Abstract
Public transcriptomic studies have shown that several genes display pronounced gender differences in their expression in the human brain, which may influence the manifestations and risk for neuronal disorders. Here, we apply a transcriptome-wide analysis to discover genes with gender-specific expression and significant alterations in public postmortem brain tissue from Alzheimer’s disease (AD) patients compared to controls. We identify the sex-linked ubiquitin-specific peptidase 9 (USP9) as an outstanding candidate gene with highly significant expression differences between the genders and male-specific underexpression in AD. Since previous studies have shown that USP9 can modulate the phosphorylation of the AD-associated protein MAPT, we investigate functional associations between USP9 and MAPT in further detail. After observing a high positive correlation between the expression of USP9 and MAPT in the public transcriptomics data, we show that USP9 knockdown results in significantly decreased MAPT expression in a DU145 cell culture model and a concentration-dependent decrease for the MAPT orthologs mapta and maptb in a zebrafish model. From the analysis of microarray and qRT-PCR experiments for the knockdown in DU145 cells and prior knowledge from the literature, we derive a data-congruent model for a USP9-dependent regulatory mechanism modulating MAPT expression via BACH1 and SMAD4. Overall, the analyses suggest USP9 may contribute to molecular gender differences observed in tauopathies and provide a new target for intervention strategies to modulate MAPT expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gender differences in the incidence and phenotypic manifestations of brain disorders like Alzheimer’s disease (AD) have been reported in several independent studies [1–5]. Findings for AD include significantly more severe pathology in women than in men [1], faster brain atrophy in females [2], as well as a higher AD incidence in females after adjusting for differential survival, which has mainly been observed in the oldest-age categories [3–5].
Some studies have suggested lifestyle differences as a contributing factor for these gender disparities, e.g., the Framingham Heart Study indicated that selective survival of men with a healthier cardiovascular risk profile and hence lower propensity to dementia could partly explain a higher lifetime risk for AD in women [6]. Moreover, hormonal mechanisms have been implicated in AD gender differences, and neuroprotective functions have been proposed for testosterone [7, 8], estrogen [9], and progesterone [10]. However, studies according to which the disease incidence rates in other neurodegenerative diseases show an almost opposite distribution between the genders as compared to AD (e.g., a greater risk for Parkinson’s disease in men than in women [11]), suggest that differences in hormonal neuroprotection without disease specificity are not the only factor influencing the observed gender differences in neurodegenerative disorders. Similarly, more severe AD-associated effects of the ε4 variant of the APOE gene, the largest known genetic risk factor for sporadic AD, reported for females [12] cannot account for the entire range of phenotypic differences in the disease between the genders, because cognitive impairment progresses faster in women than in men even when considering only individuals without APOE ε4 alleles [13].
Overall, previous studies suggest that a multitude of factors is likely to be involved in AD gender differences, and accordingly, system level analyses of biomolecular differences between the sexes in AD may help to identify and understand further relevant factors. Although gender-based disease alterations in individual biomolecules are unlikely to explain the full spectrum of observed gender-specific disease manifestations, their identification and analysis may provide new pointers to biomolecules with important regulatory functions in the studied disease, supporting the design of new therapeutic interventions.
Since major differences in brain gene expression levels between males and females have previously been reported [14], but not investigated in the context of AD, we have conducted a corresponding system-wide transcriptome analysis on public data. In subsequent experiments for the top candidate gene identified, the ubiquitin-specific peptidase 9 (USP9), we investigated which potential AD disease mediators the gene is linked to. USP9 is a gonosomal gene with a Y-chromosomal (USP9Y) and an X-chromosomal form. In humans, USP9X escapes X-inactivation [15]. The gene encodes a deubiquitinase [16], removing the ubiquitin moieties from specific proteins and thereby preventing their proteasomal degradation [15, 17]. It deubiquitinates monoubiquitinated SMAD4 and has been shown to be an essential component of the TGF-beta/BMP signaling cascade [16] (the full gene names and brief function descriptions for USP9, SMAD4, and the other main genes discussed in this manuscript is provided in Table 1).
In this study, we first describe the analyses of public transcriptomics data that led to the identification of USP9 as a gene with an outstanding gender-linked expression pattern and significant alterations in AD. We then discuss evidence for regulatory links between USP9 and the AD-associated protein tau (MAPT), derived from previous studies in the literature, as well as USP9 knockdown experiments we performed in two model systems, a zebrafish model and human DU145 prostate cancer cells. Analyzing the transcriptome alterations in the DU145 cells in response to the knockdown experiments at the level of cellular pathways and molecular networks, we find significant alterations in cytoskeleton remodeling processes and specifically, in the molecular network of microtubule-associated proteins (MAPs) and tubulins linked to MAPT. From a network analysis of this dataset, we derive a model for the involvement of USP9 in the regulation of MAPT expression and phosphorylation and discuss possible applications for developing USP9-based intervention strategies targeting tau hyperphosphorylation and aggregation in AD and other tauopathies.
Results and Discussion
USP9Y Displays Significant Gender-Linked Expression in the Adult Brain and Diminished Expression in Alzheimer’s Disease
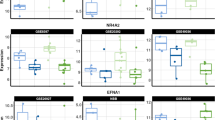
To investigate gender differences in the human brain transcriptome during adulthood which may contribute to sex differences observed in AD, we mined large-scale public datasets for genes that displayed both a generic gender-linked expression and significant expression alterations in AD. For this purpose, first, we computed male to female median gene expression level ratios using postmortem microarray data from a subset of 404 samples from 16 different brain regions across all unaffected adult individuals in the Human Brain Transcriptome Project (HBT) [14] (NCBI GEO dataset series GSE25219; the covered brain regions include orbital, dorsolateral, ventrolateral, and medial regions of the prefrontal cortex, primary motor cortex, primary somatosensory cortex, posterior inferior parietal cortex, primary auditory cortex, superior and inferior temporal cortex, primary visual cortex, hippocampus, amygdala, striatum, mediodorsal nucleus of the thalamus, and cerebellar cortex; see the “Methods” section for details on data preprocessing and analysis). As expected, the highest consistent male/female expression ratios across the 16 brain regions were observed for sex-linked genes, the top five being (ordered by decreasing ratio): PCDH11Y, RPS4Y1, DDX3Y, USP9Y, and EIF1AY (see Fig. 1a, showing a boxplot for USP9Y and USP9X as an example).
