Abstract
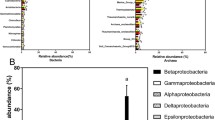
Microbial planktonic communities are critical components of marine biogeochemical pathways. Despite this, there is still limited knowledge on the dynamics of this group in warm and oligotrophic waters. We used high-throughput sequencing to characterise the bacterial (16S rRNA) and eukaryotic (18S rRNA) microbial plankton communities in two regions under the influence of anthropogenic impacts (a port and sewage outflow) and a coastal region with no direct anthropogenic disturbances in the central Red Sea. Overall, bacterial and eukaryotic components responded in a similar way to the environmental conditions. Community composition and structure were more sensitive than alpha diversity measures to environmental impacts. With the exception of eukaryotes, for which the number of OTU differed significantly between sampling periods in all the regions, environmental changes associated with anthropogenic pressures seem to be better reflected by variations in the relative dominance of microbial groups. For example, elevated proportional abundances of nitrifying and sewage-/faecal-related bacteria at the impacted sites were observed compared with the coastal region. The recently developed microgAMBI also appeared to correlate well with the level of anthropogenic impact the regions experienced, showing the potential to be applied in oligotrophic waters.








Similar content being viewed by others
References
Abu-Zied RH, Al-Dubai TAM, Bantan RA (2016) Environmental conditions of shallow waters alongside the souther corniche of Jeddah based on benthic foraminifera, physio-chemical parameters and heavy metals. J Foraminiferal Research 46 (2):149–170
Al-Farawati R (2010) Environmental conditions of the coastal waters of Southern Corinche Jeddah, Eastern red sea: Physico-chemical approach. Aust J Basic Appl Sci 4:3324–3337
Anderson M, Gorley RN, Clarke RK (2008) Permanova+ for primer: guide to software and statistical methods. Primer-E Limited,
Aylagas E, Borja Á, Tangherlini M, Dell'Anno A, Corinaldesi C, Michell CT, Irigoien X, Danovaro R, Rodríguez-Ezpeleta N (2017) A bacterial community-based index to assess the ecological status of estuarine and coastal environments. Mar Pollut Bull 114:679–688
Basaham A, Rifaat A, El-Mamoney M, El Sayed M (2009) Re-evaluation of the impact of sewage disposal on coastal sediments of the Southern Corniche, Jeddah, Saudi Arabia. Journal of King Abdulaziz University, Marine Sciences 20:109–126
Borja A (2018) Testing the efficiency of a bacterial community-based index (microgAMBI) to assess distinct impact sources in six locations around the world. Ecol Indic 85:594–602
Borja A, Franco J, Perez V (2000) A Marine Biotic Index to establish the ecological quality of soft-bottom benthos within European estuarine and coastal environments. Mar Pollut Bull 40:1100–1114
Caporaso JG et al (2010) QIIME allows analysis of high throughput community sequencing data. Nat Methods 7:335–336
Carugati L, Corinaldesi C, Dell'Anno A, Danovaro R (2015) Metagenetic tools for the census of marine meiofaunal biodiversity: An overview. Mar Genomics 24:11–20
Caruso G, la Ferla R, Azzaro M, Zoppini A, Marino G, Petochi T, Corinaldesi C, Leonardi M, Zaccone R, Fonda Umani S, Caroppo C, Monticelli L, Azzaro F, Decembrini F, Maimone G, Cavallo RA, Stabili L, Hristova Todorova N, K. Karamfilov V, Rastelli E, Cappello S, Acquaviva MI, Narracci M, de Angelis R, del Negro P, Latini M, Danovaro R (2016) Microbial assemblages for environmental quality assessment: knowledge, gaps and usefulness in the European Marine Strategy Framework Directive. Crit Rev Microbiol 42:883–904. https://doi.org/10.3109/1040841X.2015.1087380
Carvalho S, Barata M, Pereira F, Gaspar MB, da Fonseca LC, Pousão-Ferreira P (2006) Distribution patterns of macrobenthic species in relation to organic enrichment within aquaculture earthen ponds. Mar Pollut Bull 52:1573–1584
