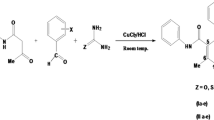
Abstract
On screening of endolithic actinobacteria from a granite rock sample of Meghalaya for antibacterial compound, a novel antibacterial compound CCp1 was isolated from the fermentation broth of Actinomadura sp. AL2. On purification of the compound based on chromatographic techniques followed by characterization with FT-IR, UV–visible, 1H NMR, 13C NMR and mass spectrometry, the molecular formula of the compound was generated as C20H17N3O2, a furopyrimidine derivative. In vitro antibacterial activity of the compound was evaluated against both Gram positive and negative bacteria by agar well diffusion assay. The compound had lowest MIC (2.00 µg/ml) for Bacillus subtilis and highest MIC (> 64 µg/ml) for Staphylococcus epidermidis and Pseudomonas aeruginosa. The study revealed that the compound has potential antibacterial activity. The mode of action of the antibacterial compound was evaluated through in silico studies for its ability to bind DNA gyrase, 30S RNA molecules, OmpF porins and N-Acetylglucosamine-1-phosphate uridyltransferase (GlmU). The antibacterial compound demonstrated more favorable docking with DNA gyrase, 30S RNA molecules and OmpF porins than GlmU which support the antibacterial compound CCp1 can be as a promising broad spectrum antibiotic agent with “multitarget” characteristics.
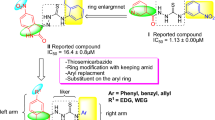
Graphical Abstract








Similar content being viewed by others
References
Abdulla H (2009) Bioweathering and biotransformation of granitic rock minerals by actinomycetes. Microb Ecol 58(4):753–761. doi:10.1007/s00248-009-9549-1
Andrusier N, Nussinov R, Wolfson HJ (2007) FireDock: fast interaction refinement in molecular docking. Proteins 69(1):139–159. doi:10.1002/prot.21495
Bajpai VK, Kang SC (2011) Isolation and characterization of biologically active secondary metabolites from Metasequoia glyptostroboides Miki ex Hu. J Food Safety 31(2):276–283. doi:10.1111/j.1745-4565.2011.00298.x
Baker D, Mocek U, Garr C (2000) Natural products versus combinatorials: a case study. In: Wrigley SA, Hayes MA, Thomas R, Chrystal EJT, Nicholson N (eds) New leads for pharmaceutical and agrochemical industries. The royal society of chemistry, Cambridge, pp 66–72
Balachandran C, Duraipandiyan V, Emi N, Ignacimuthu S (2015) Antimicrobial and cytotoxic properties of Streptomyces sp. (ERINLG-51) Isolated from Southern Western Ghats. South Ind J Biol Sci 1:7–14. doi:10.22205/sijbs/2015/v1/i1/100436
Bax BD, Chan PF, Eggleston DS, Fosberry A, Gentry DR et al (2010) Type IIA topoisomerase inhibition by a new class of antibacterial agents. Nature 466(7309):935–940. doi:10.1038/nature09197
Berdy J (2005) Bioactive microbial metabolites. J Antibiot 58:1–26. doi:10.1038/ja.2005.1
Berdy J (2012) Thoughts and facts about antibiotics: where we are now and where we are heading. J Antibiot 65:385–395. doi:10.1038/ja.2012.27
Bhattacharjee K, Joshi SR (2016) A selective medium for recovery and enumeration of endolithic bacteria. J Microbiol Methods 129:44–54. doi:10.1016/j.mimet.2016.07.026
Bhattacharjee K, Banerjee S, Joshi SR (2012) Diversity of Streptomyces spp. in Eastern Himalayan region-computational RNomics approach to phylogeny. Bioinformation 8(12):548–554. doi:10.6026/97320630008548