a Left: USP9Y normalized gene expression levels in postmortem brain tissue from adult males and females not affected by AD from the HBT dataset (as expected, the signal for the Y-chromosomal version of the gene in female samples does not significantly exceed the baseline, i.e., no expression is detected); right: USP9X normalized expression levels in postmortem brain tissue from adult males and females (expression levels are comparable across the genders, i.e., USP9X expression in females does not compensate the absence of USP9Y expression in females; see also the comparison across different age groups in Fig. 2c). b Boxplot of USP9Y gene expression levels in human postmortem hippocampus samples for incipient, moderate, and severe cases of male Alzheimer’s disease patients as compared to non-demented male controls (dataset by Blalock et al.). The reader should note that normalized expression level intensities are only comparable across different genes within one study but not across different studies
Next, we used data from an independent late-onset AD case/control transcriptomics study [19] (NCBI GEO dataset series GSE44772), covering brain samples from 690 individuals across three brain regions (prefrontal cortex, visual cortex, and cerebellum), to examine which of the identified genes with significant gender-linked expression in the adult human brain also displayed significant and consistent gender-specific expression alterations in AD across the covered brain regions. The rationale is that genes with robust expression alterations across multiple brain regions in a single study can be compared qualitatively with gene expression changes observed in other studies covering the same or further brain regions, to accumulate evidence for consistent multiregional alteration patterns. Among the identified genes with consistent multiregional gender-linked expression as well as gender-specific expression alterations in AD, the deubiquitinase USP9Y was the only gene showing a significantly reduced expression in male AD patients as compared to unaffected male subjects (adjusted p value <0.05) that matched with the multiregional changes observed on data from another AD case/control study with 161 microarray samples (p = 0.003) [20] (NCBI GEO dataset series GSE5281, covering the brain regions primary visual cortex, entorhinal cortex, hippocampus, medial temporal gyrus, posterior cingulated, and superior frontal gyrus) as well as a further AD case/control study with 30 hippocampal microarray samples [21] (NCBI GEO dataset series GSE28146; given the small sample size for the individual genders, on this third dataset significance could not be shown independently, p = 0.17, but a statistical meta-analysis across all datasets confirmed the significant underexpression across multiple brain regions, p < 2E−05). The dataset by Blalock et al. additionally groups AD patients into different stages of progression (“incipient,” “moderate,” and “severe” AD), and in all three stages, a qualitative reduction in median USP9Y expression is observed as compared to the control samples (the sample size is however too small to establish significance for the individual progression groups; see boxplot in Fig. 1b). Thus, we decided to focus our subsequent mechanistic investigation on USP9 gene regulation.
Associations Between USP9 Expression and the Expression and Phosphorylation of Tau
The AD-related microtubule-associated protein tau (MAPT) is involved in the stabilization of microtubules, which function as intracellular transport path [22]. Aggregations of paired helical filaments (PHFs) of MAPT into neurofibrillary tangles (NFTs) are known as one of the main hallmarks of AD and are commonly considered as neurotoxic but have also been proposed to represent secondary protective effects, since the reduction in microtubule assembly in AD is independent of MAPT abnormalities [23].
Similarly, different roles have been suggested for MAPT phosphorylation in AD, and potential toxic and protective effects have both been described [24, 25]. USP9 is linked mechanistically to MAPT phosphorylation. It has been shown that the X-chromosomal version of USP9 (USP9X) deubiquitinates the microtubule affinity-regulating kinase 4 (MARK4) [26], which in turn is known to phosphorylate MAPT [27]. A significant and strong increase of MARK4 expression and MARK4-MAPT interactions in AD brains, correlating with the Braak stages of the disease, has been found in postmortem human brains [28]. In a drosophila Alzheimer’s model, overexpression of the USP9 ortholog faf was reported to enhance phospho-MAPT-mediated postsynaptic toxicity of amyloid precursor protein (APP)/Aβ-42 [29]. Moreover, USP9X can increase MAPT phosphorylation via a second mechanism, by deubiquitinating the protein alpha-synuclein (SNCA) [30], which functions as a connecting mediator between the glycogen synthase kinase 3β (GSK3B) and MAPT and has been shown to stimulate MAPT phosphorylation via GSK3B in vitro [31]. While our analyses focus on the relations between USP9 and MAPT at the level of transcriptional regulation (see following sections), the prior knowledge on the involvement of USP9 in the regulation of MAPT phosphorylation provides a first line of evidence in support of a close functional relationship between these two genes/proteins.
For the Y-chromosomal version of USP9 (USP9Y), interactions with MARK4 and SNCA as observed for USP9X have not been reported so far, but USP9Y has a high sequence similarity of 93% to USP9X (quantified using the SIAS software tool with default settings, see http://imed.med.ucm.es/Tools, and the canonical protein sequences for UniProt IDs Q93008 and O00507) and is therefore expected to have largely similar structure and function (this is also confirmed by our cell culture model transcriptome analysis, comparing a USP9X-specific knockdown with a USP9X/Y knockdown; see details below).
Apart from previously reported functional relations between USP9 and MAPT at the protein level, the gene expression data analyzed here suggests a further possible association at the transcriptional level, which was analyzed in more detail via knockdown experiments in this study (see following sections). For a shared genetic probe for MAPT contained in the transcriptomics datasets by Liang et al. [20] and Blalock et al. [21], high positive correlations with USP9Y were found across the male samples (r = 0.483, p = 2.3E−07 for cases and controls combined, and r = 0.376, p = 5.5E−03 in controls only for the data by Liang et al.; r = 0.425, p = 0.169 for cases and controls combined, and r = 0.452, p = 0.368 in controls only on the small dataset by Blalock et al.). Similarly, among the unaffected male individuals of the older age group (age ≥ 40) in the HBT dataset, a high correlation between MAPT and USP9Y was observed (r = 0.480, p = 1.5E−06). Given that a higher correlation was observed in controls as compared to cases on the dataset by Blalock et al., as opposed to a lower correlation in controls compared to cases on the dataset by Liang et al., the results do not suggest that there is a general trend of higher MAPT/USP9Y correlations in either case or control samples. Generally, for larger sample sizes, improved p value significance scores were obtained for the MAPT/USP9Y correlation (i.e., when combining case and control samples as opposed to studying control samples only and when considering the HBT and Liang et al. datasets as opposed to the smaller Blalock et al. dataset).
Age-Dependent Expression of MAPT and USP9
MAPT and USP9Y expression levels were also compared across different adult age groups using brain transcriptomics data from individuals unaffected by AD from the Human Brain Transcriptome Project (HBT) project [14] (see the “Methods” section). In general, we observed a significant decrease in the levels of MAPT in the brain during healthy adult aging, matching with the results reported in an independent study [32]. Apart from this overall trend, we noted that the age-dependent decline in MAPT levels was gender-specific, with a faster decline in females as compared to males (see Fig. 2a). However, these gender differences in MAPT expression alone cannot explain the various gender differences in AD pathoetiology. Indeed, diverse types of gender differences have been observed across different MAPT-related diseases, e.g., a higher male preponderance for frontotemporal dementia [33] as opposed to an opposite risk profile in AD. While this indicates that MAPT-mediated gender intrinsic pathoetiology may be modulated also by other factors, it is perfectly in line with the role of MAPT in tauopathies.
Boxplot for the normalized gene expression levels of a MAPT, b USP9Y, and c USP9X across different age groups in males and females in the Human Brain Transcriptome dataset (covering only individuals unaffected by AD). a After 20 years of age, median MAPT levels show a general trend of continuous decline with age but decrease faster in females as compared to males (two-sided Welch test p value significance relative to the 12–20 years age group in the same gender: *p < 0.05, **p < 0.01, ***p < 0.001). Comparing the differences between males and females for each age group, females have significantly higher MAPT levels in the 12–20 years group (p = 1.27E−08), whereas no significant difference is found in the 20–40 years group, and female MAPT levels are significantly lower in the 40–60 years group (p = 0.024) and the >60 years group (p = 0.0012). b Up to the age group of 40 to 60 years, USP9Y levels tend to increase with higher age (p value significance for males relative to 12–20 years age group: *p < 0.05, **p < 0.01, *** p < 0.001). c USP9X levels show no significant differences in males and females after 20 years of age, i.e., during adulthood the absence of USP9Y expression in females is not compensated by higher USP9X expression in females as compared to males (shown are only the significant p values relative to the 12–20 years age group in the same gender: *p < 0.05, **p < 0.01, ***p < 0.001)
Given the associations between USP9 and MAPT described above, we also compared the genes in terms of the gender differences in their expression across different age groups. As shown in Fig. 2b, median USP9Y levels in males increase across the first three age groups (12 to 20, 20 to 40, and 40 to 60 years) with a slight relative decrease afterward. These changes in USP9Y were not compensated by corresponding increases in USP9X expression in females (see Fig. 2c).