Clarke KR, Gorley RN (2006) PRIMER V6: user manual-tutorial. Plymouth Marine Laboratory
Closek CJ, Sunagawa S, DeSalvo MK, Piceno YM, DeSantis TZ, Brodie EL, Weber MX, Voolstra CR, Andersen GL, Medina M (2014) Coral transcriptome and bacterial community profiles reveal distinct Yellow Band Disease states in Orbicella faveolata. The ISME journal 8:2411–2422
de Vargas C, Audic S, Henry N, Decelle J, Mahe F, Logares R, Lara E, Berney C, le Bescot N, Probert I, Carmichael M, Poulain J, Romac S, Colin S, Aury JM, Bittner L, Chaffron S, Dunthorn M, Engelen S, Flegontova O, Guidi L, Horak A, Jaillon O, Lima-Mendez G, Luke J, Malviya S, Morard R, Mulot M, Scalco E, Siano R, Vincent F, Zingone A, Dimier C, Picheral M, Searson S, Kandels-Lewis S, Tara Oceans Coordinators, Acinas SG, Bork P, Bowler C, Gorsky G, Grimsley N, Hingamp P, Iudicone D, Not F, Ogata H, Pesant S, Raes J, Sieracki ME, Speich S, Stemmann L, Sunagawa S, Weissenbach J, Wincker P, Karsenti E, Boss E, Follows M, Karp-Boss L, Krzic U, Reynaud EG, Sardet C, Sullivan MB, Velayoudon D (2015) Eukaryotic plankton diversity in the sunlit ocean. Science 348:1261605. https://doi.org/10.1126/science.1261605
Demers S, Therriault JC, Bourget E, Bah A (1987) Resuspension in the shallow sublittoral zone of a macrotidal estuarine environment: wind influence. Limnol Oceanogr 32:327–339
Done TJ (1992) Phase shifts in coral reef communities and their ecological significance. Hydrobiologia 247:121–132
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
El-Rayis OA, Moammar MO (1998) Environmental conditions of two Red Sea coastal lagoons in Jeddah: 1. Hydrochemistry. Journal King Abdulaziz University Earth Science 9:31–47
Estrada M (1991) Phytoplankton assemblages across a NW Mediterranean front: changes from winter mixing to spring stratification. Oecologia aquatica 10:157–185
Estrada M et al. (2016) Phytoplankton across tropical and subtropical regions of the Atlantic, Indian and Pacific oceans PLOS ONE 11 https://doi.org/10.1371/journal10.1371/journal.pone.0151699
Ferreira JG, Andersen JH, Borja A, Bricker SB, Camp J, Cardoso da Silva M, Garcés E, Heiskanen AS, Humborg C, Ignatiades L, Lancelot C, Menesguen A, Tett P, Hoepffner N, Claussen U (2011) Overview of eutrophication indicators to assess environmental status within the European Marine Strategy Framework Directive. Estuar Coast Shelf Sci 93:117–131. https://doi.org/10.1016/j.ecss.2011.03.014
Findlay HS (2006) Modelling of autumn plankton bloom dynamics. J Plankton Res 28:209–220. https://doi.org/10.1093/plankt/fbi114
Frouin P (2000) Effects of anthropogenic disturbances of tropical soft-bottom benthic communities. Mar Ecol Prog Ser 194:39–53
Fuhrman JA, Cram JA, Needham DM (2015) Marine microbial community dynamics and their ecological interpretation. Nat Rev Microbiol 13:133–146
Furnas M, Mitchell A, Skuza M, Brodie J (2005) In the other 90%: phytoplankton responses to enhanced nutrient availability in the Great Barrier Reef Lagoon. Mar Pollut Bull 51:253–265. https://doi.org/10.1016/j.marpolbul.2004.11.010
Gilbert JA, Steele JA, Caporaso JG, Steinbrück L, Reeder J, Temperton B, Huse S, McHardy AC, Knight R, Joint I, Somerfield P, Fuhrman JA, Field D (2012) Defining seasonal marine microbial community dynamics. ISME J 6:298–308. https://doi.org/10.1038/ismej.2011.107
Glibert P (2016) Margalef revisited: a new phytoplankton mandala incorporating twelve dimensions, including nutritional physiology. Harmful Algae 55:25–30
Guillou L, Bachar D, Audic S, Bass D, Berney C, Bittner L, Boutte C, Burgaud G, de Vargas C, Decelle J, del Campo J, Dolan JR, Dunthorn M, Edvardsen B, Holzmann M, Kooistra WHCF, Lara E, le Bescot N, Logares R, Mahé F, Massana R, Montresor M, Morard R, Not F, Pawlowski J, Probert I, Sauvadet AL, Siano R, Stoeck T, Vaulot D, Zimmermann P, Christen R (2012) The protist ribosomal reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res 41:D597–D604