Bhattacharjee K, Banerjee S, Bawitlung L, Krishnappa D, Joshi SR (2014) A study on parameters optimization for degradation of endosulfan by bacterial consortia isolated from contaminated soil. Proc Natl Acad Sci India Sect B Biol Sci 84(3):657–667. doi:10.1007/s40011-013-0223-5
Brodersen DE, Clemons WM Jr, Carter AP, Morgan-Warren RJ, Wimberly BT, Ramakrishnan V (2000) The structural basis for the action of the antibiotics tetracycline, pactamycin, and hygromycin B on the 30S ribosomal subunit. Cell 103(7):1143–1154. doi:10.1016/S0092-8674(00)00216-6
Buurman ET, Andrews B, Gao N, Hu J, Keating TA et al (2011) In vitro validation of acetyltransferase activity of GlmU as an antibacterial target in Haemophilus influenzae. J Biol Chem 286(47):40734–40742. doi:10.1074/jbc.M111.274068
Cao C, Jiang J, Sun H, Huang Y, Tao F, Lian B (2016) Carbonate mineral formation under the influence of limestone-colonizing actinobacteria: morphology and polymorphism. Front Microbiol 7:366. doi:10.3389/fmicb.2016.00366
Ceccarelli M, Danelon C, Laio A, Parrinello M (2004) Microscopic mechanism of antibiotics translocation through a porin. Biophys J 87(1):58–64. doi:10.1529/biophysj.103.037283
Choma IM, Grzelak EM (2011) Bioautography detection in thin-layer chromatography. J Chromatogr A 1218(19):2684–2691. doi:10.1016/j.chroma.2010.12.069
Coates J (2000) Interpretation of infrared spectra, a practical approach. In: Meyers RA (ed) Encyclopedia of analytical chemistry, Wiley, New York, pp 1–22
Cockell CS, Calsteren PV, Mosselmans JFW, Franchi IA, Gilmour I et al (2010) Microbial endolithic colonization and the geochemical environment in young seafloor basalts. Chem Geol 279:17–30. doi:10.1016/j.chemgeo.2010.09.015
Coldham NG, Webber M, Woodward MJ, Piddock LJ (2010) A 96-well plate fluorescence assay for assessment of cellular permeability and active efflux in Salmonella enterica serovar Typhimurium and Escherichia coli. J Antimicrob Chemother 65(8):1655–1663. doi:10.1093/jac/dkq169
Collin F, Karkare S, Maxwell A (2011) Exploiting bacterial DNA gyrase as a drug target: current state and perspectives. Appl Microbiol Biotechnol 92(3):479–497. doi:10.1007/s00253-011-3557-z
Cragg GM, Newman DJ (2013) Natural products: a continuing source of novel drug leads. Biochim Biophys Acta 1830:3670–3695. doi:10.1016/j.bbagen.2013.02.008
Cui J, Hu C, Yang Y, Wu Y, Yang L et al (2012) Facile fabrication of carbonaceous nanospheres loaded with silver nanoparticles as antibacterial materials. J Mater Chem 22:8121–8126. doi:10.1039/C2JM16441H
Demain AL (2006) From natural products discovery to commercialization: a success story. J Ind Microbiol Biotechnol 33(7):486–495. doi:10.1007/s10295-005-0076-x
Dias DA, Urban S, Roessner U (2012) A historical overview of natural products in drug discovery. Metabolites 2:303–336. doi:10.3390/metabo2020303
Dong H, Rech JA, Jiang H, Sun H, Buck BJ (2007) Endolithic cyanobacteria in soil gypsum: occurrences in Atacama (Chile), Mojave (United States), and Al-Jafr Basin (Jordan) Deserts. J Geophys Res 112:G02030. doi:10.1029/2006JG000385
Duhovny D, Nussinov R, Wolfson HJ (2002) Efficient Unbound Docking of Rigid Molecules. In: Gusfield D et al. (ed) Proceedings of the 2’nd Workshop on Algorithms in Bioinformatics(WABI) Rome, Italy, Lecture Notes in Computer Science 2452. Springer Verlag, pp 185–200
Duraipandiyan V, Al-Dhabi NA, Ignacimuthu S (2016) New antimicrobial anthraquinone 6,61-bis (1,5,7-trihydroxy-3 hydroxymethylanthraquinone) isolated from Streptomyces sp. isolate ERI-26. Saudi J Biol Sci 23(6):731–735. doi:10.1016/j.sjbs.2016.02.008