Comparing Fig. 2a, b, the increased USP9Y expression in higher age groups in males coincides with a slower age-dependent decline of MAPT in males as compared to females, in line with the observed expression level correlations between USP9Y and MAPT. Since correlations do not necessarily imply a causal regulatory relationship and do not enable a distinction between cause and effect, we investigated a possible functional role of USP9Y as a positive regulator of MAPT by conducting USP9 knockdown experiments in a zebrafish model and in the human DU145 cell culture model (see following sections).
Tau Expression Is Decreased After USP9 Knockdown in Zebrafish
Among potential model organisms for the study of MAPT regulation, the zebrafish (Danio rerio) is of particular interest, since it has two MAPT paralogs, mapta and maptb, which reflect two major groups of MAPT isoforms in humans. While maptb is predominantly expressed as an isoform with three tubulin-binding repeats (3R-tau), mapta gives rise to isoforms with four to six repeats (4-6R-tau) [34, 35], which significantly increases its affinity for microtubules. Since 3R-tau and 4R-tau are the two main types of isoforms in humans, the two zebrafish paralogs may therefore provide a means to study how other regulatory genes influence the activity of the two isoforms.
The zebrafish is a well-established vertebrate model for the study of gene function and human pathologies [36, 37] and its genome displays a high degree of homology to the human genome [38]. In this study, we used zebrafish embryos as a model organism to investigate the possible regulatory relation between USP9 and MAPT, by studying alterations in the expression of mapta and maptb in response to the knockdown of the zebrafish USP9 ortholog (usp9). The knockdown was achieved using usp9 antisense morpholino oligonucleotides (MO) blocking exon2-intron2 (e2i2) splicing. Gene expression changes were assessed via qRT-PCR (see the “Methods” section for details on the experiment and analysis). A microscopic analysis comparing uninjected control, morpholino control, and usp9 MO-injected zebrafish showed no apparent toxic or developmental effect for the knockdown (see Fig. 3).
As shown in Fig. 4 at 2 days postfertilization, gene expression in usp9 morphants displays concentration-dependent decreases in both mapta and maptb, with a significant reduction at a MO concentration of 8 ng/injection (p < 0.05 for mapta and p < 0.01 for maptb). Overall, the results match with the high positive correlations between USP9Y and MAPT in the human postmortem transcriptomics datasets, further corroborating a possible regulatory relationship at the gene expression level.
Transcriptome-Wide Analysis of USP9X/Y Knockdown Effects
Gene-Level Expression Analysis of USP9 Knockdown Effects in DU145 Cells
To assess the biomolecular effects of knocking down USP9 on a transcriptome-wide scale, the human DU145 prostate cancer cell culture model was chosen, after confirming that USP9X, USP9Y, and MAPT are robustly expressed in this model. As a tumor-derived and non-neuronal cell line, DU145 can only provide an approximate model for the regulatory network around USP9 in primary neuronal cells but has the advantage that stable expression of all genes of interest could be confirmed (see discussion of cell line choice and limitations in the “Methods” section).
ShRNA constructs were designed for the knockdown of USP9X and USP9X/Y (i.e., targeting USP9X and USP9Y jointly) to infer potential USP9Y-specific gene alterations indirectly by comparing both knockdowns. As discussed in the following paragraphs, no significant differences in the alteration patterns between the two knockdowns could be detected, prompting us to combine the knockdown data for the subsequent pathway analyses in the section “Pathway and network analysis of USP9 knockdown effects on gene expression in DU145 cells”. Multiple shRNA constructs were tested for knockdown efficiency and validated functionality via gene expression analysis in whole-cell populations. Only the best construct for each target was used for further experiments. Limitations regarding potentially undetected off-target effects when using a single shRNA construct per target, as well as the design and selection of constructs, are covered in the “Methods” section. As readout, microarray expression profiling using the Affymetrix GeneChip Human Gene 2.0 ST platform was performed with triplicate samples for each of the three considered conditions (USP9X/Y knockdown, USP9X knockdown, scrambled RNA control). This dataset has been made publicly available in the Gene Expression Omnibus (GEO) database under series GSE79376. Additionally, for target genes of interest discussed below, a qRT-PCR validation was performed (see the “Methods” section).
After preprocessing and normalization (see the “Methods” section), expression alterations were investigated first in the main genes of interest (USP9X/Y and MAPT), filtering the genetic probes mapping to these genes to retain only those with an average expression greater than the average expression across all probes on the chip. As expected, in the USP9X/Y knockdown, a marked underexpression was observed in comparison to the control samples for all 16 genetic probes mapping to the USP9X gene and fulfilling the average expression criterion (see detailed statistics in Suppl. Table S1). For USP9Y, 14 out of 15 genetic probes passing the average expression filter also showed a strong reduction in expression levels (see Suppl. Table S1; the only probe with a positive log-fold change displayed a higher than average standard deviation and is therefore likely an artifact of noise). These expression level decreases in the USP9X/Y knockdown were confirmed in the qRT-PCR validation (p = 4.6E−10 for USP9X and p = 4.77E−07 for USP9Y). For the USP9X-specific knockdown, an underexpression trend was also confirmed (see Suppl. Table S1).
Interestingly, gene expression levels for MAPT were consistently decreased for the USP9X/Y knockdown for all of eight filtered MAPT probes and in the qRT-PCR validation, as well as for all seven probes passing the abovementioned average expression filter for the USP9X knockdown (see Suppl. Table S1). These results are in line with the observed underexpression of mapta and maptb in the zebrafish usp9 knockdown (see Figs. 4 and 5). Moreover, the MAPT probe expression levels displayed positive average correlations with those for the USP9X probes (avg. Pearson correlation across all pairs of filtered probes: r = 0.479, avg. p = 0.096) and USP9Y probes (avg. r = 0.472, avg. p = 0.082). Similar correlations were also observed in the AD brain transcriptomics datasets by Liang et al. [20] (r = 0.483, p = 2.3E−07) and Blalock et al. [21] (r = 0.425, p = 0.169).