Hargrave B, Phillips G, Doucette L, White M, Milligan T, Wildish D, Cranston R (1997) Assessing benthic impacts of organic enrichment from marine aquaculture. In: The interactions between sediments and water. Springer, pp 641–650
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Gloeckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies Nucleic Acids Res 41 https://doi.org/10.1093/nar/gks808
Lallias D, Hiddink JG, Fonseca VG, Gaspar JM, Sung W, Neill SP, Barnes N, Ferrero T, Hall N, Lambshead PJD, Packer M, Thomas WK, Creer S (2015) Environmental metabarcoding reveals heterogeneous drivers of microbial eukaryote diversity in contrasting estuarine ecosystems. The ISME journal 9:1208–1221
Lapointe BE, Barile PJ, Matzie WR (2004) Anthropogenic nutrient enrichment of seagrass and coral reef communities in the Lower Florida Keys: discrimination of local versus regional nitrogen sources. J Exp Mar Biol Ecol 308:23–58
Lugoli F, Garmendia M, Lehtinen S, Kauppila P, Moncheva S, Revilla M, Roselli L, Slabakova N, Valencia V, Dromph KM, Basset A (2012) Application of a new multi-metric phytoplankton index to the assessment of ecological status in marine and transitional waters. Ecol Indic 23:338–355
Malham SK, Rajko-Nenow P, Howlett E, Tuson KE, Perkins TL, Pallett DW, Wang H, Jago CF, Jones DL, McDonald JE (2014) The interaction of human microbial pathogens, particulate material and nutrients in estuarine environments and their impacts on recreational and shellfish waters. Environ Sci Process Impacts 16:2145–2155. https://doi.org/10.1039/c4em00031e
Massana R, Gobet A, Audic S, Bass D, Bittner L, Boutte C, Chambouvet A, Christen R, Claverie JM, Decelle J, Dolan JR, Dunthorn M, Edvardsen B, Forn I, Forster D, Guillou L, Jaillon O, Kooistra WHCF, Logares R, Mahé F, Not F, Ogata H, Pawlowski J, Pernice MC, Probert I, Romac S, Richards T, Santini S, Shalchian-Tabrizi K, Siano R, Simon N, Stoeck T, Vaulot D, Zingone A, de Vargas C (2015) Marine protist diversity in European coastal waters and sediments as revealed by high-throughput sequencing. Environ Microbiol 17:4035–4049. https://doi.org/10.1111/1462-2920.12955
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Newton RJ, Bootsma MJ, Morrison HG, Sogin ML, McLellan SL (2013) A microbial signature approach to identify fecal pollution in the waters off an urbanized coast of Lake Michigan. Microb Ecol 65:1011–1023
Nogales B, Lanfranconi MP, Pina-Villalonga JM, Bosch R (2011) Anthropogenic perturbations in marine microbial communities. FEMS Microbiol Rev 35:275–298. https://doi.org/10.1111/j.1574-6976.2010.00248.x
Park H-D, Lee S-Y, Hwang S (2009) Redundancy analysis demonstration of the relevance of temperature to ammonia-oxidizing bacterial community compositions in a full-scale nitrifying bioreactor treating saline wastewater. J Microbiol Biotechnol 19:346–350. https://doi.org/10.4014/jmb.0806.399
Pastorok RA, Bilyard GR (1985) Effects of sewage pollution on coral-reef communities. Mar Ecol Prog Ser 21(1):175–189
Pearman J, Kurten S, Sarma YVB, Jones B, Carvalho S (2016) Biodiversity patterns of plankton assemblages at the extremes of the Red Sea FEMS Microbiol Ecol 92 https://doi.org/10.1093/femsec/fiw002
Pearman JK, Ellis J, Irigoien X, Sarma Y, Jones BH, Carvalho S (2017) Microbial planktonic communities in the Red Sea: high levels of spatial and temporal variability shaped by nutrient availability and turbulence. Sci Rep 7:6611
Peña-García D, Ladwig N, Turki AJ, Mudarris MS (2014) Input and dispersion of nutrients from the Jeddah Metropolitan Area, Red Sea. Mar Pollut Bull 80:41–51
Piló D, Pereira F, Carriço A, Curdia J, Pereira P, Gaspar M, Carvalho S (2015) Temporal variability of biodiversity patterns and trophic structure of estuarine macrobenthic assemblages along a gradient of metal contamination. Estuar Coast Shelf Sci 167:286–299
