Abstract
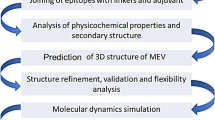
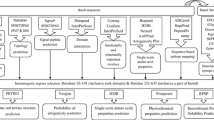
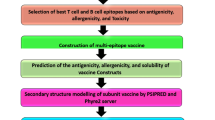
Acinetobacter baumannii is a well-recognized cause of nosocomial infections. This organism is recognized to be among the most difficult antimicrobial-resistant gram-negative bacilli to control and treat. One of the main challenges we face is Carbapenem resistance in A. baumannii. Carbapenem resistance in A. baumannii associated with the loss of an outer membrane protein designated CarO (Carbapenem resistance outer membrane protein). This protein is a membrane porin of A. baumannii. Using specific antibodies against this protein exert a bacteriostatic or bactericidal effect in vitro. Attempts should be made to discover peptides that could mimic protein epitopes and possess the same immunogenicity as the complete protein. Subsequently, bioinformatics methods for epitope prediction have been developed leading to synthesis of such peptides that are important for development of vaccine. This study provides a basis for the design of pathogen specifically, B cell epitope-based vaccine that is targeted to diseases caused by A. baumannii in the global human population. A combination of available bioinformatics tools are used to understand and characterize the Baumannii Acinetobactin utilization structure of A. baumannii and appropriate selection regions as effective B cell epitopes and functional exposed amino acids. In conclusion, amino acids 19–158 were selected as vaccine candidate.












Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Ansari HR, Raghava GP (2010) Identification of conformational B-cell epitopes in an antigen from its primary sequence. Immunome Res 6:6
Ashkenazy H, Erez E, Martz E, Pupko T, Ben-Tal N (2010) ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res 38:W529–W533
Bagos PG, Liakopoulos TD, Spyropoulos IC, Hamodrakas SJ (2004) PRED-TMBB: a web server for predicting the topology of β-barrel outer membrane proteins. Nucleic Acids Res 32:W400–W404
Bazmara H, Rasooli I, Jahangiri A, Sefid F, Astaneh SDA, Payandeh Z (2019) Antigenic properties of iron regulated proteins in Acinetobacter baumannii: an in silico approach. Int J Pept Res Ther 25:205–213
Berezin C et al (2004) ConSeq: the identification of functionally and structurally important residues in protein sequences. Bioinformatics 20:1322–1324
Blue-Hnidy DE, Allen SD (2006) Microbiology for the surgical pathologist. In: Essentials of anatomic pathology. Springer, New York, pp 155–229
Bryson K, Cozzetto D, Jones DT (2007) Computer-assisted protein domain boundary prediction using the Dom-Pred server. Curr Protein Pept Sci 8:181–188
Catel-Ferreira M et al (2011) Structure–function relationships of CarO, the carbapenem resistance-associated outer membrane protein of Acinetobacter baumannii. J Antimicrob Chemother 66:2053–2056
Dehghani E, Sefid F (2016) Evaluation of lamb linear and conformational epitopes on the vibrio Harveyi. Int J Adv Biotechnol Res 7(3):667–671
Dijkshoorn L, Nemec A, Seifert H (2007) An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat Rev Microbiol 5:939–951
El-Manzalawy Y, Dobbs D, Honavar V (2008) Predicting linear B-cell epitopes using string kernels. J Mol Recognit 21:243–255
Esmaeilkhani H, Rasooli I, Hashemi M, Nazarian S, Sefid F (2019) Immunogenicity of cork and loop domains of recombinant Baumannii acinetobactin utilization protein in murine model. Avicenna J Med Biotechnol 11:180
Fattahian Y, Rasooli I, Gargari SLM, Rahbar MR, Astaneh SDA, Amani J (2011) Protection against Acinetobacter baumannii infection via its functional deprivation of biofilm associated protein (Bap). Microb Pathog 51:402–406
García-Patiño MG, García-Contreras R, Licona-Limón P (2017) The immune response against Acinetobacter baumannii, an emerging pathogen in nosocomial infections. Front Immunol 8:441
Gordon NC, Wareham DW (2010) Multidrug-resistant Acinetobacter baumannii: mechanisms of virulence and resistance. Int J Antimicrob Agents 35:219–226
Higgins PG, Dammhayn C, Hackel M, Seifert H (2009) Global spread of carbapenem-resistant Acinetobacter baumannii. J Antimicrob Chemother 65:233–238