Normalized MAPT expression levels on log-scale in the USP9XY and USP9X knockdowns (KD) and the scrambled control (scrambled) for the MAPT genetic probe with the highest average expression in the DU145 microarray dataset. Decreased median MAPT expression levels are observed for both knockdowns (p = 0.017 for the USP9XY knockdown and p = 0.44 for the USP9X knockdown)
To identify possible molecular mechanisms behind the underexpression of MAPT in the USP9X/Y and USP9X knockdowns, we conducted an upstream network analysis, investigating transcription factors (TFs) known to be involved in the regulation of MAPT transcription for gene expression alterations in our dataset, using literature-curated TF-target relationships from the Biobase Proteome™ database (see http://www.biobase-international.com/proteome-2), the ResNet® Mammalian database 11 from Elsevier (see http://www.ariadnegenomics.com), and GeneCards [18]. MAPT transcriptional regulators were filtered according to the consistency of their expression alterations and the decrease of MAPT levels observed in the DU145 USP9X/Y knockdown gene expression data with their reported regulatory effect on MAPT. One known MAPT transcription repressor, BACH1 [39], was found to have increased expression levels in the USP9X/Y knockdown for five out of seven filtered genetic probes and for seven out of eight filtered probes for the USP9X knockdown (see Suppl. Table S1). By investigating BACH1 upstream regulators using regulatory interactions from the ResNet Mammalian database, a mechanistic link between BACH1 and USP9 was found. Since USP9 is known to deubiquitinate SMAD4 [16], loss of USP9 orthologs disables SMAD4-dependent responses in several model systems [16], and SMAD4 knockdown has been shown to constitutively activate BACH1 expression [40], and the USP9X/Y knockdown effect on BACH1 expression and its downstream target MAPT can be explained by an inhibition of the common monoubiquitinated SMAD4 (see also our proposed mechanistic model for the involvement of USP9 in MAPT expression regulation in Fig. 6b). Specifically, SMAD4 is monoubiquitinated by the E3 Monoubiquitin Ligase TRIM33 (Ectodermin/Tif1γ), inhibiting SMAD4 responses, an action which is opposed by the deubiquitinase USP9 [16].
a Role of USP9 in the regulation of MAPT phosphorylation using relations derived from the literature (see labels of the directed edges). b Proposed model for the role of USP9 in the regulation of MAPT gene expression, derived from the upstream analysis of the DU145 transcriptomics data. In both a and b, genes/proteins are represented by ellipses and colored blue for transcription factors, beige for kinases, and green for all other proteins. Small blue arrows pointing downward represent genes with decreased mRNA levels after USP9X/Y knockdowns in DU145 cells, and red arrows pointing upward highlight increased mRNA levels observed after the knockdowns
Overall, in spite of a higher variance across replicate samples in the USP9X knockdown, a highly significant Pearson correlation between the log-fold expression changes for the USP9X/Y and the USP9X knockdown was observed both across the entire DU145 transcriptomics dataset (r = 0.56, p < 2.2E−16) and in particular when considering only the top 1000 most significant shared genetic probes (r = 0.965, p < 2.2E−16), which are expected to be less strongly affected by random variation than the entire set of transcripts. Furthermore, no significant difference in the alterations between the significantly differentially expressed genes in the USP9X/Y knockdown and the USP9X knockdown could be detected. This observation may result from a shared functional profile of USP9X and USP9Y, or alternatively, from a lack of statistical power to detect smaller differences in downstream gene expression changes or an insufficient specificity of the knockdowns. In the USP9X knockdown, a small but not statistically significant decrease in USP9Y expression was observed in the qRT-PCR measurements (p = 0.759), which may indicate a weak off-target effect of this knockdown on USP9Y. Regarding the possibility of a lack of statistical power to reliably detect small gene expression differences between the knockdowns, this cannot be excluded, but the high correlation of fold changes observed across the top 1000 shared genetic probes, which include a majority of probes with small effect sizes in their alterations in the individual differential expression analyses, does not point to any significant shifts in expression pattern, and no clear outliers were observed that would indicate a biologically meaningful difference for specific genes between the knockdowns. Alterations for genetic probes ranked lower than the top 1000 have too small effect sizes or too high variation to be distinguished from random variation (these probes all have adjusted p values for differential expression of at least 0.45). In summary, we could not detect robust gene expression differences between the knockdowns or indications of such differences that would warrant a qPCR validation and could be used for biological data interpretation. Although we cannot exclude the existence of functional differences between the X- and Y-forms of USP9 due to the limitations of the omics profiling and knockdown approach, our results rather support the hypothesis that USP9-related gender differences result solely from differences in total USP9 expression levels (i.e., the sum of USP9X and USP9Y expression levels). Given the highly correlated alteration patterns at the level of single genes in the two knockdowns, the lack of detectable statistically significant differences between the knockdowns, and the high protein sequence similarity between USP9X and USP9Y of 93%, which also suggests a high structural and functional similarity, we have therefore decided to combine the USP9X/Y and USP9Y knockdown groups for the subsequent analyses to increase the statistical power for the investigation of pathway and network alterations.
Pathway and Network Analysis of USP9 Knockdown Effects on Gene Expression in DU145 Cells
To identify cellular pathways with jointly altered activity in the USP9XY and USP9X knockdowns, a pathway enrichment analysis was performed using the GeneGO software (see Methods). Table 2 reports the top ten cellular pathways with an overrepresentation of differentially expressed genes in the knockdowns. These altered pathways mainly include processes related to cytoskeleton remodeling, cell adhesion, and stress signaling (ordered by significance).
Studying the molecular network regions associated with cytoskeleton remodeling in more detail revealed pronounced expression changes in several tubulins and microtubule-associated genes (e.g., TUBA1B, TUBA3C, TUBA3E, TUBA4B, TUBB2A, TUBB4A, TUBB8, MAP2, and MAP4). Overall, tubulins tend to have reduced expression in the knockdowns with few exceptions (e.g., TUBA8 and TUBG2). Supplementary Fig. S1 shows a corresponding subnetwork from the pathway “Regulation of cytoskeleton proteins in oligodendrocyte differentiation and myelination” (adjusted p value: 2.28E−05). More specifically, the figure highlights that tubulins involved in binding interactions with microtubule-associated protein tau (MAPT) are underexpressed, matching with the decreased expression observed for MAPT itself (see above). These results suggest that the underexpression of MAPT in the USP9XY/USP9X knockdowns is part of a broader response affecting a subnetwork involving MAPT and tubulins within cytoskeleton regulation-associated processes. The increased expression of the transcription factor BACH1, which represses MAPT expression, may partly explain these changes, since multiple tubulins are predicted as further target genes repressed or activated by BACH1 according to the SABioscience DECODE database (www.sabiosciences.com), including TUBA1A, TUBA1C, TUBA4A, TUBA4B, TUBA8, TUBB, TUBB2A, TUBB2C, TUBB3, TUBB6, TUBD1, TUBG2, and TUBGCP2. The DECODE database contains binding site predictions derived from SABioscience’s proprietary Text Mining Application and manual genome annotations from the UCSC Genome Browser [41] (the database was queried via the SABioscience “Champion ChiP Transcription Factor Search Portal” at www.sabiosciences.com, searching for the transcription factor BACH1 and setting the species to “Human”). Interestingly, USP9X has previously also been found to associate with microtubules in neuronal processes and to interact in vivo with doublecortin (DCX), a microtubule-associated protein involved in neuronal migration [42].
Network Model of USP9 Knockdown Effects on MAPT Phosphorylation and Expression
In Fig. 6, we combine information from the literature on the involvement of USP9 in MAPT phosphorylation regulation (Fig. 6a) and our findings from the upstream network analysis in a mechanistic model for USP9-mediated regulation of MAPT gene expression (Fig. 6b). As outlined in Fig. 6a, previous evidence from the literature shows that USP9 can influence the phosphorylation of MAPT both by deubiquitinating the kinase MARK4 [26], which directly phosphorylates MAPT, and by deubiquitinating SNCA [30], which mediates the connection between glycogen synthase kinase 3β (GSK3B) and its phosphorylation target MAPT [31]. MAPT expression can be altered by USP9-mediated deubiquitination of the transcription factor SMAD4 [16], which represses BACH1 transcription. BACH1, in turn, encodes a transcriptional repressor for MAPT [39] (see Fig. 6b).
