, Tinker D, Reiser K, Melko M, Hyde D, Rucker RB (1989) Nutritional importance of pyrroloquinoline quinone. Science 245(4920):850–852
Latham JA, Iavarone AT, Barr I, Juthani PV, Klinman JP (2015) PqqD is a novel peptide chaperone that forms a ternary complex with the radical S-adenosylmethionine protein PqqE in the pyrroloquinoline quinone biosynthetic pathway. J Biol Chem 290(20):12908–12918
Live DH, Davis DG, Agosta WC, Cowburn D (1984) Long range hydrogen bond mediated effects in peptides: 15 N NMR study of gramicidin S in water and organic solvents. J Am Chem Soc 106:1939–1941
Matsumura H, Umezawa K, Takeda K, Sugimoto N, Ishida T, Samejima M, Ohno H, Yoshida M, Igarashi K, Nakamura N (2014) Discovery of a eukaryotic pyrroloquinoline quinone-dependent oxidoreductase belonging to a new auxiliary activity family in the database of carbohydrate-active enzymes. PLoS ONE 9(8):e104851
Montelione GT, Lyons BA, Emerson SD, Tashiro M (1992) An efficient triple resonance experiment using carbon-13 isotropic mixing for determining sequence-specific resonance assignments of isotopically-enriched proteins. J Am Chem Soc 114:10974–10975
Muhandiram DR, Kay LE (1994) Gradient-enhanced triple-resonance three-dimensional experiments with improved sensitivity. J Magn Reson B 103:203–216
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–222
Shen YQ, Bonnot F, Imsand EM, RoseFigura JM, Sjolander K, Klinman JP (2012) Distribution and properties of the genes encoding the biosynthesis of the bacterial cofactor, pyrroloquinoline quinone. Biochemistry 51(11):2265–2275
Singh AK, Pandey SK, Saha G, Gattupalli NK (2015) Pyrroloquinoline quinone (PQQ) producing Escherichia coli Nissle 1917 (EcN) alleviates age associated oxidative stress and hyperlipidemia, and improves mitochondrial function in ageing rats. Exp Gerontol 66:1–9
Steinberg FM, Gershwin ME, Rucker RB (1994) Dietary pyrroloquinoline quinone: growth and immune response in BALB/c mice. J Nutr 124(5):744–753
Steinberg F, Stites TE, Anderson P, Storms D, Chan I, Eghbali S, Rucker R (2003) Pyrroloquinoline quinone improves growth and reproductive performance in mice fed chemically defined diets. Exp Biol Med 228(2):160–166
Stites T, Storms D, Bauerly K, Mah J, Harris C, Fascetti A, Rogers Q, Tchaparian E, Satre M, Rucker RB (2006) Pyrroloquinoline quinone modulates mitochondrial quantity and function in mice. J Nutr 136(2):390–396
Unkefer CJ, Houck DR, Britt BM, Sosnick TR, Hanners JL (1995) Biogenesis of pyrroloquinoline quinone from 13C-labeled tyrosine. Methods Enzymol 258:227–235
van Kleef MA, Duine JA (1988) L-tyrosine is the precursor of PQQ biosynthesis in Hyphomicrobium X. FEBS Lett 237(1–2):91–97
Vuister GW, Bax A (1994) Measurement of four-bond HN-Hα J-couplings in staphylococcal nuclease. J Biomol NMR 4:193–200
Wishart DS, Sykes BD (1994) The 13C Chemical-Shift Index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J Biomol NMR 4:171–180
Yamazaki T, Forman-Ka JD, Kay LE (1993) Two-dimensional NMR experiments for correlating 13Cβ. and 1Hδ/ε chemical shifts of aromatic residues in 13C-labeled proteins via scalar couplings. J Am Chem Soc 115:11054–11055
Zhang J, Meruvu S, Bedi YS, Chau J, Arguelles A, Rucker R, Choudhury M (2015) Pyrroloquinoline quinone increases the expression and activity of Sirt1 and -3 genes in HepG2 cells. Nutr Res 35(9):844–849
Zimmerman DE, Kulikowski CA, Huang Y, Feng W, Tashiro M, Shimotakahar S, Chien C, Power R, Montelione GT (1997) Automated analysis of protein NMR assignments using methods from artificial intelligence. J Mol Biol 269:592–610
Acknowledgments
Financial support for this work came from the National Institutes of Health grants GM-066569 (CMW) and GM-039296 (JPK). Special thanks to the Minnesota NMR Center for both professional expertise and access to spectrometers, and to the Minnesota Supercomputing Institute for their assistance in installing and configuring the annealing software to run on the supercomputer, Mesabi (reducing 20 h anneals to 30 min).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Evans, R.L., Latham, J.A., Klinman, J.P. et al. 1H, 13C, and 15N resonance assignments and secondary structure information for Methylobacterium extorquens PqqD and the complex of PqqD with PqqA. Biomol NMR Assign 10, 385–389 (2016). https://doi.org/10.1007/s12104-016-9705-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-016-9705-8