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17(6):368–376. doi:10.1007/BF01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evol Int J Org Evol 39(4):783–791. doi:10.2307/2408678
Groth I, Vettermann R, Schuetze B, Schumann P, Saiz-Jimenez C (1999) Actinomycetes in Karstic caves of northern Spain (Altamira and Tito Bustillo). J Microbiol Methods 36(1–2):115–122. doi:10.1016/S0167-7012(99)00016-0
Groth I, Tan GY, González JM, Laiz L, Carlsohn MR et al (2007) Amycolatopsis nigrescens sp. nov., an actinomycete isolated from a Roman catacomb. Int J Syst Evol Microbiol 57(Pt 3):513–519. doi:10.1099/ijs.0.64602-0
Hazra S, Saha P, Ray J, Podder A (2010) Simple statistical and mineralogical studies as petrogenetic indicator for Neoproterozoic Mylliem porphyritic granites of East Khasi Hills, Meghalaya, Northeastern India. J Geol Soc India 75:760–768. doi:10.1007/s12594-010-0061-5
Holt JG (1994) Bergey’s manual of determinative bacteriology, 9th edn. Lippincott Williams and Wilkins, Baltimore
Hong K, Gao AH, Xie QY, Gao H, Zhuang L et al (2009) Actinomycetes for marine drug discovery isolated from mangrove soils and plants in China. Mar Drugs 7(1):24–44. doi:10.3390/md7010024
Hong W, Zeng J, Xie J (2014) Antibiotic drugs targeting bacterial RNAs. Acta Pharm Sin B 4(4):258–265. doi:10.1016/j.apsb.2014.06.012
Hossain MI, Bhuiyan MMH (2009) Synthesis and antimicrobial activities of some new thieno and furopyrimidine derivatives. J Sci Res 1:317–325. doi:10.3329/jsr.v1i2.2299
Igarashi Y, Iida T, Oku N, Watanabe H, Furihata K, Miyanouchi K (2012) Nomimicin, a new spirotetronate-class polyketide from an actinomycete of the genus Actinomadura. J Antibiot 65:355–359. doi:10.1038/ja.2012.30
Imamura K, Odagawa A, Tanabe K, Hayakawa Y, Otake N (1984) Akrobomycin, a new anthracycline antibiotic. J Antibiot 37:83–84. doi:10.7164/antibiotics.37.83
Ito Y (2005) Golden rules and pitfalls in selecting optimum conditions for high-speed counter current chromatography. J Chromatogr A 1065(2):145–168. doi:10.1016/j.chroma.2004.12.044
Karuppiah V, Li Y, Sun W, Feng G, Li Z (2015) Functional gene-based discovery of phenazines from the actinobacteria associated with marine sponges in the South China Sea. Appl Microbiol Biotechnol 99(14):5939–5950. doi:10.1007/s00253-015-6547-8
Katritzky AR, Ji Y, Fang Y, Prakash I (2001) Novel syntheses of 2,3-disubstituted Benzofurans. J Org Chem 66:5613–5615. doi:10.1021/jo010278p
Katrusiak A, Katrusiak A, Bałoniak S (1994) Reactivity of 6-chloro-4- and 5-hydrazino-2-phenyl-3(2H)-pyridazinones with Vilsmeier reagent. Tetrahedron 50(45):12933–12940. doi:10.1016/S0040-4020(01)81212-6
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical Streptomyces genetics. John Innes Foundation, Norwich
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16(2):111–120
Kimura T, Iwatsuki M, Asami Y, Ishiyama A, Hokari R et al (2016) Anti-trypanosomal compound, sagamilactam, a new polyene macrocyclic lactam from Actinomadura sp. K13-0306. J Antibiot 69(11):818–824. doi:10.1038/ja.2016.28
Kohanski MA, Dwyer DJ, Collins JJ (2010) How antibiotics kill bacteria: from targets to networks. Nat Rev Microbiol 8(6):423–435. doi:10.1038/nrmicro2333
Kotoda N, Shin-Ya K, Furihata K, Hayakawa Y, Seto H (1997) Isolation and structure elucidation of novel neuronal cell protecting substances, carbazomadurins A and B produced by Actinomadura madurae. J Antibiot 50:770–772. doi:10.7164/antibiotics.50.770