Additional functional relationships between USP9 and the cytoskeleton are suggested by the interactomes of MARK4 and BACH1. Apart from MAPT, MARK4 also phosphorylates further tau family microtubule-associated proteins (MAPs) [27]. The transcriptional MAPT repressor BACH1 has several additional tubulin targets, explaining why tubulins are underexpressed in the USP9X/Y knockdown (see Suppl. Fig. S1).
Interestingly, MARK phosphorylation of tau family MAPs has been shown to trigger microtubule disruption [43], and inhibitors of tau and MAP phosphorylation have previously been proposed as drug targets for Alzheimer’s disease (AD). For example, GSK3B inhibition via small molecule ligands has been suggested as a possible therapeutic intervention strategy to counteract MAPT hyperphosphorylation in AD [44], and heat shock protein 70 (HSP70), which suppresses alpha-synuclein (SNCA)-induced MAPT phosphorylation via GSK3B through direct binding to SNCA, has also been proposed as a therapeutic target [31]. In a similar fashion, modulation of USP9X/Y activity could be of interest as a new intervention strategy to alter both the phosphorylation of MAPT (via MARK4 or SNCA/GSK3B, as outlined in Fig. 6a) and MAPT expression levels (via SMAD4 and BACH1, see Fig. 6b).
Conclusions
The human brain transcriptome displays several significant gender differences with large effect size, which may influence the risk for brain disorders and the severity of their phenotypic manifestations. In particular, in this study, we observe gender-specific differences in the age-dependent decrease of brain gene expression levels for the microtubule-associated protein tau (MAPT), a protein playing a central role in neurodegenerative diseases referred to as tauopathies. In a joint analysis of multiple transcriptomics datasets from independent studies, we identify USP9 as a candidate MAPT regulator behind these gender differences. Previous studies have shown that USP9 can influence the phosphorylation of MAPT via deubiquitination of the MAPT-phosphorylating kinase MARK4 and indirectly via deubiquitination of SNCA, which stimulates MAPT phosphorylation by mediating its connection with the kinase GSK3B. At the gene transcription level, USP9 knockdown experiments in zebrafish and the DU145 human cell culture result in reduced MAPT levels, confirming the positive correlation between USP9 and MAPT observed in human brain transcriptomics datasets.
A mechanistic explanation for this correlation is provided by the upstream network analysis of the USP9 knockdown transcriptome, exploiting existing molecular interaction data and identifying a regulatory relation between USP9 and MAPT via the USP9 deubiquitination target SMAD4 and the MAPT transcription repressor BACH1, whose transcription is in turn repressed by SMAD4. This potential pathway merits further validation and investigation with regard to a putative relevance for AD molecular pathology and intervention strategies. Since the transcription factor BACH1 is predicted to target multiple tubulins and the kinase MARK4 is involved in the phosphorylation of multiple microtubule-associated proteins (MAPs), the alterations in these regulators also provide a parsimonious explanation for the main changes observed in the pathway and network analysis of the USP9 knockdown effects. Overall, the pathway/network analysis shows that cellular processes related to cytoskeleton remodeling, as well as tubulins and MAPs in the interaction network around MAPT, are most significantly affected.
In summary, in the context of prior knowledge from the literature, the presented transcriptomics data supports the existence of mechanistic links between USP9 and MAPT and suggests that USP9 could be of biomedical interest as a regulator modulating both the expression and phosphorylation of MAPT. For tauopathies like Alzheimer’s disease, involving aggregations of paired helical filaments of MAPT into neurofibrillary tangles, USP9 may therefore warrant further study as a target for the development of new intervention strategies.
Materials and Methods
Gene Expression Data Processing and Analyses
All raw microarray datasets from Affymetrix platforms were preprocessed and normalized using the GC-RMA approach [45]. For the dataset from the study by Zhang et al., derived from Rosetta/Merck Human 44 k 1.1 chips and not suitable for the GC-RMA procedure, the preprocessed data according to the procedures in the original publication was used [19]. Differential gene expression was scored and analyzed using the empirical Bayes moderated t-statistic [46], and the resulting p value significance scores were adjusted for multiple hypothesis testing following the approach by Benjamini and Hochberg [47].
To account for differences in brain regions covered across the microarray datasets used, we focused on genes that displayed consistent, multiregional expression changes in the HBT dataset [14] (NCBI GEO dataset series GSE25219) and the late-onset AD case/control dataset [19] (NCBI GEO dataset series GSE44772), i.e., genes with consistent signs for the logarithmic fold changes and with overall significant p values after adjustment for multiple hypothesis testing (adjusted p value <0.05). While the GSE25219 dataset covers 16 brain regions (see the section “USP9Y displays significant gender-linked expression in the adult brain and diminished expression in Alzheimer’s disease”), the GSE44772 dataset covers three regions (the prefrontal cortex, visual cortex, and cerebellum, which represent higher-level groupings of brain regions that overlap with the 16 brain regions for the GSE25219 dataset). The reason for focusing on genes with consistent expression alterations across multiple brain regions is that additional evidence for corresponding multiregional alteration patterns can be collected across datasets from other studies, which cover the same or further brain regions. For the gene of interest derived from the analyses on the GSE25219 and GSE44772 dataset, USP9Y, which showed both consistent multiregional gender-linked expression (GSE25219 dataset) and multiregional gender-specific expression alterations in AD (GSE44772 dataset), two further microarray datasets were studied to investigate whether the significantly reduced expression in male AD patients as compared to unaffected male subjects observed on the GSE44772 dataset across the prefrontal cortex, visual cortex, and cerebellum, could also be found in the brain regions covered by other AD case/control datasets. Specifically, significant multiregional expression changes for USP9Y were confirmed in the NCBI GEO dataset GSE5281 [20] (covering the brain regions primary visual cortex, entorhinal cortex, hippocampus, medial temporal gyrus, posterior cingulated, and superior frontal gyrus), and a matching qualitative change was observed in hippocampal samples of the NCBI GEO dataset GSE28146 [21] (see the section “USP9Y displays significant gender-linked expression in the adult brain and diminished expression in Alzheimer’s disease”).
A limitation of these investigations of consistent gene expression alterations across multiple brain regions is that the quantitative statistical results for specific brain regions on a single dataset cannot be directly integrated with the statistics obtained from independent datasets covering distinct brain regions to increase the statistical power. However, by using datasets with large sample sizes and coverage of multiple brain regions to identify genes with multiregional consistent expression changes, a qualitative confirmation of multiregional expression alterations is possible on independent datasets, which may partly cover distinct brain regions (comparing the statistics on the separate datasets, instead of integrating them into a single statistic).
To exclude age-related biases in the significant findings for the analyses of microarray datasets from the AD case/control studies, the significance of gene expression alterations between patients and controls was confirmed after adjustment for age using a surrogate variable analysis (R software package SVA [48]). As a separate investigation related to aging-associated changes, a transcriptome-wide analysis and comparison of brain gene expression alterations during adult aging and in the neurodegenerative disorders Alzheimer’s and Parkinson’s disease has been presented previously [49].