Hirokawa T, Boon-Chieng S, Mitaku S (1998) SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14:378–379
Jahangiri A, Rasooli I, Owlia P, Fooladi AAI, Salimian J (2017) In silico design of an immunogen against Acinetobacter baumannii based on a novel model for native structure of outer membrane protein A. Microb Pathog 105:201–210
Jespersen MC, Peters B, Nielsen M, Marcatili P (2017) BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res 45(W1):W24–W29
Kang AD, Smith KP, Eliopoulos GM, Berg AH, McCoy C, Kirby JE (2017) Invitro apramycin activity against multidrug-resistant Acinetobacter baumannii and Pseudomonas aeruginosa. Diagn Microbiol Infect Dis 88:188–191
Kempf M et al (2012) Cell surface properties of two differently virulent strains of Acinetobacter baumannii isolated from a patient. Can J Microbiol 58:311–317
Knezevic P, Aleksic V, Simin N, Svircev E, Petrovic A, Mimica-Dukic N (2016) Antimicrobial activity of Eucalyptus camaldulensis essential oils and their interactions with conventional antimicrobial agents against multi-drug resistant Acinetobacter baumannii. J Ethnopharmacol 178:125–136
Kringelum JV, Lundegaard C, Lund O, Nielsen M (2012) Reliable B cell epitope predictions: impacts of method development and improved benchmarking. PLoS Comput Biol 8:e1002829
Livermore DM, Mushtaq S, Warner M, Woodford N (2016) In-vitro activity of eravacycline against carbapenem-resistant Enterobacteriaceae and Acinetobacter baumannii. Antimicrob Agents Chemother 60(6):3840–3844
Longo DL (2006) Idiotype vaccination in follicular lymphoma: knocking on the doorway to cure. Oxford University Press, Oxford
McConnell MJ, Domínguez-Herrera J, Smani Y, López-Rojas R, Docobo-Pérez F, Pachón JJI (2011) Vaccination with outer membrane complexes elicits rapid protective immunity to multidrug-resistant Acinetobacter baumannii. Infect Immunity 79:518–526
Mussi MA, Limansky AS, Viale AM (2005) Acquisition of resistance to carbapenems in multidrug-resistant clinical strains of Acinetobacter baumannii: natural insertional inactivation of a gene encoding a member of a novel family of β-barrel outer membrane proteins. Antimicrob Agents Chemother 49:1432–1440
Mussi MA, Relling VM, Limansky AS, Viale AM (2007) CarO, an Acinetobacter baumannii outer membrane protein involved in carbapenem resistance, is essential for l-ornithine uptake. FEBS Lett 581:5573–5578
Negi SS, Schein CH, Oezguen N, Power TD, Braun W (2007) InterProSurf: a web server for predicting interacting sites on protein surfaces. Bioinformatics 23:3397–3399
Noori E, Rasooli I, Owlia P, Gargari SLM, Ebrahimizadeh W (2014) A conserved region from biofilm associated protein as a biomarker for detection of Acinetobacter baumannii. Microb Pathog 77:84–88
Notredame C, Higgins DG, Heringa J (2000) T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217
Pagdepanichkit S, Tribuddharat C, Chuanchuen RJ (2016) Distribution and expression of the Ade multidrug efflux systems in Acinetobacter baumannii clinical isolates. Can J Microbiol 62:794–801
Parkin J, Cohen B (2001) An overview of the immune system. The Lancet 357:1777–1789
Peleg AY, Seifert H, Paterson DL (2008) Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev 21:538–582
Peters B et al (2005) The immune epitope database and analysis resource: from vision to blueprint. PLoS Biol 3:e91
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Pirovano W, Feenstra KA, Heringa J (2008) PRALINE™: a strategy for improved multiple alignment of transmembrane proteins. Bioinformatics 24:492–497
Poirel L, Nordmann P (2006) Carbapenem resistance in Acinetobacter baumannii: mechanisms and epidemiology. Clin Microbiol Infect 12:826–836
Ponomarenko J, Bui H-H, Li W, Fusseder N, Bourne PE, Sette A, Peters B (2008) ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinform 9:514
Rahbar MR et al (2012) A potential in silico antibody–antigen based diagnostic test for precise identification of Acinetobacter baumannii. J Theoret Biol 294:29–39
Reimer U (2009) Prediction of linear B-cell epitopes epitope mapping protocols. Methods Mol Biol 524:335–344
Rubinstein ND, Mayrose I, Halperin D, Yekutieli D, Gershoni JM, Pupko T (2008) Computational characterization of B-cell epitopes. Mol Immunol 45:3477–3489
