Abstract
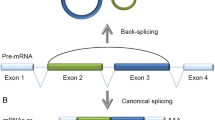
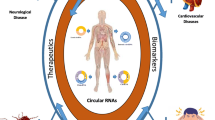
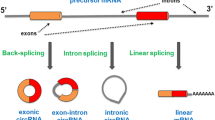
Circular RNAs (circRNAs), a type of endogenous noncoding RNAs distinct from linear forms, are produced by backsplicing events within genes. circRNAs are structurally stable, highly conserved molecules found widely in organisms, and display tissue-type and developmental-stage specific expression patterns, which reveal their significant regulatory functions in gene expression. Based on accumulating evidence, some circRNAs are now believed to be a class of competitive endogenous RNAs that regulate gene expression. For example, circRNAs may prevent microRNAs from inhibiting target RNAs acting as microRNA sponges, or interact with RNA binding proteins and thereby efficiently and post-transcriptionally regulate expression of the parental and other genes. In addition, an increasing number of studies have shown that circRNAs play important roles in the development and progression of neurological disorders. In this review, we provide a comprehensive overview on the biogenesis, characteristics, and functions of circRNAs. We also discuss the critical role of circRNAs in neurological disorders.




Similar content being viewed by others
References
Abdelmohsen K, Panda AC, Munk R, Grammatikakis I, Dudekula DB, de S, Kim J, Noh JH, Kim KM, Martindale JL, Gorospe M (2017) Identification of hur target circular RNAs uncovers suppression of pabpn1 translation by circpabpn1. RNA Biol 14(3):361–369. https://doi.org/10.1080/15476286.2017.1279788
Abe N, Hiroshima M, Maruyama H, Nakashima Y, Nakano Y, Matsuda A, Sako Y, Ito Y, Abe H (2013) Rolling circle amplification in a prokaryotic translation system using small circular RNA. Angew Chem Int Ed Engl 52(27):7004–7008. https://doi.org/10.1002/anie.201302044
Abe N, Matsumoto K, Nishihara M, Nakano Y, Shibata A, Maruyama H, Shuto S, Matsuda A, Yoshida M, Ito Y, Abe H (2015) Rolling circle translation of circular RNA in living human cells. Sci Rep 5:16435. https://doi.org/10.1038/srep16435
AbouHaidar MG, Venkataraman S, Golshani A, Liu B, Ahmad T (2014) Novel coding, translation, and gene expression of a replicating covalently closed circular RNA of 220 nt. Proc Natl Acad Sci U S A 111:14542–14547. https://doi.org/10.1073/pnas.1402814111
Armakola M, Higgins MJ, Figley MD, Barmada SJ, Scarborough EA, Diaz Z, Fang X, Shorter J, Krogan NJ, Finkbeiner S, Farese RV, Gitler AD (2012) Inhibition of RNA lariat debranching enzyme suppresses tdp-43 toxicity in ALS disease models. Nat Genet 44(12):1302–1309. https://doi.org/10.1038/ng.2434
Arnberg AC, Van Ommen GJ, Grivell LA, Van Bruggen EF, Borst P (1980) Some yeast mitochondrial RNAs are circular. Cell 19:313–319. https://doi.org/10.1016/0092-8674(80)90505-x
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S (2014) circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56:55–66. https://doi.org/10.1016/j.molcel.2014.08.019
Bahn JH, Zhang Q, Li F, Chan TM, Lin X, Kim Y, Wong DTW, Xiao X (2015) The landscape of microRNA, piwi-interacting RNA, and circular RNA in human saliva. Clin Chem 61:221–230. https://doi.org/10.1373/clinchem.2014.230433
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE (2010) Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet 6:e1001233. https://doi.org/10.1371/journal.pgen.1001233
Cao QP, Gaudette MF, Robinson DH, Crain WR (1995) Expression of the mouse testis-determining gene Sry in male preimplantation embryos. Mol Reprod Dev 40:196–204. https://doi.org/10.1002/mrd.1080400208
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, Goodfellow P, Lovell-Badge R (1993) Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73:1019–1030. https://doi.org/10.1016/0092-8674(93)90279-y
Chao CW, Chan DC, Kuo A, Leder P (1998) The mouse formin (Fmn) gene: abundant circular RNA transcripts and gene-targeted deletion analysis. Mol Med 4:614–628
Chen CY, Sarnow P (1995) Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Science 268:415–417. https://doi.org/10.1126/science.7536344