To conduct cellular pathway analyses, the differential expression analysis results for the combined knockdown samples as compared to the control samples were used as input for the GeneGO pathway analysis software [50], filtering out genetic probes with a p value above 0.05. Pathways with an overrepresentation of differentially expressed genes were ranked by increasing p value. For the upstream network analysis, literature-curated TF-target relationships from the Biobase Proteome™ database (see http://www.biobase-international.com/proteome-2) and GeneCards [18] were used, and the consistency of potentially relevant TF-target pairs with the alteration patterns observed in the transcriptomics data was checked.
For qRT-PCR measurement analyses, the data was normalized against the mean of multiple reference genes according to the method by Hellemans et al. [51] (see details on the selected reference genes and the experimental design in the “Method” sections on the zebrafish and cell culture experiments). The Welch’s t test was then applied to assess the significance of differential expression (changes with a significance of p < 0.05 are highlighted by star symbols in the corresponding boxplots, see Fig. 4).
Zebrafish Experiments
Knockdown Experiments
The usp9 antisense morpholino (MO) was designed and synthesized to target the splice donor site of exon 2 (5′-TGAAAATGGTGCTCTGACCTGGTTC-3′) and to interfere with normal pre-mRNA splicing of the zebrafish usp9 gene (ENSDART00000135384). To analyze gene expression changes after usp9 antisense morpholino MO knockdown in zebrafish for the target genes usp9, and the zebrafish MAPT paralogs mapta and maptb, titrated microinjections were performed with 4.8 and 8 ng/injection of an e2i2 splice blocking usp9 MO in one to two cell-stage zebrafish embryos. Control MO (randomized 25 N oligomer, 8 ng/injection) and wild-type embryos (non-injected controls) were processed in parallel. All MOs were designed and synthesized by Gene Tools. Embryos were maintained in standard conditions (28 °C) in embryo medium (0.3× Danieau’s). Two days after microinjection, 20 dechorionated embryos per condition were processed for RNA extraction and reverse transcription. For each target gene to be assessed via qRT-PCR gene expression measurements (LightCycler®480, Roche), three different pairs of primers were initially tested for specificity, and then the most specific pair was selected for further experiments (see Suppl. Table S2). Three zebrafish housekeeping (HK) genes were processed in parallel (β-actin1, elongation factor 1, and 60S ribosomal protein L13). The qRT-PCR was performed with five biological replicates per condition, and each biological replicate was covered by four technical replicates.
Cell Culture Experiments
In order to study USP9X/Y and its gene regulatory network in a cell culture model, relevant human cell lines were first compared in terms of the stability of USP9X, USP9Y, and MAPT and expression using information from the literature and public gene expression data. The prostate carcinoma cell line DU145 was chosen, since the public transcriptomics data showed stable expression for the genes of interest (USP9Y, USP9X, MAPT), as opposed to the alternative neuroblastoma cell line from female origin SH-SY5Y (ATCC no. CRL-2266, undifferentiated) and the human embryonic kidney cell line HEK293 (ATCC no. CRL-1573, see Suppl. Fig. S2 and S3 and Suppl. Tables S3 and S4). Since DU145 cells are derived from a non-neuronal tumor tissue, the molecular regulatory network around USP9 present in this cell line can only provide an approximation to a corresponding network in a genetically stable, neuronal cell population. Possible limitations include that yet unknown regulatory genes in this network and downstream effectors of USP9 with tissue-specific expression in primary neuronal cells may not be expressed in DU145 cells, and that like in other tumor cell lines, the metabolism may be shifted toward proliferation and growth. We therefore consider the analysis of USP9 in DU145 cells in combination with our findings in the zebrafish model, the analyses of human brain microarray data, and information from the published literature, to avoid over-reliance on a single model and information source.
DU145 cells were obtained from the American tissue culture collection (ATCC no. HTB-81). Cells were cultured in Dulbecco’s modified eagle medium (Invitrogen no. 41966-029) containing high glucose (25 mM), l-glutamine (4 mM), and sodium pyruvate (1 mM). This medium was supplemented with 10% v/v heat-inactivated fetal bovine serum (Invitrogen no. 10500-064). Cells were grown at 37 °C at 5% CO2 and saturated humidity.
Knockdown plasmids were designed to either target USP9X and USP9Y jointly or USP9X specifically. Due to the high sequence homology between USP9X and USP9Y, the production of potentially unspecific USP9Y shRNA plasmids was omitted. Hairpin sequences (Suppl. Tab. S5) and U6 promoters were subcloned into FastBac plasmids (PMID 15771966). The list of all primers tested for the knockdown experiments is provided in Suppl. Tab. S3, and the agarose gel electrophoresis results are shown in Fig. S3. While multiple shRNA constructs per target gene were designed and tested for knockdown efficiency via gene expression analysis in whole-cell populations, for each target only, the construct with the best knockdown efficiency was used for subsequent profiling. As the efficiency of gene silencing may vary for different shRNA constructs and off-target effects can occur in some cases, limitations may arise in the interpretation of data derived from the use of a single construct. However, since the highly significant inhibition of the target genes observed in both microarray and qRT-PCR expression measurements (see the section “Gene-level expression analysis of USP9 knockdown effects in DU145 cells”) was not matched or approximated by expression changes observed in any other gene in the differential expression ranking derived from the microarray data, at least a strong off-target alteration can be excluded, as well as a scenario in which a larger amount of the target transcripts escapes the inhibition to recover the normal function. Transfection into DU145 was done via Lipofectamine 2000 (Invitrogen no. 11668019) and transfected cells were incubated for 48 h. Prior to RNA extraction, the perturbed cells were enriched via fluorescence-activated cell sorting. RNA was extracted using the Qiagen RNeasy Mini Kit (Qiagen no. 74106) and treated with DNaseI (Qiagen no. 79254). Reverse transcription was done as described previously (PMID 26738520). For microarray expression profiling, RNA extracts were prepared using the GeneChip WT PLUS Reagent Kit (Affymetrix, Manual P/N 703174 Rev. 2 and UserGuide GeneChip Expression Wash, Stain and Scan for Cartridge Arrays P/N 702731 Rev. 4). RNA quality and integrity was checked using a NanoDrop ND-100 spectrophotometer (Thermo Scientific) and the 2100 Agilent Bioanalyzer (Agilent), respectively. The purified, sense-strand cDNA was fragmented by uracil-DNA glycosylase (UDG) and apurinic/apyrimidinic endonuclease 1 (APE 1) at the unnatural dUTP residues. The fragmented cDNA was labeled by terminal deoxynucleotidyl transferase (TdT) using the Affymetrix proprietary DNA Labeling Reagent that is covalently linked to biotin. Single-stranded cDNA (5.5 μg) was used for fragmentation and labeling, and the GeneChip Hybridization, Wash and Stain Kit was used to hybridize and wash the cartridges. Control Oligonucleotide B2 and 20X Eukaryotic Hybridization Controls were added to the hybridization cocktail containing the labeled sample and injected into the cartridge. The incubation lasted 16 h at 45 °C with a rotation at 60 rpm. Then, the Fluidics Station 450/250 was used to wash and stain the Affymetrix GeneChip Human Gene 2.0 ST probe arrays. The probe arrays were scanned after completion of the wash protocols using the Affymetrix GeneChip Scanner 3000.