Rubinstein ND, Mayrose I, Martz E, Pupko T (2009) Epitopia: a web-server for predicting B-cell epitopes. BMC Bioinform 10:287
Saha S, Raghava G (2004) BcePred: prediction of continuous B-cell epitopes in antigenic sequences using physico-chemical properties. In: Nicosia G, Cutello V, Bentley PJ, Timmis J (eds) Artificial immune systems. ICARIS 2004. Lecture notes in computer science, vol 3239. Springer, Berlin, Heidelberg, pp 197–204
Saha S, Raghava GP (2007) Prediction methods for B-cell epitopes. Methods Mol Biol 409:387–394
Sánchez-Encinales V et al (2017) Overproduction of outer membrane protein A by Acinetobacter baumannii as a risk factor for nosocomial pneumonia, bacteremia, and mortality rate increase. J Infect Dis 215:966–974
Sangroodi YH, Rasooli I, Nazarian S, Ebrahimizadeh W, Sefid F (2015) Immunogenicity of conserved cork and ß-barrel domains of baumannii acinetobactin utilization protein in an animal model. Turkish J Med Sci 45:1396–1402
Sefid F, Rasooli I, Jahangiri A (2013) In silico determination and validation of baumannii acinetobactin utilization a structure and ligand binding site. BioMed Res Int 2013:172784
Sefid F, Rasooli I, Jahangiri A, Bazmara H (2015) Functional exposed amino acids of BauA as potential immunogen against Acinetobacter baumannii. Acta Biotheor 63:129–149
Shaker MA, Shaaban MI (2017) Formulation of carbapenems loaded gold nanoparticles to combat multi-antibiotic bacterial resistance: in vitro antibacterial study. Int J Pharm 525:71–84
Siroy A et al (2005) Channel formation by CarO, the carbapenem resistance-associated outer membrane protein of Acinetobacter baumannii. Antimicrob Agents Chemother 49:4876–4883
Tan KP, Varadarajan R, Madhusudhan MS (2011) DEPTH: a web server to compute depth and predict small-molecule binding cavities in proteins. Nucleic Acids Res 39:W242–W248
Viklund H, Bernsel A, Skwark M, Elofsson A (2008) SPOCTOPUS: a combined predictor of signal peptides and membrane protein topology. Bioinformatics 24:2928–2929
Vita R et al (2014) The immune epitope database (IEDB) 3.0. Nucleic Acids Res 43:D405–D412
Weinstein RA, Gaynes R, Edwards JR (2005) Overview of nosocomial infections caused by gram-negative bacilli. Clin Infect Dis 41:848–854
Wong D, Nielsen TB, Bonomo RA, Pantapalangkoor P, Luna B, Spellberg B (2017) Clinical and pathophysiological overview of Acinetobacter infections: a century of challenges. Clin Microbiol Rev 30:409–447
Yao B, Zhang L, Liang S, Zhang C (2012) SVMTriP: a method to predict antigenic epitopes using support vector machine to integrate tri-peptide similarity and propensity. PLoS ONE 7:e45152
Zahn M, D’Agostino T, Eren E, Baslé A, Ceccarelli M, van den Berg BJ (2015) Small-molecule transport by CarO, an abundant eight-stranded β-barrel outer membrane protein from Acinetobacter baumannii. J Mol Biol 427:2329–2339
Zdobnov EM, Apweiler R (2001) InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17:847–848
Zhang Z, Li Y, Lin B, Schroeder M, Huang BJB (2011) Identification of cavities on protein surface using multiple computational approaches for drug binding site prediction. Bioinformatics 27:2083–2088
Acknowledgements
We thank Yazd University for granting us a permission to use its web server.
Funding
This study was supported by an unrestricted free access to Yazd University web site for data collection. The authors have received no financial support for the elaboration of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Yazd University did not play any decision-making role in the study analysis or writing of the manuscript. All authors declare no Potential Conflicts of Interest.
Ethical Approval
As the present study involved no experimental with animals or human, hence there was no need for approval by the Ethics Committee of Yazd University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tohidinia, M., Moshtaghioun, S.M., Sefid, F. et al. Functional Exposed Amino Acids of CarO Analysis as a Potential Vaccine Candidate in Acinetobacter Baumannii. Int J Pept Res Ther 26, 1185–1197 (2020). https://doi.org/10.1007/s10989-019-09923-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-019-09923-2