Kwon Y, Kim SH, Shin Y, Bae M, Kim BY, Lee SK et al (2014) A new benzofuran glycoside and indole alkaloids from a sponge-associated rare actinomycete, Amycolatopsis sp. Mar Drugs 12(4):2326–2340. doi:10.3390/md12042326
Landwehr W, Wolf C, Wink J (2016) Actinobacteria and myxobacteria - two of the most important bacterial resources for novel antibiotics. In: Stadler M, Dersch P (eds) How to overcome the antibiotic crisis, Springer, New York, pp 273–302
Lapchinskaya OA, Saburova TA, Ponomarenko VI, 1975. Production of carminomycin by auxotrophic mutants of Actinomadura carminata. J Basic Microbiol 15(5):383–385. doi:10.1002/jobm.19750150512
Lee DW, Lee SD (2010) Actinomadura scrupuli sp. nov., isolated from rock. Int J Syst Evol Microbiol 60:2647–2651. doi:10.1099/ijs.0.017608-0
Lee JY, Moon SS, Hwang BK (2003) Isolation and antifungal and antioomycete activities of aerugine produced by Pseudomonas fluorescens strain MM-B16. Appl Environ Microbiol 69(4):2023–2031. doi:10.1128/AEM.69.4.2023-2031.2003
Li S, Shi Y, Zhang Q, Liao X, Zhu L, Lou K (2013) Phylogenetic diversity of endolithic bacteria in Bole granite rock in Xinjiang. Acta Ecol Sin 33(4):178–184. doi:10.1016/j.chnaes.2013.05.003
Lima-Filho JVM, Carvalho AFFU, Freitas SM, Melo VMM (2002) Antibacterial activity of extracts of six macroalgae from the north eastern Brasilian coast. Braz J Microbiol 33:311–313. doi:10.1590/S1517-83822002000400006
Mashiach E, Schneidman-Duhovny D, Andrusier N, Nussinov R, Wolfson HJ (2008) FireDock: a web server for fast interaction refinement in molecular docking. Nucleic Acids Res 36(Web Server Issue):W229–W232. doi:10.1093/nar/gkn186
Masuda M, Abe T, Sato S, Suzuki T, Suzuki M (1997) Diversity of halogenated secondary metabolites in the red alga Laurencia nipponica (Rhodomelaceae Ceramiales). J Phycol 33(2):196–208. doi:10.1111/j.0022-3646.1997.00196.x
McGuigan C, Brancale A, Barucki H, Srinivasan S, Jones G et al (2001) Furano pyrimidines as novel potent and selective anti-VZV agents. Antivir Chem Chemother 12(2):77–89. doi:10.1177/095632020101200201
McNamara CJ, Perry TD 4th, Bearce KA, Hernandez-Duque G, Mitchell R (2006) Epilithic and endolithic bacterial communities in limestone from a Maya archaeological site. Microb Ecol 51(1):51–64. doi:10.1007/s00248-005-0200-5
Mehra R, Rani C, Mahajan P, Vishwakarma RA, Khan IA, Nargotra A (2016) Computationally guided identification of novel Mycobacterium tuberculosis GlmU inhibitory leads, their optimization, and in vitro validation. ACS Comb Sci 18(2):100–116. doi:10.1021/acscombsci.5b00019
Michener CD, Sokal RR (1957) A quantitative approach to a problem in classification. Evol Int J Org Evol 11(2):130–162. doi:10.2307/2406046
Miyadoh S (1993) Research on antibiotic screening in Japan over the last decade: a producing microorganisms approach. Actinomycetologica 9:100–106. doi:10.3209/saj.7_100
Mochalkin I, Lightle S, Zhu Y, Ohren JF, Spessard C et al (2007) Characterization of substrate binding and catalysis in the potential antibacterial target N-acetylglucosamine-1-phosphate uridyltransferase (GlmU). Protein Sci 16(12):2657–2666. doi:10.1110/ps.073135107
Olsson-Francis K, de la Torre R, Cockell CS (2010) Isolation of novel extreme-tolerant cyanobacteria from a rock-dwelling microbial community by using exposure to low earth orbit. Appl Environ Microbiol 76(7):2115–2121. doi:10.1128/AEM.02547-09