Chen LL (2016) The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol 17:205–211. https://doi.org/10.1038/nrm.2015.32
Cocquerelle C, Mascrez B, Hétuin D, Bailleul B (1993) Mis-splicing yields circular RNA molecules. FASEB J 7:155–160. https://doi.org/10.1096/fasebj.7.1.7678559
Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA, Roslan S, Schreiber AW, Gregory PA, Goodall GJ (2015) The RNA binding protein quaking regulates formation of circRNAs. Cell 160:1125–1134. https://doi.org/10.1016/j.cell.2015.02.014
Du WW, Yang W, Chen Y et al (2017) Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J 38:1402–1412. https://doi.org/10.1093/eurheartj/ehw001
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB (2016) Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res 44:2846–2858. https://doi.org/10.1093/nar/gkw027
Dubin RA, Kazmi MA, Ostrer H (1995) Inverted repeats are necessary for circularization of the mouse testis Sry transcript. Gene 167:245–248. https://doi.org/10.1016/0378-1119(95)00639-7
Ghosal S, Das S, Sen R, Basak P, Chakrabarti J (2013) Circ2Traits: a comprehensive database for circular RNA potentially associated with disease and traits. Front Genet 4:283. https://doi.org/10.3389/fgene.2013.00283
Guarnerio J, Bezzi M, Jeong JC, Paffenholz SV, Berry K, Naldini MM, Lo-Coco F, Tay Y, Beck AH, Pandolfi PP (2016) Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell 166:1055–1056
Guo JU, Agarwal V, Guo H, Bartel DP (2014) Expanded identification and characterization of mammalian circular RNAs. Genome Biol 15:409. https://doi.org/10.1186/preaccept-1176565312639289
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013b) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388. https://doi.org/10.1038/nature11993
Hansen TB, Kjems J, Damgaard CK (2013a) Circular RNA and miR-7 in cancer. Cancer Res 73:5609–5612. https://doi.org/10.1158/0008-5472.can-13-1568
Hansen TB, Wiklund ED, Bramsen JB, Villadsen SB, Statham AL, Clark SJ, Kjems J (2011) miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J 30:4414–4422. https://doi.org/10.1038/emboj.2011.359
Hentze MW, Preiss T (2013) Circular RNAs: splicing's enigma variations. EMBO J 32:923–925. https://doi.org/10.1038/emboj.2013.53
Hsu MT, Coca-Prados M (1979) Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature 280:339–340. https://doi.org/10.1038/280339a0
Ivanov A, Memczak S, Wyler E, Torti F, Porath HT, Orejuela MR, Piechotta M, Levanon EY, Landthaler M, Dieterich C, Rajewsky N (2015) Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep 10:170–177. https://doi.org/10.1016/j.celrep.2014.12.019
Jeck WR, Sharpless NE (2014) Detecting and characterizing circular RNAs. Nat Biotechnol 32:453–461. https://doi.org/10.1038/nbt.2890
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE (2013) Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19:141–157. https://doi.org/10.1261/rna.035667.112
Junn E, Lee KW, Jeong BS, Chan TW, Im JY, Mouradian MM (2009) Repression of alpha-synuclein expression and toxicity by microRNA-7. Proc Natl Acad Sci U S A 106:13052–13057. https://doi.org/10.1073/pnas.0906277106
Kefas B, Godlewski J, Comeau L, Li Y, Abounader R, Hawkinson M, Lee J, Fine H, Chiocca EA, Lawler S, Purow B (2008) microRNA-7 inhibits the epidermal growth factor receptor and the Akt pathway and is down-regulated in glioblastoma. Cancer Res 68:3566–3572. https://doi.org/10.1158/0008-5472.can-07-6639
Kelly S, Greenman C, Cook PR, Papantonis A (2015) Exon skipping is correlated with exon circularization. J Mol Biol 427:2414–2417. https://doi.org/10.1016/j.jmb.2015.02.018
Kos A, Dijkema R, Arnberg AC, van der Meide PH, Schellekens H (1986) The hepatitis delta (delta) virus possesses a circular RNA. Nature 323:558–560. https://doi.org/10.1038/323558a0
Lasda E, Parker R (2014) Circular RNAs: diversity of form and function. RNA 20:1829–1842