Data was processed as described in the “Methods” section on gene expression data processing and analysis. Both the raw and processed data have been deposited in the Gene Expression Omnibus (GEO) database under the series accession number GSE79376.
For validation of microarray-derived gene expression data in USP9 knockdown conditions, shRNA perturbations in DU145, cell sorting, RNA extraction, and reverse transcription were redone as described above. Real-Time PCR expression measurements for the target genes of interest and six reference genes were performed using a Fluidigm 48.48 integrated fluidic circuit array. The six reference genes consisted of three genes commonly used as reference for human cell culture RNA expression measurements (PPIA, GAPDH, and PDHB) and three genes displaying the lowest variation in the processed Affymetrix GeneChip microarray data for the DU145 cells (EIF4G2M, HSPA8, and UBAP2L). The assay IDs and catalog numbers for the TaqMan® gene expression assays used for the target and reference genes are provided in Suppl. Table S6. For each condition (USP9X/Y knockdown, USP9X knockdown, and three scrambled RNA control samples) measurements were obtained for five biological replicates.
To preprocess and analyze the raw data from the Fluidigm platform, an optimal subset of two reference genes (PPIA and HSPA8) was determined using the method by Vandesompele et al. [52] as implemented in the R software package SLqPCR (http://www.bioconductor.org). After determining the median across the measurements for the selected reference genes, a ΔCt calculation was performed as described in Yuan et al. [53] and the empirical Bayes moderated t-statistic [46] was applied to compare USP9X/Y knockdown samples against controls.
References
Barnes LL, Wilson RS, Bienias JL et al (2005) Sex differences in the clinical manifestations of Alzheimer disease pathology. Arch Gen Psychiatry 62:685–691. doi:10.1001/archpsyc.62.6.685
Hua X, Hibar DP, Lee S et al (2010) Sex and age differences in atrophic rates: an ADNI study with n = 1368 MRI scans. Neurobiol Aging 31:1463–1480. doi:10.1016/j.neurobiolaging.2010.04.033
Fratiglioni L, Viitanen M, von Strauss E et al (1997) Very old women at highest risk of dementia and Alzheimer’s disease: incidence data from the Kungsholmen project, Stockholm. Neurology 48:132–138. doi:10.1212/WNL.48.1.132
Miech RA, Breitner JCS, Zandi PP et al (2002) Incidence of AD may decline in the early 90s for men, later for women: the Cache County study. Neurology 58:209–218. doi:10.1212/WNL.58.2.209
Andersen K, Launer LJ, Dewey ME et al (1999) Gender differences in the incidence of AD and vascular dementia: the EURODEM studies. Neurology 53:1992–1992. doi:10.1212/WNL.53.9.1992
Chêne G, Beiser A, Au R et al (2014) Gender and incidence of dementia in the Framingham Heart Study from mid-adult life. Alzheimers Dement. doi:10.1016/j.jalz.2013.10.005
Pike CJ (2001) Testosterone attenuates beta-amyloid toxicity in cultured hippocampal neurons. Brain Res 919:160–165
Moffat SD, Zonderman AB, Metter EJ et al (2004) Free testosterone and risk for Alzheimer disease in older men. Neurology 62:188–193. doi:10.1212/WNL.62.2.188
Henderson VW, Watt L, Buckwalter JG (1996) Cognitive skills associated with estrogen replacement in women with Alzheimer’s disease. Psychoneuroendocrinology 21:421–430. doi:10.1016/0306-4530(95)00060-7
Stein DG (2008) Progesterone exerts neuroprotective effects after brain injury. Brain Res Rev 57:386–397. doi:10.1016/j.brainresrev.2007.06.012
Wooten GF, Currie LJ, Bovbjerg VE et al (2004) Are men at greater risk for Parkinson’s disease than women? J Neurol Neurosurg Psychiatry 75:637–639. doi:10.1136/jnnp.2003.020982
Damoiseaux JS, Seeley WW, Zhou J et al (2012) Gender modulates the APOE 4 effect in healthy older adults: convergent evidence from functional brain connectivity and spinal fluid tau levels. J Neurosci 32:8254–8262. doi:10.1523/JNEUROSCI.0305-12.2012
Lin KA, Choudhury KR, Rathakrishnan BG et al (2015) Marked gender differences in progression of mild cognitive impairment over 8 years. Alzheimer’s Dement Transl Res Clin Interv. doi:10.1016/j.trci.2015.07.001
Kang HJ, Kawasawa YI, Cheng F et al (2011) Spatio-temporal transcriptome of the human brain. Nature 478:483–489. doi:10.1038/nature10523
Jones MH, Furlong RA, Burkin H et al (1996) The drosophila developmental gene fat facets has a human homologue in Xp11.4 which escapes X-inactivation and has related sequences on Yq11.2. Hum Mol Genet 5:1695–1701. doi:10.1093/hmg/5.11.1695
Dupont S, Mamidi A, Cordenonsi M et al (2009) FAM / USP9x, a deubiquitinating enzyme essential for TGFb signaling, controls Smad4 monoubiquitination. Cell 136:123–135. doi:10.1016/j.cell.2008.10.051
Wood SA, Pascoe WS, Ru K et al (1997) Cloning and expression analysis of a novel mouse gene with sequence similarity to the drosophila fat facets gene. Mech Dev 63:29–38
Rebhan M, Chalifa-Caspi V, Prilusky J, Lancet D (1998) GeneCards: a novel functional genomics compendium with automated data mining and query reformulation support. Bioinformatics 14:656–664. doi:10.1093/bioinformatics/14.8.656
Zhang B, Gaiteri C, Bodea L-G et al (2013) Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153:707–720. doi:10.1016/j.cell.2013.03.030
Liang WS, Reiman EM, Valla J et al (2008) Alzheimer’s disease is associated with reduced expression of energy metabolism genes in posterior cingulate neurons. Proc Natl Acad Sci U S A 105:4441–4446. doi:10.1073/pnas.0709259105
Blalock EM, Geddes JW, Chen KC et al (2004) Incipient Alzheimer’s disease: microarray correlation analyses reveal major transcriptional and tumor suppressor responses. Proc Natl Acad Sci U S A 101:2173–2178. doi:10.1073/pnas.0308512100
Sheetz MP, Vale R, Schnapp B et al (1986) Vesicle movements and microtubule-based motors. J Cell Sci Suppl 5:181–188
Cash AD, Aliev G, Siedlak SL et al (2003) Microtubule reduction in Alzheimer’s disease and aging is independent of tau filament formation. Am J Pathol 162:1623–1627. doi:10.1016/S0002-9440(10)64296-4
Castellani RJ, Nunomura A, Lee H et al (2008) Phosphorylated tau: toxic, protective, or none of the above. J Alzheimers Dis 14:377–383. doi:10.1016/j.bbi.2008.05.010
Lee H-G, Perry G, Moreira PI et al (2005) Tau phosphorylation in Alzheimer’s disease: pathogen or protector? Trends Mol Med 11:164–169. doi:10.1016/j.molmed.2005.02.008
Al-Hakim AK, Zagorska A, Chapman L et al (2008) Control of AMPK-related kinases by USP9X and atypical Lys(29)/Lys(33)-linked polyubiquitin chains. Biochem J 411:249–260. doi:10.1042/BJ20080067