Palepu NR, Nongbri SL, Premkumar JR, Verma AK, Bhattacharjee K et al (2015) Synthesis and evaluation of new salicylaldehyde-2-picolinylhydrazone Schiff base compounds of Ru(II), Rh(III) and Ir(III) as in vitro antitumor, antibacterial and fluorescence imaging agents. J Biol Inorg Chem 20(4):619–638. doi:10.1007/s00775-015-1249-3
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM et al (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. doi:10.1002/jcc.20084
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425. doi:10.1093/oxfordjournals.molbev.a040454
Sayed GH, Sayed MA, Mahmoud MR, Shaaban SS (2002) Synthesis and reactions of new pyridazinone derivatives of expected antimicrobial activities. Egypt J Chem 45:767–776
Schmitz A, Felder S, Hover T, Kehraus S, Neu E, Lohr F, Konig GM, Schaberle TF (2013) Antibiotics from gliding bacteria. Phytochem Rev 12:507. doi:10.1007/s11101-012-9224-x
Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson HJ (2005) PatchDock and SymmDock: servers for rigid and symmetric docking. Nucl Acids Res 33:W363-367. doi:10.1093/nar/gki481
Selvin J, Shanmughapriya S, Gandhimathi R, SeghalKiran G, RajeethaRavji T et al (2009) Optimization and production of novel antimicrobial agents from sponge associated marine actinomycetes Nocardiopsis dassonvillei MAD08. Appl Microbiol Biotechnol 83(3):435–445. doi:10.1007/s00253-009-1878-y
Shirling EB, Gottlieb D (1966) Methods for Characterization of Streptomyces Species. Int J Syst Evol Microbiol 16:313–340. doi:10.1099/00207713-16-3-313
Silver LL (2011) Challenges of antibacterial discovery. Clin Microbiol Rev 24(1):71–109. doi:10.1128/CMR.00030-10
Strauss DG, Baum M, Fleck WF (1986) Butylmaduramycin, a new antibiotic from Actinomadura rubra. J Basic Microbiol 26(3):169–172. doi:10.1002/jobm.3620260307
Tiwari K, Gupta RK (2012) Rare actinobacteria: a potential storehouse for novel antibiotics. Crit Rev Biotechnol 32(2):108–132. doi:10.3109/07388551.2011.562482
Vaca I, Casqueiro J, Ullán RV, Rumbero A, Chávez R, Martín JF (2011) A preparative method for the purification of isopenicillin N from genetically blocked Acremonium chrysogenum strain TD189: studies on the degradation kinetics and storage conditions. J Antibiot 64:447–451. doi:10.1038/ja.2011.30
Vicens Q, Westhof E (2001) Crystal structure of paromomycin docked into the eubacterial ribosomal decoding A site. Structure 9(8):647–658. doi:10.1016/S0969-2126(01)00629-3
Vicens Q, Westhof E (2002) Crystal structure of a complex between the aminoglycoside tobramycin and an oligonucleotide containing the ribosomal decoding a site. Chem Biol 9(6):747–755. doi:10.1016/S1074-5521(02)00153-9
Vicens Q, Westhof E (2003) Crystal structure of geneticin bound to a bacterial 16S ribosomal RNA A site oligonucleotide. J Mol Biol 326(4):1175–1188. doi:10.1016/S0022-2836(02)01435-3
Vithani N, Bais V, Prakash B (2014) GlmU (N-acetylglucosamine-1-phosphate uridyltransferase) bound to three magnesium ions and ATP at the active site. Acta Crystallogr F Struct Biol Commun 70(Pt 6):703–708. doi:10.1107/S2053230X14008279
Waitz JA, Horan AC, Kalyanpur M, Lee BK, Loebenberg D et al (1981) Kijanimicin (Sch 25663), a novel antibiotic produced by Actinomadura kijaniata SCC 1256, Fermentation, isolation, characterization and biological properties. J Antibiot 34(9):1101–1106. doi:10.7164/antibiotics.34.1101