Lasda E, Parker R (2016) Circular RNAs co-precipitate with extracellular vesicles: a possible mechanism for circRNA clearance. PLoS One 11:e0148407. https://doi.org/10.1371/journal.pone.0148407
Laurin M, Huber J, Pelletier A, Houalla T, Park M, Fukui Y, Haibe-Kains B, Muller WJ, Cote JF (2013) Rac-specific guanine nucleotide exchange factor DOCK1 is a critical regulator of HER2-mediated breast cancer metastasis. Proc Natl Acad Sci U S A 110:7434–7439. https://doi.org/10.1073/pnas.1213050110
Legnini I, Di TG, Rossi F et al (2017) Circ-znf609 is a circular RNA that can be translated and functions in myogenesis. Mol Cell 66(1):22–37.e9
Li Z, Huang C, Bao C (2015) Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22:256–264. https://doi.org/10.1038/nsmb.2959
Liang D, Wilusz JE (2014) Short intronic repeat sequences facilitate circular RNA production. Genes Dev 28:2233–2247. https://doi.org/10.1101/gad.251926.114
Liu Q, Zhang X, Hu X, Dai L, Fu X, Zhang J, Ao Y (2016) Circular RNA related to the chondrocyte ECM regulates MMP13 expression by functioning as a MiR-136 ‘sponge’ in human cartilage degradation. Sci Rep 6:22572. https://doi.org/10.1038/srep22572
Lukiw WJ (2013) Circular RNA (circRNA) in Alzheimer's disease (AD). Front Genet 4:307. https://doi.org/10.3389/fgene.2013.00307
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338. https://doi.org/10.1038/nature11928
Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD, Kinzler KW, Vogelstein B (1991) Scrambled exons. Cell 64:607–613
Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, Shenzis S, Samson M, Dittmar G, Landthaler M, Chekulaeva M, Rajewsky N, Kadener S (2017) Translation of circrnas. Mol Cell 66(1):9–21.e7
Preußer C, Hung LH, Schneider T, Schreiner S, Hardt M, Moebus A, Santoso S, Bindereif A (2018) Selective release of circRNAs in platelet-derived extracellular vesicles. J Extracell Vesicles 7(1):1424473
Qu S, Yang X, Li X, Wang J, Gao Y, Shang R, Sun W, Dou K, Li H (2015) Circular RNA: a new star of noncoding RNAs. Cancer Lett 365:141–148
Rodríguez-Trelles F, Tarrío R, Ayala FJ (2006) Origins and evolution of spliceosomal introns. Annu Rev Genet 40:47–76. https://doi.org/10.1146/annurev.genet.40.110405.090625
Rybak-Wolf A, Stottmeister C, Glažar P, Jens M, Pino N, Giusti S, Hanan M, Behm M, Bartok O, Ashwal-Fluss R, Herzog M, Schreyer L, Papavasileiou P, Ivanov A, Öhman M, Refojo D, Kadener S, Rajewsky N (2015) Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell 58:870–885. https://doi.org/10.1016/j.molcel.2015.03.027
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO (2013) Cell-type specific features of circular RNA expression. PLoS Genet 9:e1003777. https://doi.org/10.1371/journal.pgen.1003777
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One 7:e30733. https://doi.org/10.1371/journal.pone.0030733
Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK (1976) Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A 73:3852–3856
Saydam O, Senol O, Würdinger T et al (2011) miRNA-7 attenuation in Schwannoma tumors stimulates growth by upregulating three oncogenic signaling pathways. Cancer Res 71:852–861
Seitz H (2009) Redefining microRNA targets. Curr Biol 19:870–873. https://doi.org/10.1073/pnas.73.11.3852
Shen T, Han M, Wei G, Ni T (2015) An intriguing RNA species--perspectives of circularized RNA. Protein Cell 6:871–880. https://doi.org/10.1016/j.cub.2009.03.059
Shi X, Sun M, Liu H, Yao Y, Song Y (2013) Long non-coding RNAs: a new frontier in the study of human diseases. Cancer Lett 339:159–166. https://doi.org/10.1016/j.canlet.2013.06.013
Shi Z, Chen T, Yao Q, Zheng L, Zhang Z, Wang J, Hu Z, Cui H, Han Y, Han X, Zhang K, Hong W (2017) The circular RNA ciRS-7 promotes APP and BACE1 degradation in an NF-κB-dependent manner. FEBS Jl 284(7):1096–1109. https://doi.org/10.1111/febs.14045
Song X, Zhang N, Han P, Moon BS, Lai RK, Wang K, Lu W (2016) Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res 44:e87. https://doi.org/10.1093/nar/gkw075
Suzuki H, Tsukahara T (2014) A view of pre-mRNA splicing from RNase R resistant RNAs. Int J Mol Sci 15:9331–9342. https://doi.org/10.3390/ijms15069331