Trinczek B, Brajenovic M, Ebneth A, Drewes G (2004) MARK4 is a novel microtubule-associated proteins/microtubule affinity-regulating kinase that binds to the cellular microtubule network and to centrosomes. J Biol Chem 279:5915–5923. doi:10.1074/jbc.M304528200
Gu GJ, Lund H, Wu D et al (2013) Role of individual MARK isoforms in phosphorylation of tau at Ser 262 in Alzheimer’s disease. Neruomol Med 15:458–469. doi:10.1007/s12017-013-8232-3
Lee S, Wang JW, Yu W, Lu B (2012) Phospho-dependent ubiquitination and degradation of PAR-1 regulates synaptic morphology and tau-mediated Abeta toxicity in drosophila. Nat Commun 3:1312. doi:10.1038/ncomms2278
Rott R, Szargel R, Haskin J et al (2011) α-Synuclein fate is determined by USP9X-regulated monoubiquitination. Proc Natl Acad Sci U S A 108:18666–18671. doi:10.1073/pnas.1105725108
Kawakami F, Suzuki M, Shimada N et al (2011) Stimulatory effect of α-synuclein on the tau-phosphorylation by GSK-3β. FEBS J 278:4895–4904. doi:10.1111/j.1742-4658.2011.08389.x
Hayesmoore JBG, Bray NJ, Cross WC et al (2009) The effect of age and the H1c MAPT haplotype on MAPT expression in human brain. Neurobiol Aging 30:1652–1656. doi:10.1016/j.neurobiolaging.2007.12.017
Ratnavalli E, Brayne C, Dawson K, Hodges JR (2002) The prevalence of frontotemporal dementia. Neurology 58:1615–1621. doi:10.1212/WNL.58.11.1615
Moussavi Nik SH, Newman M, Ganesan S et al (2014) Hypoxia alters expression of zebrafish microtubule-associated protein tau (mapta, maptb) gene transcripts. BMC Res Notes 7:767. doi:10.1186/1756-0500-7-767
Chen M, Martins RN, Lardelli M (2009) Complex splicing and neural expression of duplicated tau genes in zebrafish embryos. J Alzheimers Dis 18:305–317. doi:10.3233/JAD-2009-1145
Lieschke GJ, Currie PD (2007) Animal models of human disease: zebrafish swim into view. Nat Rev Genet 8:353–367. doi:10.1038/nrg2091
Xi Y, Noble S, Ekker M (2011) Modeling neurodegeneration in zebrafish. Curr Neurol Neurosci Rep 11:274–282. doi:10.1007/s11910-011-0182-2
Howe K, Clark MD, Torroja CF et al (2013) The zebrafish reference genome sequence and its relationship to the human genome. Nature 496:498–503. doi:10.1038/nature12111
Warnatz HJ, Schmidt D, Manke T et al (2011) The BTB and CNC homology 1 (BACH1) target genes are involved in the oxidative stress response and in control of the cell cycle. J Biol Chem 286:23521–23532. doi:10.1074/jbc.M111.220178
Okita Y, Kamoshida A, Suzuki H et al (2013) Transforming growth factor-beta induces transcription factors MafK and Bach1 to suppress expression of the heme oxygenase-1 gene. J Biol Chem 288:20658–20667. doi:10.1074/jbc.M113.450478
Kent WJ, Sugnet CW, Furey TS et al (2002) The human genome browser at UCSC. Genome Res 12:996–1006. doi:10.1101/gr.229102
Friocourt G, Kappeler C, Saillour Y et al (2005) Doublecortin interacts with the ubiquitin protease DFFRX, which associates with microtubules in neuronal processes. Mol Cell Neurosci 28:153–164. doi:10.1016/j.mcn.2004.09.005
Drewes G, Ebneth A, Preuss U et al (1997) MARK, a novel family of protein kinases that phosphorylate microtubule-associated proteins and trigger microtubule disruption. Cell 89:297–308. doi:10.1016/S0092-8674(00)80208-1
Hooper C, Killick R, Lovestone S (2008) The GSK3 hypothesis of Alzheimer’s disease. J Neurochem 104:1433–1439. doi:10.1111/j.1471-4159.2007.05194.x
Wu Z, Irizarry RA, Gentleman R et al (2004) A model-based background adjustment for oligonucleotide expression arrays. J Am Stat Assoc 99:909–917. doi:10.1198/016214504000000683
Smyth G (2004) Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:article 3. doi: 10.2202/1544–6115.1027
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300. doi:10.2307/2346101
Leek JT, Storey JD (2007) Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet 3:1724–1735. doi:10.1371/journal.pgen.0030161
Glaab E, Schneider R (2015) Comparative pathway and network analysis of brain transcriptome changes during adult aging and in Parkinson’s disease. Neurobiol Dis 74:1–13. doi:10.1016/j.nbd.2014.11.002
Ekins S, Nikolsky Y, Bugrim A et al (2007) Pathway mapping tools for analysis of high content data. Methods Mol Biol 356:319–350
Hellemans J, Mortier G, De Paepe A et al (2007) qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 8:R19. doi:10.1186/gb-2007-8-2-r19
Vandesompele J, De Preter K, Pattyn F, et al. (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:RESEARCH0034. doi: 10.1186/gb-2002-3-7-research0034
Yuan JS, Reed A, Chen F, Stewart CN (2006) Statistical analysis of real-time PCR data. BMC Bioinformatics 7:85. doi:10.1186/1471-2105-7-85
Acknowledgments
Acknowledgment is made to the Geoffrey Been Alzheimer’s Initiative, 21st Century Brain Trust and to the BrightFocus Foundation (grant no. C2014002) and the Luxembourg Fondation Wivine for sponsorship of this work. Bioinformatics analyses presented in this paper were carried out in part using the HPC facilities of the University of Luxembourg (see http://hpc.uni.lu).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Sandra Köglsberger, Maria Lorena Cordero-Maldonado and Paul Antony contributed equally to this study.
ELECTRONIC SUPPLEMENTARY MATERIAL
ESM 1
Supplementary Material (file: supplementary_material.pdf): The Supplementary Material contains tables of the differential expression statistics in the microarray transcriptomics data for DU145 cells, comparing USP9X/Y knockdown samples vs. controls, for the main genes discussed in the manuscript (Tab. S1), the zebrafish primer design (Tab. S2), the PCR program, primers and hairpin sequences for the DU145 cell culture experiments (Tab. S3, S4 and S5), and the assay IDs and catalog numbers for the TaqMan® gene expression assays (Tab. S6). Moreover, figures are provided for the molecular network analysis of the DU145 transcriptomics data (Fig. S1), the primer design scheme for the DU145 knockdown experiments (Fig. S2), and the agarose gel electrophoresis results for testing different primers for the cell culture experiments (Fig. S3) (PDF 573 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Köglsberger, S., Cordero-Maldonado, M.L., Antony, P. et al. Gender-Specific Expression of Ubiquitin-Specific Peptidase 9 Modulates Tau Expression and Phosphorylation: Possible Implications for Tauopathies. Mol Neurobiol 54, 7979–7993 (2017). https://doi.org/10.1007/s12035-016-0299-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-0299-z