Walker JJ, Pace NR (2007) Endolithic microbial ecosystems. Annu Rev Microbiol 61:331–347. doi:10.1146/annurev.micro.61.080706.093302
Walker JJ, Spear JR, Pace NR (2005) Geobiology of a microbial endolithic community in the Yellowstone geothermal environment. Nature 434:1011–1014. doi:10.1038/nature03447
Warren-Rhodes KA, Rhodes KL, Boyle LN, Pointing SB, Chen Y et al (2007) Lithic cyanobacterial ecology across environmental gradients and spatial scales in China’s hot and cold deserts. FEMS Microbiol Ecol 61(3):470–482. doi:10.1111/j.1574-6941.2007.00351.x
Weinstein GH, Wagman MJ (1984) Chromatography of antibiotics. Elsevier, Amsterdam
Wiegand I, Hilpert K, Hancock RE (2008) Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc 3(2):163–175. doi:10.1038/nprot.2007.521
Wilson DN (2014) Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat Rev Microbiol 12(1):35–48. doi:10.1038/nrmicro3155
Wilson ZE, Brimble MA (2009) Molecules derived from the extremes of life. Nat Prod Rep 26(1):44–71. doi:10.1039/B800164M
Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M et al (2006) DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res 34(Database issue):D668-72. doi:10.1093/nar/gkj067
Wong FK, Lau MC, Lacap DC, Aitchison JC, Cowan DA, Pointing SB (2010) Endolithic microbial colonization of limestone in a high-altitude arid environment. Microb Ecol 59(4):689–699. doi:10.1007/s00248-009-9607-8
Zheng L, Chen H, Han X, Lin W, Yan X (2005) Antimicrobial screening and active compound isolation from marine bacterium NJ6-3-1 associated with the sponge Hymeniacidon perleve. World J Microbiol Biotechnol 21(2):201–206. doi:10.1007/s11274-004-3318-6
Ziervogel BK, Roux B (2013) The binding of antibiotics in OmpF porin. Structure 21(1):76–87. doi:10.1016/j.str.2012.10.014
Acknowledgements
The NMR facility was provided by SAIF, N.E.H.U, Shillong. The UV–visible spectroscopic facility was provided by Central Instrumentation Facility, School of Life Sciences, N.E.H.U, Shillong and FT-IR facility was provided by CIF, I.A.S.S.T, Guwahati. We are also grateful to Dr. S. Sangilipandi, Centre for Advanced Studies in Chemistry, N.E.H.U, Shillong for the help during TLC and purification studies.
Authors’ contribution
KB and SJ designed and carried out the experiments, analyzed the data and drafted the manuscript. NP and KR performed the MS experiment and helped in analyzing the spectrometry data and structure elucidation. SK and PP conducted the in silico docking studies. SJ supervised the whole research work and revised the manuscript. All authors read and approved the final manuscript.
Funding
This research was partly supported by the Department of Science & Technology, Government of India FIST programme to the parent department. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. KB is thankful to UGC, New Delhi for providing financial support in the form of research fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Research involving human and animal participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhattacharjee, K., Kumar, S., Palepu, N.R. et al. Structure elucidation and in silico docking studies of a novel furopyrimidine antibiotics synthesized by endolithic bacterium Actinomadura sp. AL2. World J Microbiol Biotechnol 33, 178 (2017). https://doi.org/10.1007/s11274-017-2343-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-017-2343-1