Szabo L, Salzman J (2016) Detecting circular RNAs: bioinformatic and experimental challenges. Nature Rev Gene 17(11):679–692. https://doi.org/10.1038/nrg.2016.114
Thomas LF, Sætrom P (2014) Circular RNAs are depleted of polymorphisms at microRNA binding sites. Bioinformatics 30:2243–2246. https://doi.org/10.1093/bioinformatics/btu257
Wang PL, Bao Y, Yee MC, Barrett SP, Hogan GJ, Olsen MN, Dinneny JR, Brown PO, Salzman J (2014) Circular RNA is expressed across the eukaryotic tree of life. PLoS One 9:e90859
Wang Y, Wang Z (2015) Efficient backsplicing produces translatable circular mRNAs. RNA 21:172–179. https://doi.org/10.1261/rna.048272.114
Werfel S, Nothjunge S, Schwarzmayr T, Strom TM, Meitinger T, Engelhardt S (2016) Characterization of circular RNAs in human, mouse and rat hearts. J Mol Cell Cardiol 98:103–107. https://doi.org/10.1016/j.yjmcc.2016.07.007
Westholm JO, Miura P, Olson S, Shenker S, Joseph B, Sanfilippo P, Celniker SE, Graveley BR, Lai EC (2014) Genome-wide analysis of Drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep 9:1966–1980
Wu DG, Wang YY, Fan LG, Luo H, Han B, Sun LH, Wang XF, Zhang JX, Cao L, Wang XR, You YP, Liu N (2011) MicroRNA-7 regulates glioblastoma cell invasion via targeting focal adhesion kinase expression. Chin Med J 124:2616–2621
Yang Q, Du WW, Wu N et al (2017) A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ 24(9):1609–1620. https://doi.org/10.1038/cdd.2017.86
Yang W, Du WW, Li X, Yee AJ, Yang BB (2016) Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene 35:3919–3931. https://doi.org/10.1038/onc.2015.460
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F (2018) Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J Natl Cancer Inst 110(3):304–315. https://doi.org/10.1093/jnci/djx166
You X, Vlatkovic I, Babic A, Will T, Epstein I, Tushev G, Akbalik G, Wang M, Glock C, Quedenau C, Wang X, Hou J, Liu H, Sun W, Sambandan S, Chen T, Schuman EM, Chen W (2015) Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci 18:603–610. https://doi.org/10.1038/nn.3975
Yun Y, Fan X, Mao M et al (2017) Extensive translation of circular RNAs driven by n6-methyladenosine. Cell Res 27(5):626–641. https://doi.org/10.1038/cr.2017.31
Zeng Y, Du WW, Wu Y et al (2017) A circular RNA binds to and activates Akt phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics 7(16):3842–3855
Zhang M, Huang N, Yang X et al (2018) A novel protein encoded by the circular form of the shprh gene suppresses glioma tumorigenesis. Oncogene 37(13):1805–1814. https://doi.org/10.7150/thno.19764
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L (2014) Complementary sequence-mediated exon circularization. Cell 159:134–147. https://doi.org/10.1016/j.cell.2014.09.001
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, Zhu S, Yang L, Chen LL (2013) Circular intronic long noncoding RNAs. Mol Cell 51:792–806
Zhao M, Gao F, Zhang D, Wang S, Zhang Y, Wang R, Zhao J (2017) Altered expression of circular RNAs in moyamoya disease. J Neurol Sci 381:25–31. https://doi.org/10.1016/j.jns.2017.08.011
Zhao Y, Alexandrov PN, Jaber V, Lukiw WJ (2016) Deficiency in the ubiquitin conjugating enzyme UBE2A in Alzheimer's disease (AD) is linked to deficits in a natural circular miRNA-7 sponge (circRNA; ciRS-7). Genes 7:E116. https://doi.org/10.3390/genes7120116
Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G, Liang L, Gu J, He X, Huang S (2016) Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun 7:11215. https://doi.org/10.1038/ncomms11215
Acknowledgments
We would like to thank the Editage (www.editage.com) for the English language editing and Publication Support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Wang, Q., Qu, L., Chen, X. et al. Progress in Understanding the Relationship Between Circular RNAs and Neurological Disorders. J Mol Neurosci 65, 546–556 (2018). https://doi.org/10.1007/s12031-018-1125-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-018-1125-z