Primpas I, Karydis M (2011) Scaling the trophic index (TRIX) in oligotrophic marine environments. Environ Monit Assess 178:257–269
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Gloeckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196. https://doi.org/10.1093/nar/gkm864
Sanchez O, Ferrera I, Gonzalez JM, Mas J (2013) Assessing bacterial diversity in a seawater-processing wastewater treatment plant by 454-pyrosequencing of the 16S rRNA and amoA genes. Microb Biotechnol 6:435–442. https://doi.org/10.1111/1751-7915.12052
Santo Domingo JW, Rice EW (2009) Detection of microbial water quality indicators and fecal waterborne pathogens in environmental waters: a review of methods, Appl Limitations https://doi.org/10.1002/9780470027318.a9053
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/aem.01541-09
Simonato F, Gomez-Pereira PR, Fuchs BM, Amann R (2010) Bacterioplankton diversity and community composition in the Southern Lagoon of Venice. Syst Appl Microbiol 33:128–138. https://doi.org/10.1016/j.syapm.2009.12.006
Stoeck T, Bass D, Nebel M, Christen R, Jones MD, Breiner HW, Richards TA (2010) Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol Ecol 19(Suppl 1):21–31. https://doi.org/10.1111/j.1365-294X.2009.04480.x
Strong JA, Andonegi E, Bizsel KC, Danovaro R, Elliott M, Franco A, Garces E, Little S, Mazik K, Moncheva S, Papadopoulou N, Patrício J, Queirós AM, Smith C, Stefanova K, Solaun O (2015) Marine biodiversity and ecosystem function relationships: the potential for practical monitoring applications. Estuar Coast Shelf Sci 161:46–64
Team RC (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, p 2014
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wear SL, Thurber RV (2015) Sewage pollution: mitigation is key for coral reef stewardship. Ann N Y Acad Sci 1355:15–30
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer,
Yu K, Zhang T (2012) Metagenomic and metatranscriptomic analysis of microbial community structure and gene expression of activated sludge. PLoS One 7:e38183. https://doi.org/10.1371/journal.pone.0038183
Zalack JT, Smucker NJ, Vis ML (2010) Development of a diatom index of biotic integrity for acid mine drainage impacted streams. Ecol Indic 10:287–295
Zhang R, Liu B, Lau SC, Ki JS, Qian PY (2007) Particle-attached and free-living bacterial communities in a contrasting marine environment: Victoria Harbor, Hong Kong. FEMS Microbiol Ecol 61:496–508. https://doi.org/10.1111/j.1574-6941.2007.00353.x
Ziegler M, Roik A, Porter A, Zubier K, Mudarris MS, Ormond R, Voolstra CR (2016) Coral microbial community dynamics in response to anthropogenic impacts near a major city in the central Red Sea. Mar Pollut Bull 105:629–640
Acknowledgements
The authors would like to thank Richard Payumo, Saskia Kürten and various members of the Coastal & Marine Resources Core Lab for their help in collecting the samples. Further, the authors would like to express their thanks to Saskia Kürten and Pedro Ruiz Compean for their help with the nutrient analysis. The authors would also like to extend their gratitude to Ute Langner for her work in producing the maps used in this manuscript. We would also like to express our gratitude to the reviewers for their insightful comments, which improved this manuscript.
Funding
JKP and SC are funded by the Saudi Aramco/KAUST Center for Marine Environmental Observations. The funding source had no involvement in the design, undertaking or analysis of this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Rights and permissions
About this article
Cite this article
Pearman, J.K., Afandi, F., Hong, P. et al. Plankton community assessment in anthropogenic-impacted oligotrophic coastal regions. Environ Sci Pollut Res 25, 31017–31030 (2018). https://doi.org/10.1007/s11356-018-3072-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-018-3072-1