Abstract
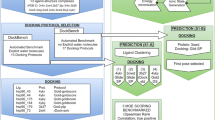
Drug Design Data Resource (D3R) continues to release valuable benchmarking datasets to promote improvement and development of computational methods for new drug discovery. We have developed several methods for protein–ligand binding mode prediction during the participation in the D3R challenges. In the present study, these methods were integrated, automated, and systematically tested using the large-scale data from Continuous Evaluation of Ligand Pose Prediction (CELPP) and a subset of Grand challenge 3 (GC3). The results show that current molecular docking methods benefit from the increasing number of protein–ligand complex structures deposited in Protein Data Bank. Using an appropriate protein structure for docking significantly improves the success rate of the binding mode prediction. The results of our template-based method and docking method are compared and discussed. Our future direction include the combination of these two methods for binding mode prediction.



Similar content being viewed by others
References
Kitchen DB, Decornez H, Furr JR, Bajorath J (2004) Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov 3:935–947
Grinter SZ, Zou X (2014) Challenges, applications, and recent advances of protein-ligand docking in structure-based drug design. Molecules 19:10150–10176
Xu X, Huang M, Zou X (2018) Docking-based inverse virtual screening: methods, applications, and challenges. Biophys Rep 4:1–16
Brooijmans N, Kuntz ID (2003) Molecular recognition and docking algorithms. Ann Rev Biophys Biomol Struct 32(1):335–373
Huang SY, Grinter SZ, Zou X (2014) Scoring functions and their evaluation methods for protein-ligand docking: recent advances and future directions. Phys Chem Chem Phys 12:12899–12908
Gathiaka S, Liu S, Chiu M, Yang H, Stuckey JA, Kang YN, Delproposto J, Kubish G, Dunbar JB, Carlson HA, Burley SK (2016) D3R grand challenge 2015: evaluation of protein–ligand pose and affinity predictions. J Comput Aided Mol Des 30:651–668
Gaieb Z, Liu S, Gathiaka S, Chiu M, Yang H, Shao C, Feher VA, Walters WP, Kuhn B, Rudolph MG, Burley SK (2018) D3R Grand Challenge 2: blind prediction of protein–ligand poses, affinity rankings, and relative binding free energies. J Comput Aided Mol Des 32:1–20
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
Smith RD, Dunbar JB Jr, Ung PM et al (2011) CSAR benchmark exercise of 2010: combined evaluation across all submitted scoring functions. J Chem Inf Model 51:2115–2131
Damm-Ganamet KL, Smith RD, Dunbar JB Jr et al (2013) CSAR benchmark exercise 2011–2012: evaluation of results from docking and relative ranking of blinded congeneric series. J Chem Inf Model 53:1853–1870
Smith RD, Damm-Ganamet KL, Dunbar JB Jr et al (2016) CSAR benchmark exercise 2013: evaluation of results from a combined computational protein design, docking, and scoring/ranking challenge. J Chem Inf Model 56:1022–1031
Carlson HA, Smith RD, Damm-Ganamet KL et al (2016) CSAR 2014: a benchmark exercise using unpublished data from pharma. J Chem Inf Model 56:1063–1077
Xu X, Yan C, Zou X (2017) Improving binding mode and binding affinity predictions of docking by ligand-based search of protein conformations: evaluation in D3R grand challenge 2015. J Comput Aided Mol Des 31:689–699
Duan R, Xu X, Zou X (2018) Lessons learned from participating in D3R 2016 grand challenge 2: compounds targeting the farnesoid X receptor. J Comput Aided Mol Des 32:103–111
Yan C, Grinter SZ, Merideth BR, Ma Z, Zou X (2016) Iterative knowledge-based scoring functions derived from rigid and flexible decoy structures: evaluation with the 2013 and 2014 CSAR benchmarks. J Chem Inf Model 56:1013–1021
Grinter SZ, Yan C, Huang SY, Jiang L, Zou X (2013) Automated large-scale file preparation, docking, and scoring: Evaluation of ITScore and STScore using the 2012 community structure–activity resource benchmark. J Chem Inf Model 53:1905–1914
Huang SY, Zou X (2011) Scoring and lessons learned with the CSAR benchmark using an improved iterative knowledge-based scoring function. J Chem Inf Model 51:2097–2106
Huang SY, Zou X (2011) Construction and test of ligand decoy sets using MDock: community structure–activity resource benchmarks for binding mode prediction. J Chem Inf Model 51:2107–2114
Huang SY, Zou X (2007) Ensemble docking of multiple protein structures: considering protein structural variations in molecular docking. Proteins 66:399–421
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Huang S, Zou X (2006) An iterative knowledge-based scoring function to predict protein-ligand interactions: I. Derivation of interaction potentials. J Comput Chem 27:1866–1875
Huang S, Zou X (2006) An iterative knowledge-based scoring function to predict protein–ligand interactions: II. Validation of the scoring function. J Comput Chem 27:1876–1882
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinform 10:421
Liu X, Jiang H, Li H (2011) SHAFTS: a hybrid approach for 3D molecular similarity calculation. 1. Method and assessment of virtual screening. J Chem Inf Model 51:2372–2385
Hawkins PC, Skillman AG, Warren GL, Ellingson BA, Stahl MT (2010) Conformer generation with omega: algorithm and validation using high quality structures from the protein databank and cambridge structural database. J Chem Inf Model 50:572–584
Hawkins PC, Nicholls A (2012) Conformer generation with OMEGA: learning from the data set and the analysis of failures. J Chem Inf Model 52:2919–2936
Cheng T, Li X, Li Y, Liu Z, Wang R (2009) Comparative assessment of scoring functions on a diverse test set. J Chem Inf Model 49:1079–1093
Wang R, Fang X, Lu Y, Yang CY, Wang S (2005) The PDBbind database: methodologies and updates. J Med Chem 48:4111–4119
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Efron B (1979) Bootstrap methods: another look at the jackknife. Ann Stat 7:1–26
Pattengale ND, Alipour M, Bininda-Emonds OR, Moret BM, Stamatakis A (2010) How many bootstrap replicates are necessary? J Comput Biol 17:337–354
Acknowledgements
Support to XZ from OpenEye Scientific Software Inc. (Santa Fe, NM, http://www.eyesopen.com) is gratefully acknowledged. This work was supported by NIH R01GM109980 (PI: XZ), NIH R01HL126774 and NIH R01HL142301 (PI: Cui) to XZ. The computations were performed on the high performance computing infrastructure supported by NSF CNS-1429294 (PI: Chi-Ren Shyu) and the HPC resources supported by the University of Missouri Bioinformatics Consortium (UMBC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, X., Ma, Z., Duan, R. et al. Predicting protein–ligand binding modes for CELPP and GC3: workflows and insight. J Comput Aided Mol Des 33, 367–374 (2019). https://doi.org/10.1007/s10822-019-00185-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-019-00185-0